Abstract
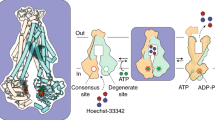
ATP binding cassette (ABC) transporters of the exporter class harness the energy of ATP hydrolysis in the nucleotide-binding domains (NBDs) to power the energetically uphill efflux of substrates by a dedicated transmembrane domain (TMD)1,2,3,4. Although numerous investigations have described the mechanism of ATP hydrolysis and defined the architecture of ABC exporters, a detailed structural dynamic understanding of the transduction of ATP energy to the work of substrate translocation remains elusive. Here we used double electron–electron resonance5,6 and molecular dynamics simulations to describe the ATP- and substrate-coupled conformational cycle of the mouse ABC efflux transporter P-glycoprotein (Pgp; also known as ABCB1), which has a central role in the clearance of xenobiotics and in cancer resistance to chemotherapy7. Pairs of spin labels were introduced at residues selected to track the putative inward-facing to outward-facing transition. Our findings illuminate how ATP energy is harnessed in the NBDs in a two-stroke cycle and elucidate the consequent conformational motion that reconfigures the TMD, two critical aspects of Pgp transport mechanism. Along with a fully atomistic model of the outward-facing conformation in membranes, the insight into Pgp conformational dynamics harmonizes mechanistic and structural data into a novel perspective on ATP-coupled transport and reveals mechanistic divergence within the efflux class of ABC transporters.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Higgins, C. F. & Linton, K. J. The ATP switch model for ABC transporters. Nature Struct. Mol. Biol. 11, 918–926 (2004)
Rees, D. C., Johnson, E. & Lewinson, O. ABC transporters: the power to change. Nature Rev. Mol. Cell Biol. 10, 218–227 (2009)
Jones, P. M. & George, A. M. Mechanism of the ABC transporter ATPase domains: catalytic models and the biochemical and biophysical record. Crit. Rev. Biochem. Mol. Biol. 48, 39–50 (2013)
Sharom, F. J. ABC multidrug transporters: structure, function and role in chemoresistance. Pharmacogenomics 9, 105–127 (2008)
Mchaourab, H. S., Steed, P. R. & Kazmier, K. Toward the fourth dimension of membrane protein structure: insight into dynamics from spin-labeling EPR spectroscopy. Structure 19, 1549–1561 (2011)
Jeschke, G. DEER distance measurements on proteins. Annu. Rev. Phys. Chem. 63, 419–446 (2012)
Gottesman, M. M., Fojo, T. & Bates, S. E. Multidrug resistance in cancer: role of ATP-dependent transporters. Nature Rev. Cancer 2, 48–58 (2002)
Sharom, F. J., Yu, X., Chu, J. W. K. & Doige, C. A. Characterization of the ATPase activity of P-glycoprotein from multidrug-resistant Chinese hamster ovary cells. Biochem. J. 308, 381–390 (1995)
Urbatsch, I. L., Sankaran, B., Weber, J. & Senior, A. E. P-glycoprotein is stably inhibited by vanadate-induced trapping of nucleotide at a single catalytic site. J. Biol. Chem. 270, 19383–19390 (1995)
Kerr, K. M., Sauna, Z. E. & Ambudkar, S. V. Correlation between steady-state ATP hydrolysis and vanadate-induced ADP trapping in human P-glycoprotein. Evidence for ADP release as the rate-limiting step in the catalytic cycle and its modulation by substrates. J. Biol. Chem. 276, 8657–8664 (2001)
Ward, A., Reyes, C. L., Yu, J., Roth, C. B. & Chang, G. Flexibility in the ABC transporter MsbA: alternating access with a twist. Proc. Natl Acad. Sci. USA 104, 19005–19010 (2007)
Zou, P., Bortolus, M. & Mchaourab, H. S. Conformational cycle of the ABC transporter MsbA in liposomes: detailed analysis using double electron-electron resonance spectroscopy. J. Mol. Biol. 393, 586–597 (2009)
Aller, S. G. et al. Structure of P-glycoprotein reveals a molecular basis for poly-specific drug binding. Science 323, 1718–1722 (2009)
Li, J., Jaimes, K. F. & Aller, S. G. Refined structures of mouse P-glycoprotein. Protein Sci. 23, 34–46 (2014)
Jin, M. S., Oldham, M. L., Zhang, Q. & Chen, J. Crystal structure of the multidrug transporter P-glycoprotein from Caenorhabditis elegans . Nature 490, 566–569 (2012)
Verhalen, B. & Wilkens, S. P-glycoprotein retains drug-stimulated ATPase activity upon covalent linkage of the two nucleotide binding domains at their C-terminal ends. J. Biol. Chem. 286, 10476–10482 (2011)
Holland, I. B. & Blight, M. A. ABC-ATPases, adaptable energy generators fuelling transmembrane movement of a variety of molecules in organisms from bacteria to humans. J. Mol. Biol. 293, 381–399 (1999)
Dawson, R. J. & Locher, K. P. Structure of a bacterial multidrug ABC transporter. Nature 443, 180–185 (2006)
Ambudkar, S. V., Kim, I. W., Xia, D. & Sauna, Z. E. The A-loop, a novel conserved aromatic acid subdomain upstream of the Walker A motif in ABC transporters, is critical for ATP binding. FEBS Lett. 580, 1049–1055 (2006)
Carrier, I., Julien, M. & Gros, P. Analysis of catalytic carboxylate mutants E552Q and E1197Q suggests asymmetric ATP hydrolysis by the two nucleotide-binding domains of P-glycoprotein. Biochemistry 42, 12875–12885 (2003)
Sauna, Z. E., Müller, M., Peng, X.-H. & Ambudkar, S. V. Importance of the conserved Walker B glutamate residues, 556 and 1201, for the completion of the catalytic cycle of ATP hydrolysis by human P-glycoprotein (ABCB1). Biochemistry 41, 13989–14000 (2002)
Tombline, G. et al. Properties of P-glycoprotein with mutations in the “catalytic carboxylate” glutamate residues. J. Biol. Chem. 279, 46518–46526 (2004)
Tombline, G., Bartholomew, L. A., Urbatsch, I. L. & Senior, A. E. Combined mutation of catalytic glutamate residues in the two nucleotide binding domains of P-glycoprotein generates a conformation that binds ATP and ADP tightly. J. Biol. Chem. 279, 31212–31220 (2004)
Tombline, G. & Senior, A. E. The occluded nucleotide conformation of P-glycoprotein. J. Bioenerg. Biomembr. 37, 497–500 (2005)
Senior, A. E., al-Shawi, M. K. & Urbatsch, I. L. The catalytic cycle of P-glycoprotein. FEBS Lett. 377, 285–289 (1995)
Siarheyeva, A., Liu, R. & Sharom, F. J. Characterization of an asymmetric occluded state of P-glycoprotein with two bound nucleotides: implications for catalysis. J. Biol. Chem. 285, 7575–7586 (2010)
Moradi, M. & Tajkhorshid, E. Mechanistic picture for conformational transition of a membrane transporter at atomic resolution. Proc. Natl Acad. Sci. USA 110, 18916–18921 (2013)
Wen, P. C., Verhalen, B., Wilkens, S., Mchaourab, H. S. & Tajkhorshid, E. On the origin of large flexibility of P-glycoprotein in the inward-facing state. J. Biol. Chem. 288, 19211–19220 (2013)
Mishra, S. et al. Conformational dynamics of the nucleotide binding domains and the power stroke of a heterodimeric ABC transporter. eLife 3, e02740 (2014)
Tombline, G. et al. Expression, purification, and characterization of cysteine-free mouse P-glycoprotein. Arch. Biochem. Biophys. 445, 124–128 (2006)
Smriti, Z., Zou, P. & Mchaourab, H. S. Mapping daunorubicin-binding sites in the ATP-binding cassette transporter MsbA using site-specific quenching by spin labels. J. Biol. Chem. 284, 13904–13913 (2009)
Jeschke, G. & Polyhach, Y. Distance measurements on spin-labelled biomacromolecules by pulsed electron paramagnetic resonance. Phys. Chem. Chem. Phys. 9, 1895–1910 (2007)
Polyhach, Y., Bordignon, E. & Jeschke, G. Rotamer libraries of spin labelled cysteines for protein studies. Phys. Chem. Chem. Phys. 13, 2356–2366 (2011)
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. Chapter 2, Unit 2.9 (2007)
Simossis, V. A. & Heringa, J. PRALINE: a multiple sequence alignment toolbox that integrates homology-extended and secondary structure information. Nucleic Acids Res. 33, W289–W294 (2005)
Pirovano, W., Feenstra, K. A. & Heringa, J. PRALINETM: a strategy for improved multiple alignment of transmembrane proteins. Bioinformatics 24, 492–497 (2008)
Zaitseva, J., Jenewein, S., Jumpertz, T., Holland, I. B. & Schmitt, L. H662 is the linchpin of ATP hydrolysis in the nucleotide-binding domain of the ABC transporter HlyB. EMBO J. 24, 1901–1910 (2005)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005)
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013)
Feller, S. E. & MacKerell, A. D. Jr. An improved empirical potential energy function for molecular simulations of phospholipids. J. Phys. Chem. B 104, 7510–7515 (2000)
Foloppe, N. & MacKerell, A. D. Jr. All-atom empirical foce field for nucleic acids: I. parameter optimization based on small molecule and condensed phase macromolecular target data. J. Comput. Chem. 21, 86–104 (2000)
Jo, S., Kim, T. & Im, W. Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS ONE 2, e880 (2007)
Jo, S., Lim, J. B., Klauda, J. B. & Im, W. CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys. J. 97, 50–58 (2009)
Wu, E. L. et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014)
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008)
Brooks, B. R. et al. CHARMM: the biomolecular simulation program. J. Comput. Chem. 30, 1545–1614 (2009)
Acknowledgements
We thank R. Stein for advice on the global analysis of DEER data, I. L. Urbatsch for donation of Pichia pastoris expression construct, and S. Wilkens for providing three expressed mutants (615–1276, 627–1260 and 613–1258) in P. pastoris. This work was supported by National Institutes of Health grants U54-GM087519 (to H.S.M., R.K.N. and E.T.) and P41-GM104601 (to E.T.). We also acknowledge computing resources provided by Blue Waters at National Center for Supercomputing Applications, Department of Energy Innovative and Novel Computational Impact on Theory and Experiment, and Extreme Science and Engineering Discovery Environment (grant TG-MCA06N060 to E.T.).
Author information
Authors and Affiliations
Contributions
H.S.M., B.V. and R.K.N. designed the experimental studies. B.V. and R.D. purified the mutants and performed the experiments. R.D. and B.V. analysed the data. S.T. and E.T. designed and performed the molecular dynamics simulations. H.S.M., R.D., B.V., S.T. and E.T. wrote the paper. Y.P. cloned and expressed the mutants. H.A.K. and B.V. initially contributed in the optimization, cloning and expression of Pgp.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 ATPase activity.
a, Representative ATPase assays on Pgp mutants. Basal (black) and Ver-stimulated (blue) ATP hydrolysis were measured by Pi-release. The red dots represent the release rate under Vi-trapping condition (5 mM ATP, 1 mM Vi). b, c, The maximum activity (Vmax) values for mutants with intact NBSs (b) and with catalytic glutamate substitutions (c) are determined by fitting the data with simple Michaelis–Menten equation. Error bars are fitting errors or standard deviation (Cys-less Pgp).
Extended Data Figure 2 NBDs form a closed dimer in the Vi-trapped transition state.
From left to right, ribbon representation of NBDs viewed from the cytosolic side (PDB accession 4M1M) with residues mutated to Cys represented as purple sticks, and DEER distance distributions. Inward-facing and outward-facing (OF) distance distributions predicted from the X-ray structure of the inward-facing, and from the outward-facing model, are shown as dotted black and red traces, respectively. Residues 627 and 1276 were not resolved in the X-ray structure. The datasets on spin-labelled pairs 607–1252 and 613–1258 are repeated from Fig. 1b.
Extended Data Figure 3 Opening of the extracellular side of the TMD in the transition state.
a, b, Spin-labelled pairs designed to monitor distances between structurally nonequivalent helices (a) as well as equivalent helices (b) in the two leaflets are represented as purple spheres. Changes in the distance distributions are predominantly indicative of increased distances as predicted by the MsbA outward-facing (OF) structure and consistent with the Pgp outward-facing model. However, even for residues where distance changes are observed, the distributions are broad and components similar to those of the apo state are present, suggesting a heterogeneous transitions state. Inward-facing and outward-facing distance distributions predicted from the X-ray structure of the inward-facing and from the outward-facing model are shown as dotted black and red traces, respectively.
Extended Data Figure 4 Local asymmetry of the NBSs.
a, Positions of the mutated residues in each NBS are represented as purple sticks. b–e, Distance distributions monitoring each NBS in the wild-type (WT; b), E552Q (c), E1197Q (d) and E552Q/E1197Q (e) backgrounds. Spin-labelled pairs 400–1121 and 478–1043 have greatly reduced ATPase activity (see Extended Data Fig. 1). However, comparing the distributions of these spin-labelled pairs to those of 400–1156 and 511–1043, the pattern of changes in distance distributions is consistent, confirming the asymmetry is localized to the region of the A-loop. The data sets on spin-labelled pairs 400–1156 and 511–1043 are repeated from Fig. 3.
Extended Data Figure 5 Assembly of the NBSs upon Vi-trapping of the transition state.
a, Ribbon representation of NBSs viewed from the side with the mutated residues represented as purple sticks. b–e, Distance distributions monitoring each NBS in the wild-type (WT; b, repeated from Fig. 2b), E552Q (c), E1197Q (d) and E552Q/E1197Q (e) backgrounds. The pattern and amplitude of the distance changes at these locations are similar and consistent with the proximity of the Walker A and signature motifs in the outward-facing (OF) conformation as previously observed for MsbA and Sav1866. Introduction of the catalytic glutamate substitutions does not affect the distance changes. Moreover, these substitutions stabilize the transitions state so that it is populated under turnover conditions—that is, in the presence of ATP/Mg2+ and Ver (ATP-Ver, blue). These results demonstrate that the asymmetry identified in Extended Data Fig. 4 and its dependence on the catalytic activity of each NBS is localized to the A-loop region.
Extended Data Figure 6 The mutations of the catalytic glutamates do not alter the TMD conformational changes but stabilize the transition state.
a, Cytoplasmic (left three panels) and extracellular (right panel) TMD regions with spin-labelled residues represented as purple sticks. Distance distributions monitoring cytoplasmic and extracellular closing and opening in the wild-type (WT; b, repeated from Fig. 1c, d), E552Q (c), E1197Q (d) and E552Q/E1197Q (e) backgrounds. In the Vi-trapped transition state (ADP-Vi-Ver, red), the distance distributions are consistent with population of the outward-facing (OF) conformation. In the E-to-Q backgrounds, ATP/Mg2+ is sufficient to stabilize Pgp to outward-facing.
Extended Data Figure 7 Structural modelling of the unknown outward-facing state.
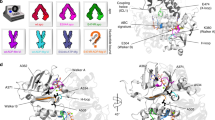
a, Structural alignment of all the TM helices between mouse Pgp and MsbA, which is used as a structural template in model construction. Regions highlighted with black dashed boxes indicate the missing structural segments from the template. TM helices of MsbA are coloured silver whereas TM helices 1–6 and 7–12 of Pgp are coloured yellow and cyan, respectively. The residues of TM helices and intracellular helices (IH) are labelled in colours corresponding to their structures. b, Incomplete model of Pgp outward-facing state resulted from the alignment without fixing the missing segments. Incomplete structural regions are highlighted in black dashed boxes. c, A complete model of Pgp outward-facing state after rebuilding the incomplete regions with structural information obtained from the inward-facing crystal structure of mouse Pgp (PDB accession 4M1M).
Extended Data Figure 8 Collective variables and structural features used to obtain the outward-facing state.
a–g, Description of the collective variables (CVs) used to obtain a reliable outward-facing state of Pgp (a–d) and tracking important structural features to verify the stability of the outward-facing state (e–g). a, Orientation quaternion (β) describing the angle between the two bundles of TM helices that separate to form the outward-facing state. Distance between K185 and D993, a charged residue pair located within the translocation chamber. b, CVs used to form accurate NBD-based interactions, which include NBD–ATP interactions and X-loop interactions. Walker A (WA1 and WA2), Walker B (WB1 and WB2) and LSGGQ (L1 and L2) motifs are shown in purple, yellow and green new cartoon representations, respectively. Y397/1040 from the A-loop (white) and ATPs (cyan) are shown in stick representations, whereas Mg2+ ions and Cα carbons of X-loop residues are displayed as grey and white beads, respectively. c, Metrics used in evaluating the basic structural elements that are key to any outward-facing ABC exporter, namely, dimerized NBDs (dNBD), closed cytoplasmic (α), and opened extracellular/periplasmic (β) sides. d, Sim1 (light colours) failed to maintain these basic structural requirements within 10 ns, whereas Sim2 (dark colours) results in a stable outward-facing state for up to 300 ns. Solid and dotted horizontal lines represent the corresponding values in inward-facing and outward-facing conformations, respectively, based on crystal structures of Pgp (PDB accession 4M1M) and MsbA (PDB accession 3B60). e, Description and time series of centre of mass distance between extended TM helical regions of TM3 (V164–V175) and TM10 (E887–E898) shown in orange, and TM4 (A244–A255) and TM9 (T806–D817) shown in pink, describing the tight closing of cytoplasmic side. f, Description and time series of centre of mass distance between the residues forming the top half of TMDs that open at the extracellular side (shown with pink and orange beads). g, Salt bridge interaction between K185 (TM3) and D993 (TM12). These calculations are compared between five different simulations.
Supplementary information
Supplementary Information
This file contains Supplementary Text and additional references. Experimental design and interpretation is expanded to include rationale, detailed analysis of DEER data, and the IF/OF alternating access model. The overarching mechanistic implications of the ABC transporter diversity is discussed in relation to literature and this work. (PDF 668 kb)
Dynamics of outward-facing (OF) state of Pgp in membrane.
Dynamics of OF state of Pgp within membrane is shown from both the front and side views. Also, the restrained and free (unbiased) parts of the simulations are labeled and shown in different color modes, with the first 30 ns (restrained) part in a dimmer and the following 270 ns of free simulation shown in a brighter representations. (MP4 14257 kb)
Rights and permissions
About this article
Cite this article
Verhalen, B., Dastvan, R., Thangapandian, S. et al. Energy transduction and alternating access of the mammalian ABC transporter P-glycoprotein. Nature 543, 738–741 (2017). https://doi.org/10.1038/nature21414
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21414
This article is cited by
-
Design, synthesis and bioactivity study on oxygen-heterocyclic-based pyran analogues as effective P-glycoprotein-mediated multidrug resistance in MCF-7/ADR cell
Scientific Reports (2024)
-
Orientational Selectivity in Pulsed-EPR Does Not Have to be Complicated
Applied Magnetic Resonance (2024)
-
Spectroscopically Orthogonal Labelling to Disentangle Site-Specific Nitroxide Label Distributions
Applied Magnetic Resonance (2024)
-
On the interplay between lipids and asymmetric dynamics of an NBS degenerate ABC transporter
Communications Biology (2023)
-
Asymmetric conformations and lipid interactions shape the ATP-coupled cycle of a heterodimeric ABC transporter
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.