Abstract
The human glucagon receptor, GCGR, belongs to the class B G-protein-coupled receptor family and plays a key role in glucose homeostasis and the pathophysiology of type 2 diabetes. Here we report the 3.0 Å crystal structure of full-length GCGR containing both the extracellular domain and transmembrane domain in an inactive conformation. The two domains are connected by a 12-residue segment termed the stalk, which adopts a β-strand conformation, instead of forming an α-helix as observed in the previously solved structure of the GCGR transmembrane domain. The first extracellular loop exhibits a β-hairpin conformation and interacts with the stalk to form a compact β-sheet structure. Hydrogen–deuterium exchange, disulfide crosslinking and molecular dynamics studies suggest that the stalk and the first extracellular loop have critical roles in modulating peptide ligand binding and receptor activation. These insights into the full-length GCGR structure deepen our understanding of the signalling mechanisms of class B G-protein-coupled receptors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
19 June 2017
The Reviewer Information section was corrected.
References
Lagerström, M. C. & Schiöth, H. B. Structural diversity of G protein-coupled receptors and significance for drug discovery. Nat. Rev. Drug Discov. 7, 339–357 (2008)
Hollenstein, K. et al. Insights into the structure of class B GPCRs. Trends Pharmacol. Sci. 35, 12–22 (2014)
Finan, B. et al. Chemical hybridization of glucagon and thyroid hormone optimizes therapeutic impact for metabolic disease. Cell 167, 843–857.e14 (2016)
Longuet, C. et al. The glucagon receptor is required for the adaptive metabolic response to fasting. Cell Metab. 8, 359–371 (2008)
Parthier, C., Reedtz-Runge, S., Rudolph, R. & Stubbs, M. T. Passing the baton in class B GPCRs: peptide hormone activation via helix induction? Trends Biochem. Sci. 34, 303–310 (2009)
Koth, C. M. et al. Molecular basis for negative regulation of the glucagon receptor. Proc. Natl Acad. Sci. USA 109, 14393–14398 (2012)
Yang, L. et al. Conformational states of the full-length glucagon receptor. Nat. Commun. 6, 7859 (2015)
Siu, F. Y. et al. Structure of the human glucagon class B G-protein-coupled receptor. Nature 499, 444–449 (2013)
Jazayeri, A. et al. Extra-helical binding site of a glucagon receptor antagonist. Nature 533, 274–277 (2016)
Hollenstein, K. et al. Structure of class B GPCR corticotropin-releasing factor receptor 1. Nature 499, 438–443 (2013)
Song, G. et al. Human GLP-1 receptor transmembrane domain structure in complex with allosteric modulators. Nature http://dx.doi.org/10.1038/nature22378 (2017)
Grace, C. R. et al. NMR structure and peptide hormone binding site of the first extracellular domain of a type B1 G protein-coupled receptor. Proc. Natl Acad. Sci. USA 101, 12836–12841 (2004)
Pioszak, A. A., Parker, N. R., Suino-Powell, K. & Xu, H. E. Molecular recognition of corticotropin-releasing factor by its G-protein-coupled receptor CRFR1. J. Biol. Chem. 283, 32900–32912 (2008)
Ballesteros, J. & Weinstein, H. In Methods in Neuroscience Vol. 25 (ed, Sealfon, S. ) 366–428 (Elsevier, 1995)
Wootten, D., Simms, J., Miller, L. J., Christopoulos, A. & Sexton, P. M. Polar transmembrane interactions drive formation of ligand-specific and signal pathway-biased family B G protein-coupled receptor conformations. Proc. Natl Acad. Sci. USA 110, 5211–5216 (2013)
Mann, R., Wigglesworth, M. J. & Donnelly, D. Ligand-receptor interactions at the parathyroid hormone receptors: subtype binding selectivity is mediated via an interaction between residue 23 on the ligand and residue 41 on the receptor. Mol. Pharmacol. 74, 605–613 (2008)
Unson, C. G. et al. Roles of specific extracellular domains of the glucagon receptor in ligand binding and signaling. Biochemistry 41, 11795–11803 (2002)
Roberts, D. J., Vertongen, P. & Waelbroeck, M. Analysis of the glucagon receptor first extracellular loop by the substituted cysteine accessibility method. Peptides 32, 1593–1599 (2011)
Byrne, E. F. et al. Structural basis of Smoothened regulation by its extracellular domains. Nature 535, 517–522 (2016)
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protocols 4, 706–731 (2009)
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010)
Liu, W., Ishchenko, A. & Cherezov, V. Preparation of microcrystals in lipidic cubic phase for serial femtosecond crystallography. Nat. Protocols 9, 2123–2134 (2014)
Weierstall, U. et al. Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography. Nat. Commun. 5, 3309 (2014)
Liang, M. et al. The Coherent X-ray Imaging instrument at the Linac Coherent Light Source. J. Synchrotron Radiat. 22, 514–519 (2015)
Siewert, F. et al. Ultra-precise characterization of LCLS hard X-ray focusing mirrors by high resolution slope measuring deflectometry. Opt. Express 20, 4525–4536 (2012)
White, T. A. et al. Crystallographic data processing for free-electron laser sources. Acta Crystallogr. D Biol. Crystallogr. 69, 1231–1240 (2013)
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Crystallogr. 45, 335–341 (2012)
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D Biol. Crystallogr. 67, 271–281 (2011)
Duisenberg, A. J. M. Indexing in single-crystal diffractometry with an obstinate list of reflections. J. Appl. Crystallogr. 25, 92–96 (1992)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997)
Smart, O. S. et al. Exploiting structure similarity in refinement: automated NCS and target-structure restraints in BUSTER. Acta Crystallogr. D Biol. Crystallogr. 68, 368–380 (2012)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010)
Pascal, B. D. et al. HDX workbench: software for the analysis of H/D exchange MS data. J. Am. Soc. Mass Spectrom. 23, 1512–1521 (2012)
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013)
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010)
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007)
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals – a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981)
Miyamoto, S. & Kollman, P. A. Settle – an analytical version of the shake and rattle algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992)
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997)
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010)
Acknowledgements
This work was supported by the National Basic Research Program of China grants 2014CB910400 (B.W.) and 2015CB910304 (H.J.), CAS Strategic Priority Research Program XDB08020000 and CAS grant QYZDB-SSW-SMC024 (B.W.), the National Science Foundation of China grants 31422017 (B.W.), 81525024 (Q.Z.), 81230076 (H.J.) and 81573479 (D.Y.), the National Health and Family Planning Commission grants 2012ZX09304-011, 2013ZX09401003-005, 2013ZX09507001 and 2013ZX09507-002 (M.-W.W.), Shanghai Science and Technology Development Fund 15DZ2291600 (M.-W.W.), the National Institutes of Health grants R01 GM108635 (V.C.) and R21 DA042298 (W.L.), the National Science Foundation STC award 1231306 (G.N., U.W., W.L., T.D.G., V.C.), the GPCR Consortium (B.W., G.S., R.C.S.), Shanghai local government (G.S., R.C.S.), the Netherlands eScience Center (NLeSC)/NWO (Enabling Technologies project: 3D-e-Chem, grant 027.014.201) (C.d.G.), and C.d.G. participates in the European Cooperation in Science and Technology Action CM1207 (GLISTEN). The authors thank A. Walker for assistance with manuscript preparation, Y. Feng and C. Ji for technical assistance, M. Hunter, A. Batyuk, A. Ishchenko, L. Johansson, B. Stauch, M. Audet, M. Liang, M. Seaberg and P. Walter for their help with XFEL data collection, B. Yu for help with mAb1 protein sample preparation, and the special program for applied research on super computation of the NSFC-Guangdong Joint Fund (the second phase). The synchrotron radiation experiments were performed at the BL41XU of SPring-8 with approval of the Japan Synchrotron Radiation Research Institute (proposal no. 2016A2517, 2016A2518, 2016B2517 and 2016B2518). We thank the beamline staff members K. Hasegawa, H. Okumura, and H. Murakami of the BL41XU for help on X-ray data collection. Use of the LCLS, SLAC National Accelerator Laboratory, was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under contract no. DE-AC02-76SF00515. Parts of the sample injector used at LCLS for this research were funded by the National Institutes of Health, P41GM103393, formerly P41RR001209.
Author information
Authors and Affiliations
Contributions
Ha.Z. and A.Q. optimized the construct, developed the purification procedure and purified the GCGR proteins for crystallization, performed crystallization trials and optimized crystallization conditions. D.Y. designed, performed and analysed disulfide crosslinking, ligand binding and cAMP assays of wild-type and mutant GCGRs. L.Y. performed and analysed molecular dynamics simulations. V.D. performed and analysed HDX data. Hu.Z. collected the X-ray diffraction data at the synchrotron. G.W.H. helped with structure determination and refinement. T.D.G. processed the XFEL data. R.G.S., U.W., G.N. and W.L. helped with XFEL sample preparation and data collection. Y.W. helped with construct and crystal optimization. L.M. expressed the GCGR proteins. A.D., X.C. and G.L. helped with disulfide crosslinking and ligand binding assays. X.W. prepared mAb1 protein sample. Z.G. and Y.D. helped with data analysis. C.d.G., S.R.-R., G.S., J.L., V.C. and M.A.H. helped with structure analysis/interpretation and commented on the manuscript. P.R.G. oversaw HDX assay. V.C. oversaw collection of the XFEL data. H.Y. and H.J. oversaw molecular dynamics simulations. M.-W.W. oversaw the disulfide crosslinking, ligand binding and cAMP assays. R.C.S. helped to initiate the project. R.C.S, H.J. and M.-W.W. oversaw structure analysis/interpretation and edited the manuscript. B.W. and Q.Z. initiated the project, planned and analysed experiments, solved the structures, supervised the research and wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
S.R.-R., X.W. and J.L. are employees of Novo Nordisk, a pharmaceutical company focused on class B GPCRs for type 2 diabetes. R.C.S. is a founder and board member of Bird Rock Bio, a company focused on GPCR therapeutic antibodies. The other authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks G. Lebon, M. Sansom, T. Schwartz and C. Siebold for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Snake plot of the GCGR construct used for crystallization and crystal packing of the GCGR–NNC0640–mAb1 complex structure.
a, Snake plot of the GCGR–T4L fusion construct used for crystallization. The eight cysteine residues forming disulfide bonds are shown in yellow with four yellow lines illustrating the disulfide bonds. b, Crystal packing of the GCGR–NNC0640–mAb1 complex structure. GCGR and mAb1 are shown in cartoon representation. The ECD, stalk and TMD of the receptor are coloured in orange, green and blue, respectively. The T4L fusion is in grey and mAb1 is in cyan. NNC0640 is displayed as magenta spheres. The unit cell is shown as a red box.
Extended Data Figure 2 Binding of [3H]-NNC0640 and 125I-labelled glucagon to wild-type and mutant GCGRs and glucagon-induced cAMP assays.
a, Binding of [3H]-NNC0640 to membrane preparations from Sf9 cells expressing wild-type (WT) and the engineered GCGR used for crystallization. Data are shown as means ± s.e.m. from three independent experiments performed in duplicate. ‘Construct’ indicates the construct used for crystallization, containing the T4L fusion at ICL2 and 45 residues truncated at the C terminus of the receptor. b, Binding of [3H]-NNC0640 to membrane preparations from HEK293T cells expressing wild-type or mutant GCGRs. Data are shown as means ± s.e.m. from three independent experiments performed in duplicate. The IC50 values for the wild-type and mutant GCGRs from at least three independent experiments are listed in Extended Data Table 2. c, Glucagon-induced cAMP accumulation measurement of the mutants V130C/L210C, V130C and L210C and the wild-type GCGR. d, Glucagon-induced cAMP accumulation measurement of the mutants V130C/L210C, V130C and L210C and the wild-type GCGR in the presence of 1 mM DTT. Dose–response curves of cAMP accumulation assays generated from three independent experiments performed in duplicate. Data are shown as means ± s.e.m. e–g, Disulfide cross linking assays of the GCGR mutant Q113C/D209C (e) and the controls, single-site mutants Q113C (f) and D209C (g). Dose–response curves of 125I-labelled-glucagon-binding assay generated from three independent experiments performed in duplicate. Data are shown as means ± s.e.m.
Extended Data Figure 3 Electron densities for the stalk and ECL1 in the full-length GCGR crystal structure.
a, Electron densities for the stalk. The stalk region is shown as sticks and coloured in green. The rest of the receptor is shown in cartoon representation and coloured in orange (ECD), magenta (ECL1) and blue (TMD). Electron densities are contoured at 1.0σ from a composite omit map and coloured in blue. b, Electron densities for ECL1. ECL1 is shown as sticks and coloured in magenta. The rest of the receptor is shown in cartoon representation and coloured in orange (ECD), green (stalk) and blue (TMD). Electron densities are contoured at 1.0σ from a composite omit map and coloured in blue.
Extended Data Figure 4 Comparison between the full-length GCGR crystal structure and previously published structure and model.
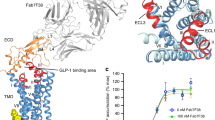
a, b, Comparison between the GCGR–mAb1 complex structure and the previously published model of the GCGR–glucagon complex7. Only the receptors in the GCGR–mAb1 complex structure and the model are shown in cartoon representation and coloured in blue and yellow, respectively. The ECDs are also shown in surface representation. The stalk in the full-length GCGR structure is in green, and the stalk in the model is coloured magenta. a, Side view; b, top view. c, Comparison between the full-length GCGR structure and crystal structure of the ECD of GLP-1R bound to its endogenous ligand GLP-1. Structural superimposition shows spatial clashes between ECL1 and helix II of GCGR and GLP-1. The receptor in the full-length GCGR structure is shown as blue cartoon, and the ligand NNC0640 is displayed as magenta spheres. The complex structure of the ECD of GLP-1R bound to GLP-1 (PDB ID: 3IOL) is shown in cartoon representation. The ECD of GLP-1R is coloured green, and GLP-1 is in red.
Extended Data Figure 5 HDX studies for the NNC0640-stabilized GCGR in complex with mAb1 or mAb23.
a, Interaction between GCGR and mAb1. The receptor is shown in grey cartoon representation. The regions of αA helix (residues L32–L38) and β4–L5 (residues K98–Q105) in the ECD and stalk (residues I128–M137), which showed increased protection in the antibody-bound GCGRs, are coloured in red, blue and green, respectively. The antibody mAb1 is shown as cyan surface and cartoon. b–d, HDX plots for the regions of the ECD αA helix (b), ECD β4–L5 (c) and the stalk (d) in the antibody-free and mAb1-bound GCGRs. e–g, HDX plots for the regions of the ECD αA helix (e), ECD β4–L5 (f) and the stalk (g) in the antibody-free and mAb23-bound GCGRs. HDX data are plotted as means ± s.d. of three independent experiments.
Extended Data Figure 6 Differential perturbation heat map view of the HDX studies.
a, Heat map view of the GCGR–NNC0640–mAb1 complex coloured according to the heat map colouring scheme used by the software HDX Workbench36. Each bar represents a peptide showing the average difference (across six time points) in D2O uptake between the receptor–antibody complex and the antibody-free receptor with the standard deviation between replicates and the peptide charge states shown in parentheses. The regions that revealed statistically significant reductions in deuterium uptake in the receptor–antibody complex compared to the antibody-free receptor are coloured green and boxed in red. The D2O difference between the antibody-bound and antibody-free GCGRs at two consecutive time points has a P value < 0.05 or a single time point has a P value < 0.01. The regions with no significant change are in grey and the regions that have no peptides covering the sequence in MSMS and HDX runs are shown as gaps. b, Heat map view of the GCGR–NNC0640–mAb23 complex.
Extended Data Figure 7 MD simulations of the apo GCGR.
a, Main chain r.m.s.d. values of the ECD versus simulation time in the three 1-μs molecular dynamics simulations. The values were calculated from snapshots at 100-ps intervals. All the structures were superimposed to the crystal structure of full-length GCGR using the main chain atoms of residues S150–L160 (helix I), I176–V193 (helix II), A227–G246 (helix III), G271–P275 (helix IV), V311–I321 (helix V), T351–L358 (helix VI) and Q392–Y400 (helix VII). b–d, Comparison between the results of simulations and the full-length GCGR crystal structure. The full-length GCGR crystal structure is shown as grey cartoon. The results of the three simulations are shown in cartoon representation, and coloured in yellow, cyan and orange, respectively. The ECDs of the receptors are also shown in surface representation. The N-terminal portion of stalk (residues G125–Q131) and the ECL1 region (residues T200–D218), which are analysed in panel e, are coloured green and magenta, respectively. The red arrows indicate the movements of the ECDs and ECL1 (d) in the simulations. e, Secondary structure as a function of time for the stalk (residues G125–Q131) and ECL1 (residues T200–D218) regions in the crystal structure and simulations.
Extended Data Figure 8 Interactions between the ECD and stalk/ECL1 in molecular dynamics simulations and comparison with the binding modes of glucagon and mAb1.
a, Interactions between the ECD and stalk/ECL1 in one typical molecular dynamics simulation snapshot. The residues Y202, K205 and I206 on the N-terminal half of ECL1 make hydrophobic contacts with a hydrophobic core formed by residues L32, F33, W36, Y65, Y84, L85, P86 and W87 on the αA helix, L2 and L5 of the ECD. Additionally, the negatively charged residues E127 and E129 on the stalk and D208 and D209 on the tip of ECL1 tend to form salt bridges with the basic residues R111 and R116 on the L5 of the ECD. The receptor is shown in cartoon representation. The ECD, stalk, ECL1 and TMD are coloured orange, green, magenta and blue, respectively. The residues involved in the interaction are shown as sticks. b–d, Interaction interfaces on the ECD for the stalk/ECL1 in the molecular dynamics simulations (b), glucagon in the GCGR–glucagon complex model7 (c) and mAb1 in the GCGR–NNC0640–mAb1 complex structure (d). The ECDs of the receptor are shown as grey cartoon and surface. The residues involved in the interactions are shown as sticks and coloured orange.
Rights and permissions
About this article
Cite this article
Zhang, H., Qiao, A., Yang, D. et al. Structure of the full-length glucagon class B G-protein-coupled receptor. Nature 546, 259–264 (2017). https://doi.org/10.1038/nature22363
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22363
This article is cited by
-
G protein-coupled receptors (GPCRs): advances in structures, mechanisms, and drug discovery
Signal Transduction and Targeted Therapy (2024)
-
Cryo-electron microscopy for GPCR research and drug discovery in endocrinology and metabolism
Nature Reviews Endocrinology (2024)
-
Molecular features of the ligand-free GLP-1R, GCGR and GIPR in complex with Gs proteins
Cell Discovery (2024)
-
Structure-based drug discovery of a corticotropin-releasing hormone receptor 1 antagonist using an X-ray free-electron laser
Experimental & Molecular Medicine (2023)
-
Tail engagement of arrestin at the glucagon receptor
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.