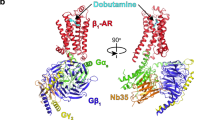
Abstract
G-protein-coupled receptor (GPCR)-mediated signal transduction is central to human physiology and disease intervention, yet the molecular mechanisms responsible for ligand-dependent signalling responses remain poorly understood. In class A GPCRs, receptor activation and G-protein coupling entail outward movements of transmembrane helix 6 (TM6). Here, using single-molecule fluorescence resonance energy transfer imaging, we examine TM6 movements in the β2 adrenergic receptor (β2AR) upon exposure to orthosteric ligands with different efficacies, in the absence and presence of the Gs heterotrimer. We show that partial and full agonists differentially affect TM6 motions to regulate the rate at which GDP-bound β2AR–Gs complexes are formed and the efficiency of nucleotide exchange leading to Gs activation. These data also reveal transient nucleotide-bound β2AR–Gs species that are distinct from known structures, and provide single-molecule perspectives on the allosteric link between ligand- and nucleotide-binding pockets that shed new light on the G-protein activation mechanism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Pierce, K. L., Premont, R. T. & Lefkowitz, R. J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell Biol. 3, 639–650 (2002)
Vilardaga, J. P. et al. GPCR and G proteins: drug efficacy and activation in live cells. Mol. Endocrinol. 23, 590–599 (2009)
Kenakin, T. New concepts in pharmacological efficacy at 7TM receptors: IUPHAR review 2. Br. J. Pharmacol. 168, 554–575 (2013)
Manglik, A. & Kobilka, B. The role of protein dynamics in GPCR function: insights from the β2AR and rhodopsin. Curr. Opin. Cell Biol. 27, 136–143 (2014)
Baker, J. G. The selectivity of β-adrenoceptor agonists at human β1-, β2- and β3-adrenoceptors. Br. J. Pharmacol. 160, 1048–1061 (2010)
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011)
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013)
Huang, W. et al. Structural insights into μ-opioid receptor activation. Nature 524, 315–321 (2015)
Carpenter, B., Nehmé, R., Warne, T., Leslie, A. G. & Tate, C. G. Structure of the adenosine A2A receptor bound to an engineered G protein. Nature 536, 104–107 (2016)
Yao, X. J. et al. The effect of ligand efficacy on the formation and stability of a GPCR–G protein complex. Proc. Natl Acad. Sci. USA 106, 9501–9506 (2009)
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015)
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013)
Cooper, M. et al. Cy3B: improving the performance of cyanine dyes. J. Fluoresc. 14, 145–150 (2004)
Vafabakhsh, R., Levitz, J. & Isacoff, E. Y. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015)
Kim, H. D. et al. Mg2+-dependent conformational change of RNA studied by fluorescence correlation and FRET on immobilized single molecules. Proc. Natl Acad. Sci. USA 99, 4284–4289 (2002)
Lamichhane, R. et al. Single-molecule view of basal activity and activation mechanisms of the G protein-coupled receptor β2AR. Proc. Natl Acad. Sci. USA 112, 14254–14259 (2015)
Bond, R. A. et al. Physiological effects of inverse agonists in transgenic mice with myocardial overexpression of the β2-adrenoceptor. Nature 374, 272–276 (1995)
Ferguson, A. et al. Functional dynamics within the human ribosome regulate the rate of active protein synthesis. Mol. Cell 60, 475–486 (2015)
Galés, C. et al. Real-time monitoring of receptor and G-protein interactions in living cells. Nat. Methods 2, 177–184 (2005)
Ernst, O. P., Gramse, V., Kolbe, M., Hofmann, K. P. & Heck, M. Monomeric G protein-coupled receptor rhodopsin in solution activates its G protein transducin at the diffusion limit. Proc. Natl Acad. Sci. USA 104, 10859–10864 (2007)
Traut, T. W. Physiological concentrations of purines and pyrimidines. Mol. Cell. Biochem. 140, 1–22 (1994)
Hein, P. et al. Gs activation is time-limiting in initiating receptor-mediated signaling. J. Biol. Chem. 281, 33345–33351 (2006)
Galés, C. et al. Probing the activation-promoted structural rearrangements in preassembled receptor-G–protein complexes. Nat. Struct. Mol. Biol. 13, 778–786 (2006)
Qin, K., Dong, C., Wu, G. & Lambert, N. A. Inactive-state preassembly of Gq-coupled receptors and Gq heterotrimers. Nat. Chem. Biol. 7, 740–747 (2011)
Westfield, G. H. et al. Structural flexibility of the Gαs α-helical domain in the β2-adrenoceptor Gs complex. Proc. Natl Acad. Sci. USA 108, 16086–16091 (2011)
Damian, M. et al. Ghrelin receptor conformational dynamics regulate the transition from a preassembled to an active receptor:Gq complex. Proc. Natl Acad. Sci. USA 112, 1601–1606 (2015)
Murayama, T. & Ui, M. [3H]GDP release from rat and hamster adipocyte membranes independently linked to receptors involved in activation or inhibition of adenylate cyclase. Differential susceptibility to two bacterial toxins. J. Biol. Chem. 259, 761–769 (1984)
Ceruso, M. A., Periole, X. & Weinstein, H. Molecular dynamics simulations of transducin: interdomain and front to back communication in activation and nucleotide exchange. J. Mol. Biol. 338, 469–481 (2004)
Herrmann, R. et al. Sequence of interactions in receptor-G–protein coupling. J. Biol. Chem. 279, 24283–24290 (2004)
Herrmann, R. et al. Rhodopsin-transducin coupling: role of the Gα C-terminus in nucleotide exchange catalysis. Vision Res. 46, 4582–4593 (2006)
Oldham, W. M., Van Eps, N., Preininger, A. M., Hubbell, W. L. & Hamm, H. E. Mechanism of the receptor-catalyzed activation of heterotrimeric G proteins. Nat. Struct. Mol. Biol. 13, 772–777 (2006)
Kapoor, N., Menon, S. T., Chauhan, R., Sachdev, P. & Sakmar, T. P. Structural evidence for a sequential release mechanism for activation of heterotrimeric G proteins. J. Mol. Biol. 393, 882–897 (2009)
Kaya, A. I. et al. A conserved phenylalanine as a relay between the α5 helix and the GDP binding region of heterotrimeric Gi protein α subunit. J. Biol. Chem. 289, 24475–24487 (2014)
Dror, R. O. et al. Signal transduction. Structural basis for nucleotide exchange in heterotrimeric G proteins. Science 348, 1361–1365 (2015)
Flock, T. et al. Universal allosteric mechanism for Gα activation by GPCRs. Nature 524, 173–179 (2015)
Acknowledgements
We thank M. Howarth for the gift of trans-divalent streptavidin, and C. Stern in the laboratory of J. Chodera for constructing the CHARMM-consistent parameters for the dyes used in the molecular dynamics simulations. Computational resources are gratefully acknowledged: an XSEDE allocation at the Texas Advanced Computing Center at the University of Texas at Austin (Stampede supercomputer, project TG MCB120008), support from resources at the Oak Ridge Leadership Computing Facility (ALCC allocation BIP109) at the Oak Ridge National Laboratory that is supported by the Office of Science of the US Department of Energy under contract no. DE-AC05-00OR22725; and the resources of the David A. Cofrin Center for Biomedical Information in the HRH Prince Alwaleed Bin Talal Bin Abdulaziz Alsaud Institute for Computational Biomedicine at Weill Cornell Medicine. This work was supported in part by National Institutes of Health (NIH) grants GM098859 (S.C.B.), R21DA0354585 (J.A.J., S.C.B. and G.G.G.), K05DA022413 and R01 MH54137 (J.A.J.), R01GM083118 and R01NS028471 (B.K.K.), and U54GM087519 (H.W. and J.M.P.-A.), the German Academic Exchange Service (DAAD) (D.H.), the American Heart Association Postdoctoral fellowship (15POST22700020) (M.M.), and the Novo Nordisk Foundation Center for Basic Metabolic Research (M.H.).
Author information
Authors and Affiliations
Contributions
G.G.G., M.M., D.H., B.K.K. and S.C.B. designed single-molecule experiments. G.G.G. labelled receptor and performed all single-molecule experiments. G.G.G. analysed single-molecule data, with support from D.S.T. M.J. and D.S.T. developed the imaging and analysis platform. M.M. expressed, purified and characterized receptor constructs. D.H. expressed, purified and biotinylated Gs, and performed GTP turnover assays. H.Z. and Z.Z. synthesized the fluorophores. J.M.P.-A. performed molecular dynamics simulations under the supervision of H.W. M.H. and S.M. performed cell-based G-protein-coupling assays under the supervision of J.A.J. G.G.G., M.M., D.H., J.A.J., H.W., B.K.K. and S.C.B. interpreted all the data and wrote the manuscript. B.K.K. and S.C.B. provided overall project supervision.
Corresponding authors
Ethics declarations
Competing interests
S.C.B. has an equity interest in Lumidyne Technologies.
Additional information
Reviewer Information Nature thanks M. Lohse and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Ligand-binding properties of β2∆6-148C/266C.
a, [3H]dihydroalprenolol (DHA) saturation binding on purified β2AR, comparing wild type (WT; blue squares) versus unlabelled (orange circles) and labelled (green triangles) β2∆6-148C/266C. Affinities (Kd values) are shown in the table. Non-specific binding controls are shown as corresponding open symbols. Error bars denote the s.e.m. from triplicate measurements. b, [3H]DHA–isoproterenol competition binding on purified β2AR, comparing wild type (blue circles) versus unlabelled (orange squares) and labelled (green triangles) β2∆6-148C/266C. Affinities (Ki values) are reported in the table. Error bars represent the s.e.m. from triplicate measurements. c, Dose–response curves of the BRET-based cAMP biosensor CAMYEL with wild-type β2AR (black triangles) and the mutants β2∆6 (blue squares) and β2∆6-148C/266C (red circles). Data from untransfected cells are shown in green triangles. Data from five independent experiments were normalized to the maximal isoproterenol response by wild-type β2AR in each experiment and globally fit to the entire dataset, with the error bars representing the s.e.m., as reported in the table. d, Skeletal structures of ligands used in the current study. e, [3H]DHA competition binding on unlabelled β2∆6-148C/266C for ligands used in the current study, except for carazolol and BI-167107, for which the ultra-high affinities reported (32 pM and 84 pM, respectively) would not allow us to determine them accurately in our assay. Instead, we used concentrations of 1 μM for both in all our measurements. The calculated Ki values are shown on the table.
Extended Data Figure 2 Fluorophore structures and properties.
a, b, Skeletal structures of the modified Cy3B* donor (a) and Cy7* acceptor (b) fluorophores. c, d, Normalized donor fluorophore emission (Cy3: green; Cy3B*: dark green) and acceptor fluorophore absorbance (Cy5: red; Cy7*: dark magenta) spectra for Cy3/Cy5 (c) and Cy3B*/Cy7* (d) FRET pairs. The spectral overlap integral (shaded region) was calculated and used to determine the Förster distance (R0) values for each pair. e, Inter-dye FRET efficiencies of the Cy3/Cy5 (black) and Cy3B*/Cy7* (blue) donor and acceptor fluorophore pairs as a function of inter-dye distances calculated based on R0 values. f, Bulk anisotropy measurements on Cy3B*-labelled β2∆6-148C/266C.
Extended Data Figure 3 smFRET experimental controls.
a, Site-specific labelling. SDS–PAGE gels under green (540 nm) or near infrared (740 nm) illumination for fluorescence visualization of Cy3B* or Cy7* labelling of β2∆6 and β2∆6-148C/266C. Coomassie-stained gel image is shown as a gel-loading control. Digestion with factor Xa protease and deglycosylation with PNGase F leads to separation of the 148C and 266C labelling sites on generated N-terminal and C-terminal fragments, respectively. For gel source data, see Supplementary Fig. 1. b, Quantification of Cy3B*/Cy7*-labelling specificity of full-length β2∆6-148C/266C. Data are normalized to β2∆6-148C/266C labelling. c, Specificity of streptavidin-mediated receptor immobilization. Frame capture from immobilization movies showing labelled β2∆6-148C/266C on streptavidin-free (−SA) or streptavidin-coated (+SA) surfaces. Bar graph shows the number of immobilized, labelled β2AR in these conditions. Error bars represent the s.d. from two replicates. d, Schematic of labelled β2AR immobilization via biotinylated alprenolol (alp-biotin). e, FRET population contour plot and histogram for alp-biotin-immobilized receptor shows correspondence with the FRET population distribution of biotin-M1-Fab-immobilized, alprenolol-bound, labelled β2AR (Fig. 2b). Histogram error bars represent the s.d. from four replicates, with n total molecules analysed. f, FRET population contour plots (top) and histograms (bottom) for adrenaline titration on biotin-M1-Fab-immobilized, labelled β2∆6-148C/266C (Fig. 2c). Dashed lines (blue) highlight the mean FRET values for the lowest (2 nM; top dashed line) and highest (200 μM; bottom dashed line) adrenaline concentrations tested. Histogram error bars represent the s.d. from three replicates with n total molecules analysed. The scale bar on the right indicates relative populations for the contour plots.
Extended Data Figure 4 TM6 motions within ligand-bound β2AR.
a, Sample fluorescence (green for Cy3B*; magenta for Cy7*) and FRET (blue) time traces for biotin-M1-Fab-immobilized, labelled β2Δ6-148C/266C imaged in the absence and presence of saturating ligands. b, Same as Fig. 2b but with the full FRET efficiency range (0–1) shown for the population histograms (bottom). c, Plots showing the mean cross correlation values of donor and acceptor fluorescence as a function of lag time for the ensemble of individual fluorescence traces obtained from experiments shown in Fig. 2b. d, Plots showing the mean cross correlation values of donor and acceptor fluorescence as a function of lag time for an ensemble of simulated fluorescence trajectories rapidly fluctuating between high (0.75) and intermediate (0.55) FRET values with varying low- to high-FRET transition rate constants (coloured lines), where the high- to low-FRET transition rate is held constant at 100 s−1 (Supplementary Methods). e, FRET distributions of the simulated data, as described in d.
Extended Data Figure 5 All-atom molecular dynamics simulations of the Cy3B*/Cy7*-labelled β2AR in a detergent micelle.
a, Time evolution of the distance between the dyes. Time dependence of the distances between the midpoints of the dyes along the simulation trajectories is shown for the β2AR–carazolol (grey) and β2AR–BI-167107/Gs (black) systems. The distributions are displayed as histograms on the right; grey bars: β2AR–carazolol; clear bars: β2AR–BI-167107/Gs. The estimated inter-dye distances derived from the experimental mean FRET values (Figs 2b, 3b; Extended Data Fig. 2e) are indicated by solid lines topped with circles (β2AR–carazolol: red; β2AR–BI-167107/Gs: blue). b, Time evolution of the distance between Cα carbons at the labelling site. Cα–Cα distances for β2AR–carazolol: red and β2AR–BI-167107/Gs: blue. c, The simulated dye-tethered β2AR–BI-167107/Gs system embedded in a n-dodecyl-β-d-maltoside (DDM) micelle (grey sticks). β2AR is rendered in grey, with TM6 and the agonist (BI-167107) highlighted in blue. The Gs protein is rendered in wheat colour surrounded by its molecular surface to indicate the excluded volume for dye movements. The Cy3B* and Cy7* dyes are coloured green and magenta, respectively. Water molecules, ions and detergent molecules distant from the β2AR structure are omitted. d, Positions explored by the midpoints of the dyes during the simulations are shown as clusters of dots in the context of the β2AR–carazolol (transparent red dots) and β2AR–BI-167107/Gs (transparent blue dots) complexes; the centre of mass of each collection of dots is indicated by a solid sphere. Cα carbons for labelled positions 1484.40 and 2666.28, are shown as magenta and green spheres, respectively.
Extended Data Figure 6 smFRET imaging of biotin-Gs-immobilized, labelled β2AR.
a, Schematic of labelled β2AR immobilization via biotinylated Gs heterotrimer. b, c, Representative fluorescence (green for Cy3B*; magenta for Cy7*) and FRET (blue) time traces for biotin-Gs-immobilized, labelled β2Δ6-148C/266C imaged in the presence of clenbuterol (b) or adrenaline (c) in nucleotide-free conditions (apyrase-treated). d, FRET population contour plots (top) and histograms (bottom) for biotin-Gs-immobilized β2Δ6-148C/266C imaged in the presence of partial and full agonists in nucleotide-free conditions. The dashed line indicates the invariant mean FRET value (~0.38) for all agonists tested. Histogram error bars represent the s.d. from three replicates, with n total molecules analysed. e, f, FRET population contour plots of biotin-Gs-immobilized β2Δ6-148C/266C in the presence of agonists exhibiting FRET transitions upon rapid addition (arrow) of 30 μM GDP (e) or 100 μM GTP (f). Scale bar on the right indicates the relative population for the contour plots.
Extended Data Figure 7 Ligand-dependent TM6 dynamics in the presence of Gs, GDP and GTP.
a, Sample fluorescence and FRET time traces for biotin-M1-Fab-immobilized, labelled β2Δ6-148C/266C imaged in the absence and presence of saturating ligands plus 8 μM Gs in nucleotide-free (after apyrase treatment) conditions. b, c, FRET population contour plots for biotin-M1-Fab-immobilized, labelled β2∆6-148C/266C imaged in the presence of the agonists clenbuterol (b) or adrenaline (c), 8 μM Gs and increasing concentrations of GDP (Fig. 5a) or GTP in the presence of saturating GDP (30 μM) (Fig. 5e), with n indicating the total number of molecules analysed from two replicates. Nucleotide-free (0 μM GDP; from Fig. 3b) and ligand-only (from Fig. 2b) conditions are included as references. Scale bar on the right indicates the relative population for the contour plots.
Extended Data Figure 8 Steady-state dynamics within β2AR related to nucleotide exchange.
a, b, Transition rates from high- to low-FRET states (khigh→low) of labelled β2AR with different agonists, saturating Gs (8 μM) and increasing GDP concentrations (2–100 μM) (a) or increasing GTP concentrations (0–100 μM) in 30 μM GDP (b). c, Low-FRET state distributions for labelled β2AR with different agonists in 100 μM GDP and saturating Gs showing their overlap with the distribution in the presence of adrenaline and saturating Gs in nucleotide-free conditions (apyrase) (Fig. 3b). The dashed line shows the mean FRET value for apyrase. d, Mean low FRET values from c. e, The percentage change in the area of the low-FRET state population distributions for clenbuterol (black squares) and adrenaline (dark yellow triangles) (as shown in Fig. 5b and c, respectively) was plotted with increasing GDP concentrations (0–100 μM) and fitted to a single exponential decay function to derive the GDP EC50 value. f, g, Sample FRET traces (blue line) of labelled β2AR in the presence of clenbuterol (f) or adrenaline (g) plus 100 μM GDP and saturating Gs. Dashed lines indicate each ligand’s corresponding mean high FRET value (black), mean low FRET value (light green) and mean FRET value in nucleotide-free conditions (dark green). h, The ratio of the low-FRET state lifetime of β2AR in the presence of saturating Gs and GDP (τGDP) over the low-FRET state lifetime in saturating GTP plus 30 μM GDP (τGDP + GTP) (Fig. 5f) is shown for different agonists. All error bars represent s.d. from two replicates.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, additional references and the uncropped gels. (PDF 2031 kb)
Rights and permissions
About this article
Cite this article
Gregorio, G., Masureel, M., Hilger, D. et al. Single-molecule analysis of ligand efficacy in β2AR–G-protein activation. Nature 547, 68–73 (2017). https://doi.org/10.1038/nature22354
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22354
This article is cited by
-
Ligand efficacy modulates conformational dynamics of the µ-opioid receptor
Nature (2024)
-
Computational drug development for membrane protein targets
Nature Biotechnology (2024)
-
Time-resolved cryo-EM of G-protein activation by a GPCR
Nature (2024)
-
Stepwise activation of a metabotropic glutamate receptor
Nature (2024)
-
Binding kinetics drive G protein subtype selectivity at the β1-adrenergic receptor
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.