Abstract
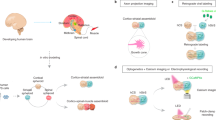
The development of the nervous system involves a coordinated succession of events including the migration of GABAergic (γ-aminobutyric-acid-releasing) neurons from ventral to dorsal forebrain and their integration into cortical circuits. However, these interregional interactions have not yet been modelled with human cells. Here we generate three-dimensional spheroids from human pluripotent stem cells that resemble either the dorsal or ventral forebrain and contain cortical glutamatergic or GABAergic neurons. These subdomain-specific forebrain spheroids can be assembled in vitro to recapitulate the saltatory migration of interneurons observed in the fetal forebrain. Using this system, we find that in Timothy syndrome—a neurodevelopmental disorder that is caused by mutations in the CaV1.2 calcium channel—interneurons display abnormal migratory saltations. We also show that after migration, interneurons functionally integrate with glutamatergic neurons to form a microphysiological system. We anticipate that this approach will be useful for studying neural development and disease, and for deriving spheroids that resemble other brain regions to assemble circuits in vitro.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kepecs, A. & Fishell, G. Interneuron cell types are fit to function. Nature 505, 318–326 (2014)
Wonders, C. P. & Anderson, S. A. The origin and specification of cortical interneurons. Nat. Rev. Neurosci. 7, 687–696 (2006)
Silbereis, J. C., Pochareddy, S., Zhu, Y., Li, M. & Sestan, N. The cellular and molecular landscapes of the developing human central nervous system. Neuron 89, 248–268 (2016)
Bartolini, G., Ciceri, G. & Marín, O. Integration of GABAergic interneurons into cortical cell assemblies: lessons from embryos and adults. Neuron 79, 849–864 (2013)
Rubenstein, J. L. & Merzenich, M. M. Model of autism: increased ratio of excitation/inhibition in key neural systems. Genes Brain Behav. 2, 255–267 (2003)
Marín, O. Interneuron dysfunction in psychiatric disorders. Nat. Rev. Neurosci. 13, 107–120 (2012)
Tabar, V. & Studer, L. Pluripotent stem cells in regenerative medicine: challenges and recent progress. Nat. Rev. Genet. 15, 82–92 (2014)
Pas¸ca, S. P., Panagiotakos, G. & Dolmetsch, R. E. Generating human neurons in vitro and using them to understand neuropsychiatric disease. Annu. Rev. Neurosci. 37, 479–501 (2014)
Maroof, A. M. et al. Directed differentiation and functional maturation of cortical interneurons from human embryonic stem cells. Cell Stem Cell 12, 559–572 (2013)
Nicholas, C. R. et al. Functional maturation of hPSC-derived forebrain interneurons requires an extended timeline and mimics human neural development. Cell Stem Cell 12, 573–586 (2013)
Pas¸ca, A. M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015)
Uylings, H. B. M., Delalle, I., Petanjek, Z. & Koenderink, M. J. T. Structural and immunocytochemical differentiation of neurons in prenatal and postnatal human prefrontal cortex. Neuroembryology 1, 176–186 (2002)
Fan, H. C., Fu, G. K. & Fodor, S. P. Expression profiling. Combinatorial labeling of single cells for gene expression cytometry. Science 347, 1258367 (2015)
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008)
Potter, G. B. et al. Generation of Cre-transgenic mice using Dlx1/Dlx2 enhancers and their characterization in GABAergic interneurons. Mol. Cell. Neurosci. 40, 167–186 (2009)
Sun, Y. et al. A deleterious Nav1.1 mutation selectively impairs telencephalic inhibitory neurons derived from Dravet syndrome patients. eLife 5, e13073 (2016)
Bortone, D. & Polleux, F. KCC2 expression promotes the termination of cortical interneuron migration in a voltage-sensitive calcium-dependent manner. Neuron 62, 53–71 (2009)
Nadarajah, B., Alifragis, P., Wong, R. O. & Parnavelas, J. G. Ventricle-directed migration in the developing cerebral cortex. Nat. Neurosci. 5, 218–224 (2002)
Anderson, S. A., Marín, O., Horn, C., Jennings, K. & Rubenstein, J. L. Distinct cortical migrations from the medial and lateral ganglionic eminences. Development 128, 353–363 (2001)
Bellion, A., Baudoin, J. P., Alvarez, C., Bornens, M. & Métin, C. Nucleokinesis in tangentially migrating neurons comprises two alternating phases: forward migration of the Golgi/centrosome associated with centrosome splitting and myosin contraction at the rear. J. Neurosci. 25, 5691–5699 (2005)
Stumm, R. K. et al. CXCR4 regulates interneuron migration in the developing neocortex. J. Neurosci. 23, 5123–5130 (2003)
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013)
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014)
Splawski, I. et al. CaV1.2 calcium channel dysfunction causes a multisystem disorder including arrhythmia and autism. Cell 119, 19–31 (2004)
Splawski, I. et al. Severe arrhythmia disorder caused by cardiac L-type calcium channel mutations. Proc. Natl Acad. Sci. USA 102, 8089–8096 (2005)
Pas¸ca, S. P. et al. Using iPSC-derived neurons to uncover cellular phenotypes associated with Timothy syndrome. Nat. Med. 17, 1657–1662 (2011)
Yarotskyy, V. et al. Roscovitine binds to novel L-channel (CaV1.2) sites that separately affect activation and inactivation. J. Biol. Chem. 285, 43–53 (2010)
Yarotskyy, V. & Elmslie, K. S. Roscovitine, a cyclin-dependent kinase inhibitor, affects several gating mechanisms to inhibit cardiac L-type (Ca(V)1.2) calcium channels. Br. J. Pharmacol. 152, 386–395 (2007)
Zechel, S., Zajac, P., Lönnerberg, P., Ibáñez, C. F. & Linnarsson, S. Topographical transcriptome mapping of the mouse medial ganglionic eminence by spatially resolved RNA-seq. Genome Biol. 15, 486 (2014)
Batista-Brito, R., Machold, R., Klein, C. & Fishell, G. Gene expression in cortical interneuron precursors is prescient of their mature function. Cereb. Cortex 18, 2306–2317 (2008)
Mao, L., Takamiya, K., Thomas, G., Lin, D. T. & Huganir, R. L. GRIP1 and 2 regulate activity-dependent AMPA receptor recycling via exocyst complex interactions. Proc. Natl Acad. Sci. USA 107, 19038–19043 (2010)
Mardinly, A. R. et al. Sensory experience regulates cortical inhibition by inducing IGF1 in VIP neurons. Nature 531, 371–375 (2016)
Lacaria, M., Gu, W. & Lupski, J. R. Circadian abnormalities in mouse models of Smith–Magenis syndrome: evidence for involvement of RAI1. Am. J. Med. Genet. A 161, 1561–1568 (2013)
Blake, D. J. et al. TCF4, schizophrenia, and Pitt–Hopkins syndrome. Schizophr. Bull. 36, 443–447 (2010)
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013)
Mariani, J. et al. Modeling human cortical development in vitro using induced pluripotent stem cells. Proc. Natl Acad. Sci. USA 109, 12770–12775 (2012)
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl Acad. Sci. USA 110, 20284–20289 (2013)
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016)
Krey, J. F. et al. Timothy syndrome is associated with activity-dependent dendritic retraction in rodent and human neurons. Nat. Neurosci. 16, 201–209 (2013)
Tian, Y. et al. Alteration in basal and depolarization induced transcriptional network in iPSC derived neurons from Timothy syndrome. Genome Med. 6, 75 (2014)
Yazawa, M. et al. Using induced pluripotent stem cells to investigate cardiac phenotypes in Timothy syndrome. Nature 471, 230–234 (2011)
Gallego Romero, I. et al. A panel of induced pluripotent stem cells from chimpanzees: a resource for comparative functional genomics. eLife 4, e07103 (2015)
Birey, F ., Andersen, J. & Pas¸ca, S. P. Generation and assembly of forebrain spheroids from human pluripotent stem cells. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.044 (2017)
Deverman, B. E. et al. Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain. Nat. Biotechnol. 34, 204–209 (2016)
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016)
Venkataramanappa, S., Simon, R. & Britsch, S. Ex utero electroporation and organotypic slice culture of mouse hippocampal tissue. J. Vis. Exp. (97), e52550 (2015)
Matsuda, T. & Cepko, C. L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl Acad. Sci. USA 101, 16–22 (2004)
Lui, J. H. et al. Radial glia require PDGFD–PDGFRβ signalling in human but not mouse neocortex. Nature 515, 264–268 (2014)
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014)
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013)
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013)
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015)
Micheva, K. D., Busse, B., Weiler, N. C., O’Rourke, N. & Smith, S. J. Single-synapse analysis of a diverse synapse population: proteomic imaging methods and markers. Neuron 68, 639–653 (2010)
Jang, H. J. et al. The development of phasic and tonic inhibition in the rat visual cortex. Korean J. Physiol. Pharmacol. 14, 399–405 (2010)
Acknowledgements
We thank J. Y. Park, K. Sabatini, T. Khan, S. Yoon, H. Gai and L. Elahi at Stanford University, R.E. Dolmetsch at Novartis Institutes for Biomedical Research, and N. Bansal and J. Fan at BD Genomics for advice and support. This work was supported by grants from NIH/National Institute of Mental Health (NIMH) grants R01MH100900 and R01MH100900-02S1, NIMH BRAINS Award R01MH107800, the California Institute of Regenerative Medicine (CIRM), the MQ Fellow Award, the Donald E. and Delia B. Baxter Foundation Faculty Award, the Kwan Research Fund and Stanford Start-up Funds (to S.P.P.), Child Research Health Institute Postdoctoral Fellowship (CHRI) (to F.B., N.H.), Walter V. and Idun Berry Postdoctoral Fellowship (to J.A.) and the Stanford Medicine Dean’s Fellowship (to F.B., J.A., N.H.), the American Epilepsy Society and Wishes for Elliott Foundation Fellowship (to C.D.M.), NIH 5P01HG00020526 (to L.M.S.), the UCSF Program for Breakthrough Biomedical Research and Sandler Foundation (to G.P.).
Author information
Authors and Affiliations
Contributions
F.B. and J.A. developed the differentiation platform and assays. C.D.M. and J.R.H. designed, conducted and analysed the electrophysiological experiments. F.B., J.A., N.H. and N.T. contributed to neural differentiation, live cell imaging experiments and analysis. F.B. and J.A. were able to replicate the differentiation and spheroid assembly independently. S.I., W.W., H.C.F., K.R.C.M. and L.M.S. contributed to the single-cell experiments. N.A.O. performed the array tomography. J.A.B. and J.H. recruited and characterized the subjects. G.P., J.A. and F.B. performed or analysed the electroporation experiments. F.B, J.A. and S.P.P. conceived the project, designed experiments and wrote the paper with input from all authors. S.P.P. supervised all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
Stanford University has filed a provisional patent application that covers the generation of region-specific neural spheroids and their assembly for studying human development and disease (US patent application number 62/477,858). H.C.F. and K.R.C.M. were employees of BD Genomics during this study.
Additional information
Reviewer Information Nature thanks G. Fishell, A. Goffinet and B. Treutlein for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Immunostaining of hSS in cryosections showing parvalbumin neurons.
Two anti-parvalbumin antibodies have been used to validate specificity to parvalbumin (PV); co-localization is shown with the neuronal marker DCX (day 209).
Extended Data Figure 2 Single-cell gene expression of hCS and hSS at day 105 of differentiation.
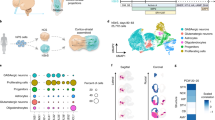
Single-cell gene expression analysis using the BD Resolve system (n = 11,838 cells). a–c, Distribution of expression of the neuronal marker STMN2 (a), the progenitor marker VIM (b), and of a set of genes (c), which are associated with the M cell cycle phase (AURKB, TPX2, UBE2C, HMMR, TOP2A, CCNB1, NUSAP1, NUF2, CDC6, HIST1H4C, BIRC5 and CKS2). d, Box plots for genes enriched in each tSNE cluster shown in Fig. 1j. e–l, Top 25 genes in each of the eight clusters shown in Fig. 1j (proportion of molecules per cells). m, Scatter plot showing the number of genes detected versus the number of useful reads.
Extended Data Figure 3 Characterization of hSS differentiation conditions.
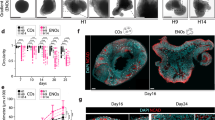
a, Schematic illustrating the differentiation conditions for deriving hCS or hSS: IS (IWP-2 and SAG), ISA (IWP-2, SAG and allopregnanolone (AlloP)) and ISRA (IWP-2, SAG, allopregnanolone (AlloP) and retinoic acid (RA)). Medium for all three conditions contained dorsomorphin (DM) and SB-431542 (SB), FGF2, EGF, BDNF and NT3 at the indicated time points. b, Representative traces of intracellular calcium measurements (Fluo-4) demonstrating spontaneous activity in hSS at around day 50 of differentiation. ΔF/F indicates the fluorescence intensity over baseline fluorescence. c, Average calcium spike frequency in three distinct hSS differentiation conditions: IS (n = 114 cells), ISA (n = 327 cells), ISRA (n = 136 cells); cells were derived from three hiPS cell lines; one-way ANOVA, F3,719 = 5.86, ***P < 0.001. d, Gene expression (qPCR, normalized to GAPDH) levels, showing downregulation of OCT4 and the lack of mesoderm (brachyury, BRACH) and endoderm (SOX17) markers following differentiation of hiPS cells into hCS and hSS conditions. e, Gene expression (qPCR, fold change versus hiPS cells and normalized to GAPDH) levels, showing upregulation of forebrain markers (SIX3 and FOXG1) but not midbrain (LMX1B), hypothalamus (RAX) or spinal cord (HOXB4) markers. f, Expression of ventral forebrain genes in hSS and hCS (qPCR, normalized to GAPDH) at day 25. g, Average percentage of the proportion of NKX2-1 by immunostaining in dissociated hCS or hSS at day 25. h, Expression of ventral forebrain genes in hSS (qPCR, normalized to GAPDH) at day 60. i, Expression of glutamatergic, GABAergic, dopaminergic and cholinergic neurotransmitter identity genes in hSS (qPCR, normalized to GAPDH) at day 60. j, Average percentage of the proportion of MAP2/Hoechst and GAD67/MAP2 by immunostaining in dissociated hSS at day 60. k, l, Expression of interneuron subtype genes and markers associated with interneuron migration in hSS (qPCR, normalized to GAPDH) at day 60. Number of hPS lines used indicated on each column. Data are mean ± s.e.m.
Extended Data Figure 4 Electrophysiological recordings of hCS and hSS.
a, Representative EPSC traces of recordings from hCS neurons (sliced preparation) before (black) and during (green) exposure to the glutamate receptor blocker kynurenic acid (Kyn, 1 mM) (Mann–Whitney U-test, **P = 0.007). b, Overlap of averaged EPSCs (red) recorded in hCS neurons (n = 6 cells) and averaged IPSCs (black) recorded in hSS (n = 5 cells). Data are mean ± s.d.
Extended Data Figure 5 Migration of Dlxi1/2::eGFP+ cells in fused hSS–hCS.
a, b, Representative immunostaining in cryosections of hSS showing co-expression of Dlxi1/2::eGFP and GAD67 (a) or GABA (b). c, Quantification by immunostaining of the proportion of Dlxi1/2/2::eGFP+ cells that co-express GAD67 in hSS derived using the ISA or ISRA condition (n = 3 hiPS cell lines; t-test, P = 0.35). d, Proportion of Dlxi1/2::eGFP+ and GAD67+ cells in hSS derived using the ISA or ISRA condition that co-express somatostatin (SST) (t-test, P = 0.48), calretinin (CR) (t-test, *P = 0.04) or calbindin (CB) (t-test, P = 0.43); n = 3 hiPS cell lines. e, Representative images of hSS–hCS at day 60 showing migration of Dlxi1/2::eGFP+ cells (from fluorescently labelled hSS) in fused hSS–hCS, but not in hSS–hSS over time. f, The number of Dlxi1/2::eGFP+ (hSS-derived) or hSYN1::mCherry+ cells (hCS-derived) that moved in hSS–hCS or hSS–hSS was quantified in snapshots of live, intact spheroids at different time points (from day 3 to 25). The values shown are the absolute number of cells that migrated to the other side (approximately the same area and thickness was imaged in each session); one-way ANOVA for cells at 17 days after assembly; F2,32 = 8.24, P = 0.001. g, Representative images of fused hSS–hCS at day 91 showing migration of Dlxi1/2::eGFP+ cells (from fluorescently labelled hSS) into fused hSS–hCS. h, Representative image of an hSS that was plated on a glass coverslip and cultured for around 7 days. i, Percentage of Dlxi1/2::eGFP inside the coverslip-plated hSS, in the rim (0–200 μm) or beyond this region (>200 μm). j, Quantification of the number of saltations of Dlxi1/2::eGFP+ cells (n = 32 cells) inside the one-week coverslip-plated hSS, in the rim and outside this region (one-way ANOVA, interaction F2,30 = 22.12, P < 0.001; Bonferroni post hoc test, ***P < 0.0001). k, Diagram showing the angle of movement of migrating Dlxi1/2::eGFP+ cells at 8–9 days after assembly of hSS–hCS. The angle was calculated between the leading process of Dlxi1/2::eGFP+ cells that have moved into hCS and the fusion interface (n = 92 cells from five hiPS cell lines). l, Histogram showing the distribution of the distance of migrated Dlxi1/2::eGFP+ cells relative to the edge of hCS in hSS–hCS at 30–50 days after assembly. The distance was measured in fixed 18-μm cryosections (n = 73 cells from two hiPS cell lines). m–o, Representative examples of migrated Dlxi1/2::eGFP+ cells in the hCS that moved within a ventricular zone-like region. The ventricular zone-like region contains GFAP-expressing cells, is surrounded by TBR1+ cells and the migrated cells show GABA expression. Supplementary Video 2 shows movement of Dlxi1/2::eGFP+ cells that is reminiscent of the ventricular-directed migration described in rodents.
Extended Data Figure 6 Single-cell gene expression of Dlxi1/2::eGFP+ cells in hSS and hCS.
a, Schematic showing the isolation by dissociation and fluorescence-activated cell sorting (FACS) of Dlxi1/2::eGFP+ cells from hSS or hCS for single-cell transcriptional analysis. b, Violin plots showing expression in Dlxi1/2::eGFP+ cells of selected genes associated with cortical, striatal and olfactory interneurons in hSS (light green, n = 123 cells) or hCS (dark green, n = 106 cells) at two weeks after assembly of hSS–hCS. c, Violin plots showing expression in Dlxi1/2::eGFP+ cells (at four weeks after assembly of hSS–hCS) in clusters A, B, and C (likelihood ratio test; GAD1, CELF4: P > 0.05; PBX3: P < 1 × 10−7 for A versus B and C; NNAT: P < 1 × 10−16 for C versus A and B, P < 1 × 10−16 for B versus A and C; MALAT1: P < 1 × 10−9 for C versus A and B; SOX11: P < 1 × 10−16 for B versus A and C, P < 1 × 10−9 for A versus B and C; GRIP2: P < 1 × 10−8 for B versus A and C). d, Scatter plot showing the number of genes detected (≥10 reads cut-off) versus the number of reads (n = 410 cells from combined single-cell RNA-sequencing experiments after two or four weeks of assembly in hSS–hCS). e, Graph illustrating biologically variable transcripts (red circles) and non-variable transcripts (black circles) along with the technical noise from the ERCC spike in RNAs (blue dots). Green line shows the technical noise fit.
Extended Data Figure 7 Immunocytochemistry and pharmacology in fused hSS–hCS.
a–d, Representative images of immunostainings for somatostatin (SST), GAD67, GABA, calretinin (CR) and calbindin (CB) in Dlxi1/2::eGFP+ cells after migration in fused hSS–hCS. e, Schematic illustrating the pharmacological manipulation of Dlxi1/2::eGFP+ cells that are migrating in hSS–hCS. f–i, Quantification of Dlxi1/2::eGFP+ cell migration before and after exposure to 100 nM of the CXCR4 antagonist AMD3100 (n = 8 cells from two hiPS cell lines; paired t-tests, *P = 0.03 for number of saltations (f), **P = 0.006 for saltation length (g), **P = 0.006 for speed when mobile (h), *P = 0.02 for path directness(i)). j, Plot illustrating the trajectory of Dlxi1/2::eGFP+ cells in fused hSS–hCS before and after exposure to AMD3100. Data are mean ± s.e.m.
Extended Data Figure 8 Migration of Dlxi1/2::eGFP+ cells in mouse and human forebrain slices versus hSS–hCS.
a–c, Representative images of human fetal cortex at GW20 showing immunostaining with antibodies against GFAP (a), CTIP2 (b) and GABA (c). d, Representative image showing cell labelling with the Dlxi1/2::eGFP reporter in human forebrain at GW18 (six days after lentivirus infection). e, f, Representative images of immunostained cryosections of human tissue at GW18 showing co-localization of Dlxi1/2::eGFP with NKX2-1 (e) and GABA (f). g, Representative images showing cell labelling with the Dlxi1/2::eGFP reporter in hSS–hCS (nine days after assembly), in human forebrain (GW18) and in mouse slice cultures (E18). h, i, Comparison of Dlxi1/2::eGFP+ cell size and quantification of the ratio of soma diameter to the length of the leading process in fused hSS–hCS (n = 25 cells from four hiPS cell lines), human forebrain at GW18 (n = 19 cells, black) and GW20 (n = 15 cells, grey), hSS-derived cells cultured on E14 mouse forebrain slices (n = 14 cells), and E18 mouse forebrain slices (n = 30 cells from two litters) (one-way ANOVA, interaction F3,97 = 11.61, P = 0.001; Bonferroni post hoc test *P < 0.05, ***P < 0.001). j–l, Comparison of the number of saltations (n = 56 cells from two hiPS cell lines; one-way ANOVA, interaction F2,103 = 29.27, P = 0.001, Bonferroni post hoc test ***P < 0.001), saltation length (n = 44 cells from three hiPS cell lines; one-way ANOVA, interaction F2,91 = 3.0, P = 0.50), speed when mobile (n = 38 cells from three hiPS cell lines; one-way ANOVA, interaction F2,83 = 11.38, P = 0.001, Bonferroni post hoc test ***P < 0.001) for Dlxi1/2::eGFP+ in fused hSS–hCS, human fetal forebrain (GW18, n = 19 cells; GW20, n = 15 cells), and E18 mouse forebrain slices (n = 14 cells for saltation length and speed, n = 16 cells for number of saltations from two litters). Data are mean ± s.e.m.
Extended Data Figure 9 Derivation of TS hSS, migration and electroporation.
a, Sequencing of PCR-amplified DNA showing the p.G406R mutation in exon 8a of CACNA1C in TS (subject 8303). b, Representative images of hiPS cell colonies expressing pluripotency markers (OCT4, SSEA4) in one TS subject. c, Level of gene expression (qPCR, normalized to GAPDH) for NKX2-1 showing no defects in ventral forebrain induction in TS (two-way ANOVA; interaction F2,15 = 0.20, P = 0.81; TS versus control F1,15 = 0.16, P = 0.68). n.s., not significant. d–g, Representative images of immunostained cryosections of TS hSS (day 60) showing expression of NKX2-1, GABA, MAP2, GAD67, somatostatin (SST) and calretinin (CR). h, Calcium imaging (Fura-2) in dissociated hCS derived from TS subjects and controls (control, n = 81 cells from two subjects; TS, n = 147 cells from two subjects). Quantification of residual intracellular calcium ([Ca2+]i) following 67 mM KCl depolarization of control and TS cells in hCS cells. Residual [Ca2+]i was calculated by dividing the plateau calcium (C − A) level by the peak calcium elevation (B − A) (t-test, ***P < 0.001). i, Quantification of [Ca2+]i following depolarization of control and TS cells in hSS (t-test, ***P < 0.001). j, Representative image of fused TS hSS–hCS showing Dlxi1/2::eGFP expression and migration. k, l, Quantification of the number of saltations and saltation length of Dlxi1/2::eGFP cells in fused hSS–hCS across multiple control and TS lines (related to Fig. 3d, e). m, Quantification of the speed when mobile of Dlxi1/2::eGFP cells in fused hSS–hCS (control, n = 21 cells from three hiPS cell lines derived from three subjects; TS, n = 29 cells from three hiPS cell lines derived from three subjects; TS–control hybrid, n = 12 cells from three combinations of five hiPS cell lines from five subjects; one-way ANOVA with Dunnett’s multiple comparison test, ***P < 0.001). n, Electroporation of cDNA encoding the TS and wild-type (WT) CaV1.2 channels into slices of mouse E14 ganglionic eminences (GE). o, Representative example of time-lapse live cell imaging depicting the saltatory migration of GFP+ cells in slices electroporated with CAG::GFP and either wild-type or TS CACNA1C. p, q, Quantification of the number of saltations (p, t-test, **P < 0.01) and saltation length (q, t-test, ***P < 0.001) of GFP+ cells in electroporated mouse forebrain slices (wild type, n = 33 cells; TS, n = 23 cells; from three litters). r, Schematic illustrating pharmacological manipulation of LTCCs during live cell imaging of fused hSS–hCS. s, Quantification of speed when mobile following exposure to the LTCC blocker nimodipine (5 μM) (paired t-test; control, n = 13 cells from three hiPS cell lines derived from three subjects, ***P < 0.001; TS, n = 12 cells from two hiPS cell lines derived from two subjects, **P < 0.005). t, Quantification of saltation length following exposure to roscovitine (15 μM) (paired t-test; control, n = 7 cells from two hiPS cell lines derived from two subjects, **P < 0.005; TS, n = 12 cells from two hiPS cell lines derived from two subjects; ***P < 0.001). u, Quantification of speed when mobile following exposure to roscovitine (15 μM) (paired t-tests; control, n = 9 cells from two hiPS cell lines derived from two subjects, ***P < 0.001; TS, n = 12 cells from two hiPS cell lines derived from two subjects; P = 0.05). Data are mean ± s.e.m.
Extended Data Figure 10 Characterization of Dlxi1/2::eGFP+ cells after migration.
a, Representative images of three-dimensional reconstructed Dlxi1/2::eGFP+ cell morphologies before and after migration from hSS into hCS. b, Quantification of dendritic branching of Dlxi1/2b::eGFP+ cells in hSS (n = 58 cells) and in hCS (n = 55 cells) of fused hSS–hCS (two-way ANOVA; interaction F2,129 = 11.29, P < 0.001; Bonferroni post hoc test *P < 0.05, ***P < 0.001). c, Representative examples of action potentials (slice recordings) in Dlxi1/2::eGFP+ cells in unfused hSS, in hSS of fused hSS–hCS and in hCS after migration in fused hSS–hCS. d, Array tomography showing expression of the GABAergic synapse marker GPHN (green) colocalized with SYN1 (red) in hCS of fused hSS–hCS but not in unfused hCS; the glutamatergic marker PSD95 (cyan) colocalized with SYN1 is found in both fused and unfused hCS (equal volumes 1.2-μm deep). e, Array tomography of a Dlxi1/2::eGFP+ synapse illustrating the colocalization with SYN1 (red), GPHN (cyan) and VGAT (white); five consecutive 70-nm sections (3 × 3 μm). f, Representative examples of whole-cell voltage-clamp recordings of IPSCs and EPSCs from Dlxi1/2::eGFP+ cells in unfused hSS, in fused hSS–hCS or after migration in hCS. (g) Representative examples of whole-cell voltage-clamp recordings of IPSCs and EPSCs in cells recorded from unfused hCS cells and fused hCS cells. h, Electrical stimulation and patch-clamp recording in fused hSS–hCS showing evoked EPSCs and IPSCs before (black) and after exposure to 10 μM gabazine (red). i, Average peri-stimulus synaptic events (IPSCs and EPSCs) in Dlxi1/2::eGFP+ cells recorded in the hCS side of fused hSS–hCS before and after electrical stimulation (paired t-test, *P < 0.05). Data are mean ± s.e.m.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5. (PDF 462 kb)
Live imaging (14.2 hrs) showing migration of Dlx2i1/2b::eGFP+ cells in fused hSS-hCS (day 12 after fusion).
hCS is located in the upper side part of the fused spheroids. Yellow arrowheads point to cells undergoing saltatory migration steps. (MOV 9697 kb)
Live imaging showing migration of Dlx2i1/2b::eGFP+ cells in a hSS plated on a coverslip (example #1: 22.7 hrs; example #1: 6.7 hrs).
Arrows indicate representative Dlx2i1/2b::eGFP+ cells. (MOV 1664 kb)
Live imaging (15.2 hrs) showing ventricle directed migration of Dlx2i1/2b::eGFP+ cells in fused hSS-hCS
A VZ-like region in the hCS has been identified by bright field microscopy and indicated with two concentrical circles. (MOV 1282 kb)
Live imaging (18.2 hrs; low magnification) showing migration of Dlx2i1/2b::eGFP+ cells in slices of human fetal forebrain at GW20
Human fetal slices were labeled with the reporter 5–6 days before imaging. (MOV 12487 kb)
Live imaging (18.9 hrs) showing examples of migrating Dlx2i1/2b::eGFP+ cells in slices of human fetal forebrain at GW18 versus fused hSS-hCS
Human fetal slices were labeled with the reporter 5–6 days before imaging. (MOV 3241 kb)
Live imaging showing migration of Dlx2i1/2b::eGFP+ cells in fused hSS-hCS before (9.6 hrs) and during exposure to the CXCR4 receptor antagonist AMD3100 (24.4 hrs total imaging time)
Live imaging showing migration of Dlx2i1/2b::eGFP+ cells in fused hSS-hCS before (9.6 hrs) and during exposure to the CXCR4 receptor antagonist AMD3100 (24.4 hrs total imaging time). (MOV 5379 kb)
Live imaging (4.6 hrs) showing migration of Dlx2i1/2b::eGFP+ cells in fused hSS-hCS derived from TS subject (lower panel) and a control subject (upper panel)
Live imaging (4.6 hrs) showing migration of Dlx2i1/2b::eGFP+ cells in fused hSS-hCS derived from TS subject (lower panel) and a control subject (upper panel). (MOV 1780 kb)
Rights and permissions
About this article
Cite this article
Birey, F., Andersen, J., Makinson, C. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017). https://doi.org/10.1038/nature22330
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22330
This article is cited by
-
Modelling the complex nature of the tumor microenvironment: 3D tumor spheroids as an evolving tool
Journal of Biomedical Science (2024)
-
iPSC-derived models of PACS1 syndrome reveal transcriptional and functional deficits in neuron activity
Nature Communications (2024)
-
Complex activity and short-term plasticity of human cerebral organoids reciprocally connected with axons
Nature Communications (2024)
-
A patient-specific lung cancer assembloid model with heterogeneous tumor microenvironments
Nature Communications (2024)
-
A patterned human neural tube model using microfluidic gradients
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.