Abstract
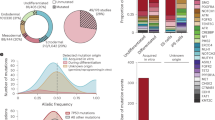
Human pluripotent stem cells (hPS cells) can self-renew indefinitely, making them an attractive source for regenerative therapies. This expansion potential has been linked with the acquisition of large copy number variants that provide mutated cells with a growth advantage in culture1,2,3. The nature, extent and functional effects of other acquired genome sequence mutations in cultured hPS cells are not known. Here we sequence the protein-coding genes (exomes) of 140 independent human embryonic stem cell (hES cell) lines, including 26 lines prepared for potential clinical use4. We then apply computational strategies for identifying mutations present in a subset of cells in each hES cell line5. Although such mosaic mutations were generally rare, we identified five unrelated hES cell lines that carried six mutations in the TP53 gene that encodes the tumour suppressor P53. The TP53 mutations we observed are dominant negative and are the mutations most commonly seen in human cancers. We found that the TP53 mutant allelic fraction increased with passage number under standard culture conditions, suggesting that the P53 mutations confer selective advantage. We then mined published RNA sequencing data from 117 hPS cell lines, and observed another nine TP53 mutations, all resulting in coding changes in the DNA-binding domain of P53. In three lines, the allelic fraction exceeded 50%, suggesting additional selective advantage resulting from the loss of heterozygosity at the TP53 locus. As the acquisition and expansion of cancer-associated mutations in hPS cells may go unnoticed during most applications, we suggest that careful genetic characterization of hPS cells and their differentiated derivatives be carried out before clinical use.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
04 May 2017
New references 29 and 30 were added, and subsequent citations were renumbered accordingly.
References
The International Stem Cell Initiative. ISCI. Screening ethnically diverse human embryonic stem cells identifies a chromosome 20 minimal amplicon conferring growth advantage. Nat. Biotechnol . 29, 1132–1144 (2011)
Avery, S. et al. BCL-XL mediates the strong selective advantage of a 20q11.21 amplification commonly found in human embryonic stem cell cultures. Stem Cell Rep. 1, 379–386 (2013)
Nguyen, H. T. et al. Gain of 20q11.21 in human embryonic stem cells improves cell survival by increased expression of Bcl-xL. Mol. Hum. Reprod. 20, 168–177 (2014)
Unger, C., Skottman, H., Blomberg, P., Dilber, M. S. & Hovatta, O. Good manufacturing practice and clinical-grade human embryonic stem cell lines. Hum. Mol. Genet. 17, R48–R53 (2008)
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014)
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015)
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014)
Adewumi, O. et al. Characterization of human embryonic stem cell lines by the International Stem Cell Initiative. Nat. Biotechnol. 25, 803–816 (2007)
Baker, D. et al. Detecting genetic mosaicism in cultures of human pluripotent stem cells. Stem Cell Rep . 7, 998–1012 (2016)
Schwartz, S. D. et al. Human embryonic stem cell-derived retinal pigment epithelium in patients with age-related macular degeneration and Stargardt’s macular dystrophy: follow-up of two open-label phase 1/2 studies. Lancet 385, 509–516 (2015)
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016)
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res . 43, D805–D811 (2015)
Zhang, J . et al. International Cancer Genome Consortium data portal—a one-stop shop for cancer genomics data. Database 2011, bar026 (2011)
Bouaoun, L. et al. TP53 variations in human cancers: new lessons from the IARC TP53 database and genomics data. Hum. Mutat. 37, 865–876 (2016)
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000)
Rideout, W. M. III, Coetzee, G. A., Olumi, A. F. & Jones, P. A. 5-methylcytosine as an endogenous mutagen in the human LDL receptor and p53 genes. Science 249, 1288–1290 (1990)
Cho, Y., Gorina, S., Jeffrey, P. D. & Pavletich, N. P. Crystal structure of a p53 tumor suppressor-DNA complex: understanding tumorigenic mutations. Science 265, 346–355 (1994)
Willis, A., Jung, E. J., Wakefield, T. & Chen, X. Mutant p53 exerts a dominant negative effect by preventing wild-type p53 from binding to the promoter of its target genes. Oncogene 23, 2330–2338 (2004)
Malkin, D. Li–Fraumeni syndrome. Genes Cancer 2, 475–484 (2011)
Xu, J. et al. Heterogeneity of Li–Fraumeni syndrome links to unequal gain-of-function effects of p53 mutations. Sci. Rep. 4, 4223 (2014)
Hindson, B. J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83, 8604–8610 (2011)
Marión, R. M. et al. A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature 460, 1149–1153 (2009)
Zhao, Y. et al. Two supporting factors greatly improve the efficiency of human iPSC generation. Cell Stem Cell 3, 475–479 (2008)
Amir, H. et al. Spontaneous single-copy loss of TP53 in human embryonic stem cells markedly increases cell proliferation and survival. Stem Cells (2016)
Forster, R. et al. Human intestinal tissue with adult stem cell properties derived from pluripotent stem cells. Stem Cell Rep . 2, 838–852 (2014)
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011)
Xie, R. et al. Dynamic chromatin remodeling mediated by polycomb proteins orchestrates pancreatic differentiation of human embryonic stem cells. Cell Stem Cell 12, 224–237 (2013)
Garber, K. RIKEN suspends first clinical trial involving induced pluripotent stem cells. Nat. Biotechnol. 33, 890–891 (2015)
Laurent, L. C. Dynamic changes in the copy number of pluripotency and cell proliferation genes in human ESCs and iPSCs during reprogramming and time in culture. Cell Stem Cell 8, 106–118 (2011)
Merkle, F. T. & Eggan, K. Culturing human pluripotent stem cells from diverse culture histories. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.087 (2017)
Ludwig, T. E. et al. Derivation of human embryonic stem cells in defined conditions. Nat. Biotechnol. 24, 185–187 (2006)
Chen, G. et al. Chemically defined conditions for human iPSC derivation and culture. Nat. Methods 8, 424–429 (2011)
Merkle, F. T. et al. Efficient CRISPR–Cas9-mediated generation of knockin human pluripotent stem cells lacking undesired mutations at the targeted locus. Cell Reports 11, 875–883 (2015)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011)
Jun, G. et al. Detecting and estimating contamination of human DNA samples in sequencing and array-based genotype data. Am. J. Hum. Genet. 91, 839–848 (2012)
Wang, K. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007)
Ganna, A. et al. Ultra-rare disruptive and damaging mutations influence educational attainment in the general population. 19, 1563–1565 (2016)
Case, D. A. et al. AMBER 2016
Wheeler, D. A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008)
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013)
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res . 29, 308–311 (2001)
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011)
Kent, W. J., Zweig, A. S., Barber, G., Hinrichs, A. S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. 26, 2204–2207 (2010)
Acknowledgements
We thank the many institutions and investigators world-wide that provided their cell lines and supported the publication of the results. We are indebted to D. Santos, M. Smith, K. Elwell, M. A. Yram, S. Ellender, L. Bevilacqua, and D. Gage for their assistance with the regulatory and logistical efforts required to acquire and sequence hES cell lines. We also thank K. Lilliehook for her comments, I. Yildirim for his assistance with the molecular modelling of P53 mutations, and C. Usher for help with figure schematics. We regret the omission of any relevant references or discussion due to space limitations. The Genomics Platform at the Broad Institute performed sample preparation, sequencing, and data storage. Y.A. is a Clore Fellow. N.B. is the Herbert Cohn Chair in Cancer Research and was partially supported by The Rosetrees Trust and The Azrieli Foundation. Costs associated with acquiring and sequencing hES cell lines were supported by HHMI and the Stanley Center for Psychiatric Research. F.T.M., S.A.M., and K.E. were supported by grants from the NIH (5P01GM099117, 5K99NS08371). K.E. was supported by the Miller consortium of the HSCI, and F.T.M. is currently supported by funds from the Wellcome Trust, the Medical Research Council (MR/P501967/1), and the Academy of Medical Sciences (SBF001\1016).
Author information
Authors and Affiliations
Contributions
F.T.M., S.G., S.A.M., and K.E. conceived the project. F.T.M. and K.E. acquired hES cell lines with the assistance of M.C. and G.S., who also assisted with regulatory issues pertaining to sequencing and data distribution. F.T.M. cultured and banked hES cell lines, prepared them for sequencing, and coordinated efforts to interpret and visualize sequencing data with the assistance of S.G. S.G. performed computational data analysis and visualization with the help of G.G., R.E.H., and S.K. Y.A. preformed the analysis of TP53 mutations in the RNA-seq database with the assistance of S.B. and N.B. Data were interpreted by F.T.M., S.G., N.K., G.G., Y.A., S.B., N.B., S.A.M., and K.E. N.K., J.M., and C.M. designed, performed and analysed experiments to measure the mosaic nature and competitive expansion of TP53 mutations. S.M. derived HUES 68, 69, 70, 74, 75, and D.I. provided the KCL lines. F.T.M., S.G., S.A.M., and K.E. prepared drafts of the manuscript text and figures with contributions and comments from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Replicates of cell competition assays carried out at earlier starting passages.
Note that while the mutant allelic fractions for lines CHB11 and WA26 approach fixation, that the fraction of mutant cells unexpectedly decreases for ESI035 over several passages, indicating a potential selective disadvantage that co-segregates with the TP53 mutation in this experiment. The number of replicate wells is indicated in each graph. Values depict the mean and error bars depict s.e.m.
Extended Data Figure 2 Summary of all observed P53 mutations.
a, b, Graphical representation of each of the 9 mutated bases in P53 observed across the 252 whole-exome sequenced (WES) and RNA-seq hPS cell lines depicting their allele frequency in ExAC (a) and the incidence with which the relevant codons are mutated in human cancer (b). c, The 15 instances of these mutations in 12 distinct cell lines and the method used to detect them are pictured. Although the M237I event is seen in two distinct hiPS cell lines, it is conservatively counted as a single event as the two affected clones may be derived from a common reprogrammed progenitor.
Extended Data Figure 3 Analysis of loss of heterozygosity in RNA sequencing samples.
a, Polymorphic sites on chromosome 17 in different hPS cells with mutations in TP53. WIBR3 cells with H193R mutation and H9 cells with both P151S and R248Q mutations show less polymorphism in the distal part of chromosome 17p compared to the proximal part of 17p and 17q. Asterisk indicates samples with less than 25 reads. b, Ratio between the fraction of polymorphic alleles in the distal part of chromosome 17p or the reminder of chromosome 17 (proximal 17p + 17q) compared to that fraction for the entire chromosome 17. Values shown depict mean. Where present, error bars depict s.e.m. for 2–22 replicate samples. ***P < 0.001, one-sided Z-score test for the two population proportion. WIBR3 cells with H193R mutation and H9 cells with both P151S and R248Q mutations have a significantly different proportion between the two parts of the chromosome, implying LOH. c, A schematic representation of possible allele states of TP53 in cultured hPS cells with all observed mutations depicted. Depending on the percentage of mutant reads in a culture, one can deduce if the culture is homogenous or mosaic for a mutation, and whether, in addition to a point mutation, LOH has occurred in the TP53 locus. MAF, minor allele frequency.
Extended Data Figure 4 Culture and passaging method employed for samples bearing P53 mutations.
a, P53 mutations were observed in hPS cells grown in a broad array of culture media including media supplemented with knockout serum replacement (KOSR), and defined, commercial media such as E8. b, P53 mutations were observed from cells grown with feeder cells or under feeder-free conditions. c, As passaging hPS cells can introduce stresses or clonal bottlenecks, we examined whether P53 mutations were consistently seen when a particular passaging method was used and observed a wide variety of passaging methods associated with these mutations. Note that the interpretation of these data are complicated by the fact that the culture methods employed in the final published study may not reflect the previous culture history of that cell line, which may have previously passed through multiple laboratories, as well as by the lack of detail about culture methods present in some published studies. d, P53 mutations are seen in studies that either include or exclude supplements such as the rock inhibitor Y-27632 (10 μM) at the time of passaging.
Supplementary information
Supplementary Table 1
Considered and whole exome sequenced hESC lines. Tab 1. We considered hESC lines for WES if they were listed on the NIH Human Embryonic Stem Cell Registry (http://grants.nih.gov/stem_cells/registry/current.htm) or if they were prepared under GMP conditions. Cell lines were typically excluded from consideration if they were unavailable for distribution or contained known karyotypic abnormalities in more than 10% of analyzed cells or disease-causing mutations identified by PGD. Cell lines with MTAs that restricted our ability to work with the cell lines, that could not be recovered upon thawing, or proved to be unavailable upon request were also excluded. Passage number at the time of request, the number of passages and time in culture from thaw to passaging, and the passaging method, media, and substrate, are provided, as is mean sequencing coverage and % cross sample contaminated per cell line. GMP, good manufacturing practice; MTA, material transfer agreement; PGD, pre-implantation genetic diagnosis; WES, whole exome sequencing. Tab 2. Summary of number of cells considered and sequenced, including reasons for exclusion. These data are presented graphically in Figure 1b-e. (XLSX 57 kb)
Supplementary Table 2
Identification of candidate mosaic variants present in sequenced hESCs. Tab 1. Filters used to identify likely mosaic variants among all heterozygous variants present among the sequenced exomes of 140 hESCs. Tab 2. List of 263 candidate mosaic variants passing quality control filters and present no more than two times among the 140 sequenced hESC lines. Variants are arranged by chromosome position and annotated by likely functional impact and frequency in the general population (ExAC AC). Tab 3. Variants from the list in Tab 2 predicted to have either a high or damaging impact on gene function based on a consensus of 7 bioinformatic algorithms. See Materials and Methods for further details. Tab 4. In addition to mosaic variants identified using these stringent filters, we provide an inclusive list of all high confidence somatic variants (n=36,396) that pass the binomial test with a P value of <0.01. SNP, single nucleotide polymorphism; CHROM, chromosome number; POS, genomic position (hg19); REF, reference allele; ALT, alternate allele; HESC, human embryonic stem cell line; REFC, count of reference alleles; ALTC, count of alternate alleles; FILTER, high confidence variant score; EXACAC, allele count in the Exome Aggregation Consortium (ExAC) database; IMPACT, predicted effect of mutation; HESCAC, allele count in hESCs; TOTALC, REFC+ALTC; AF, allelic faction (ALTC/TOTALC); P, P value for binomial test on allelic fraction. (XLSX 4082 kb)
Supplementary Table 3
Characteristics of TP53 mutations identified in hESCs by WES and RNAseq. Tab 1. Summary of all 15 instances of TP53 mutations observed by WES and RNAseq with details of read depth, allelic fraction, P value, reference, and culture method. Note that all observed mutations are frequently seen in human cancer, and that most mutations have evidence of mosaicism, indicating that they were likely culture-derived. bFGF, basic fibroblast growth factor (FGF2); COSMIC, Catalogue of Somatic Mutations in Cancer (http://cancer.sanger.ac.uk/cosmic); ExAC, Exome Aggregation Consortium (http://exac.broadinstitute.org/); Freq., frequency; GMP, good manufacturing practice, IARC, International Agency for Research on Cancer (http://p53.iarc.fr/); ICGC, International Cancer Genome Consortium (http://icgc.org/); MEF, mouse embryonic fibroblast; Seq., sequencing; SNL, SNL mouse fibroblast feeder cell line; WES, whole exome sequencing. Errors denote SEM. Tab 2. Breakdown of the incidence of P53 mutations by culture media, substrate, and passaging method. (XLSX 51 kb)
Supplementary Table 4
Primer and probe sequences used for ddPCR-based determination of P53 variant allele frequency. (XLSX 30 kb)
Supplementary Table 5
Calculation of selective advantage conferred by three distinct TP53 variants. The allelic fraction of TP53 variants was measured at several passages by ddPCR in hESCs cultured under standard conditions. Replicate experiments per passage are shown in grey, and average values are shown in black. The observed increase in allelic frequency of each of the variants across time in culture is consistent with a substantial growth or survival advantage in all but one instance. See Materials and Methods for details on ddPCR and the calculation of the effect per passage. (XLSX 39 kb)
Supplementary Table 6
Large copy number variants in hESCs identified by the human Psych Array. Tab 1. Summary of hESC lines with large copy number variants (>500kb) as ascertained by SNP array analysis. Two of the five cell lines with acquired TP53 mutations harbored large structural alternations (HUES71 and MShef10). Five separate cell lines (CSES25, ESI051, MShef3, UM78-1 and WA21) had an amplification at the pericentromeric region of chromosome 20 (Chr20q11.21). Tab 2. Complete list of large deletions or duplications (>500kb) identified across the 140 hESC lines. (XLSX 57 kb)
Supplementary Table 7
Identification of TP53 mutations in hPSCs by RNA sequencing and WES. Tab 1. List of all RNA sequenced samples from hPSCs. Five of these samples (cell2-7) were removed since they were from single stem cells rather than cell lines. Tab 2. Summary of the number of samples and studies generated from each cell line. Tab 3. List of all samples harboring TP53 mutations, their chromosomal location, and the relevant study. Tab 4. Summary of all affected cell lines and studies. Tab 5. Summary of affected samples, cell lines, and number of mutations seen in hESCs and hiPSCs by WES and RNAseq. (XLSX 68 kb)
Rights and permissions
About this article
Cite this article
Merkle, F., Ghosh, S., Kamitaki, N. et al. Human pluripotent stem cells recurrently acquire and expand dominant negative P53 mutations. Nature 545, 229–233 (2017). https://doi.org/10.1038/nature22312
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22312
This article is cited by
-
Chimeric antigen receptor-based natural killer cell immunotherapy in cancer: from bench to bedside
Cell Death & Disease (2024)
-
High prevalence of acquired cancer-related mutations in 146 human pluripotent stem cell lines and their differentiated derivatives
Nature Biotechnology (2024)
-
Unlocking the Potential of Extracellular Vesicles as the Next Generation Therapy: Challenges and Opportunities
Tissue Engineering and Regenerative Medicine (2024)
-
The origins of human pluripotent stem cells: the road from a cancer to regenerative medicine
In Vitro Cellular & Developmental Biology - Animal (2024)
-
Epigenetic repression of CHCHD2 enhances survival from single cell dissociation through attenuated Rho A kinase activity
Cellular and Molecular Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.