Abstract
Aggregation of amyloid-β peptides into fibrils or other self-assembled states is central to the pathogenesis of Alzheimer’s disease. Fibrils formed in vitro by 40- and 42-residue amyloid-β peptides (Aβ40 and Aβ42) are polymorphic, with variations in molecular structure that depend on fibril growth conditions1,2,3,4,5,6,7,8,9,10,11,12. Recent experiments1,13,14,15,16 suggest that variations in amyloid-β fibril structure in vivo may correlate with variations in Alzheimer’s disease phenotype, in analogy to distinct prion strains that are associated with different clinical and pathological phenotypes17,18,19. Here we investigate correlations between structural variation and Alzheimer’s disease phenotype using solid-state nuclear magnetic resonance (ssNMR) measurements on Aβ40 and Aβ42 fibrils prepared by seeded growth from extracts of Alzheimer’s disease brain cortex. We compared two atypical Alzheimer’s disease clinical subtypes—the rapidly progressive form (r-AD) and the posterior cortical atrophy variant (PCA-AD)—with a typical prolonged-duration form (t-AD). On the basis of ssNMR data from 37 cortical tissue samples from 18 individuals, we find that a single Aβ40 fibril structure is most abundant in samples from patients with t-AD and PCA-AD, whereas Aβ40 fibrils from r-AD samples exhibit a significantly greater proportion of additional structures. Data for Aβ42 fibrils indicate structural heterogeneity in most samples from all patient categories, with at least two prevalent structures. These results demonstrate the existence of a specific predominant Aβ40 fibril structure in t-AD and PCA-AD, suggest that r-AD may relate to additional fibril structures and indicate that there is a qualitative difference between Aβ40 and Aβ42 aggregates in the brain tissue of patients with Alzheimer’s disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Petkova, A. T. et al. Self-propagating, molecular-level polymorphism in Alzheimer’s β-amyloid fibrils. Science 307, 262–265 (2005)
Paravastu, A. K., Leapman, R. D., Yau, W. M. & Tycko, R. Molecular structural basis for polymorphism in Alzheimer’s β-amyloid fibrils. Proc. Natl Acad. Sci. USA 105, 18349–18354 (2008)
Paravastu, A. K., Qahwash, I., Leapman, R. D., Meredith, S. C. & Tycko, R. Seeded growth of β-amyloid fibrils from Alzheimer’s brain-derived fibrils produces a distinct fibril structure. Proc. Natl Acad. Sci. USA 106, 7443–7448 (2009)
Lu, J. X. et al. Molecular structure of β-amyloid fibrils in Alzheimer’s disease brain tissue. Cell 154, 1257–1268 (2013)
Bertini, I., Gonnelli, L., Luchinat, C., Mao, J. & Nesi, A. A new structural model of Aβ40 fibrils. J. Am. Chem. Soc. 133, 16013–16022 (2011)
Schutz, A. K. et al. Atomic-resolution three-dimensional structure of amyloid β fibrils bearing the Osaka mutation. Angew. Chem. Int. Edn Engl. 54, 331–335 (2015)
Xiao, Y. et al. Aβ(1–42) fibril structure illuminates self-recognition and replication of amyloid in Alzheimer’s disease. Nat. Struct. Mol. Biol. 22, 499–505 (2015)
Sgourakis, N. G., Yau, W. M. & Qiang, W. Modeling an in-register, parallel “Iowa” aβ fibril structure using solid-state NMR data from labeled samples with Rosetta. Structure 23, 216–227 (2015)
Goldsbury, C., Frey, P., Olivieri, V., Aebi, U. & Müller, S. A. Multiple assembly pathways underlie amyloid-β fibril polymorphisms. J. Mol. Biol. 352, 282–298 (2005)
Meinhardt, J., Sachse, C., Hortschansky, P., Grigorieff, N. & Fändrich, M. Aβ(1-40) fibril polymorphism implies diverse interaction patterns in amyloid fibrils. J. Mol. Biol. 386, 869–877 (2009)
Zhang, R. et al. Interprotofilament interactions between Alzheimer’s Aβ1–42 peptides in amyloid fibrils revealed by cryoEM. Proc. Natl Acad. Sci. USA 106, 4653–4658 (2009)
Kodali, R., Williams, A. D., Chemuru, S. & Wetzel, R. Aβ(1–40) forms five distinct amyloid structures whose β-sheet contents and fibril stabilities are correlated. J. Mol. Biol. 401, 503–517 (2010)
Meyer-Luehmann, M. et al. Exogenous induction of cerebral β-amyloidogenesis is governed by agent and host. Science 313, 1781–1784 (2006)
Langer, F. et al. Soluble Aβ seeds are potent inducers of cerebral β-amyloid deposition. J. Neurosci. 31, 14488–14495 (2011)
Stöhr, J. et al. Distinct synthetic Aβ prion strains producing different amyloid deposits in bigenic mice. Proc. Natl Acad. Sci. USA 111, 10329–10334 (2014)
Cohen, M. L. et al. Rapidly progressive Alzheimer’s disease features distinct structures of amyloid-β. Brain 138, 1009–1022 (2015)
Bessen, R. A. & Marsh, R. F. Distinct PrP properties suggest the molecular basis of strain variation in transmissible mink encephalopathy. J. Virol. 68, 7859–7868 (1994)
Collinge, J., Sidle, K. C. L., Meads, J., Ironside, J. & Hill, A. F. Molecular analysis of prion strain variation and the aetiology of ‘new variant’ CJD. Nature 383, 685–690 (1996)
Safar, J. et al. Eight prion strains have PrPSc molecules with different conformations. Nat. Med. 4, 1157–1165 (1998)
Gath, J. et al. Unlike twins: an NMR comparison of two α-synuclein polymorphs featuring different toxicity. PLoS One 9, e90659 (2014)
van der Wel, P. C. A., Lewandowski, J. R. & Griffin, R. G. Solid-state NMR study of amyloid nanocrystals and fibrils formed by the peptide GNNQQNY from yeast prion protein Sup35p. J. Am. Chem. Soc. 129, 5117–5130 (2007)
Collinge, J. & Clarke, A. R. A general model of prion strains and their pathogenicity. Science 318, 930–936 (2007)
Tang-Wai, D. F. et al. Clinical, genetic, and neuropathologic characteristics of posterior cortical atrophy. Neurology 63, 1168–1174 (2004)
Schmidt, C. et al. Rapidly progressive Alzheimer’s disease: a multicenter update. J. Alzheimers Dis. 30, 751–756 (2012)
Henry, E. R. & Hofrichter, J. Singular value decomposition: application to analysis of experimental data. Methods Enzymol. 210, 129–192 (1992)
Qiang, W., Yau, W. M., Luo, Y., Mattson, M. P. & Tycko, R. Antiparallel β-sheet architecture in Iowa-mutant β-amyloid fibrils. Proc. Natl Acad. Sci. USA 109, 4443–4448 (2012)
Colvin, M. T. et al. Atomic resolution structure of monomorphic Aβ42 amyloid fibrils. J. Am. Chem. Soc. 138, 9663–9674 (2016)
Wälti, M. A. et al. Atomic-resolution structure of a disease-relevant Aβ(1-42) amyloid fibril. Proc. Natl Acad. Sci. USA 113, E4976–E4984 (2016)
Guo, J. L. et al. Distinct α-synuclein strains differentially promote tau inclusions in neurons. Cell 154, 103–117 (2013)
Sanders, D. W. et al. Distinct tau prion strains propagate in cells and mice and define different tauopathies. Neuron 82, 1271–1288 (2014)
Acknowledgements
This work was supported by the Intramural Research Program of the National Institute of Diabetes and Digestive and Kidney Diseases of the US National Institutes of Health, by the UK Medical Research Council and by the National Institute of Health Research (NIHR) UCLH/UCL Biomedical Research Centre. We are grateful for the assistance of S. Mead, O. Avwenagha, and J. Wadsworth at the MRC Prion Unit in selection and processing of tissue samples. We thank UK neurologists for referral of rapidly progressive dementias to the NHS National Prion Clinic, National Hospital for Neurology and Neurosurgery (NHNN), University College London Hospitals NHS Foundation Trust (UCLH). We thank the Queen Square Brain Bank for Neurological Disorders (supported by the Reta Lila Weston Trust for Medical Research, the Progressive Supranuclear Palsy [Europe] Association and the Medical Research Council) at the UCL Institute of Neurology, for provision of the human brain tissue samples. We thank all patients and their families for consent to use tissues in research.
Author information
Authors and Affiliations
Contributions
W.Q., J.-X.L., J.C., and R.T. designed experiments, including selection of tissue samples, development of protocols for preparation of brain-seeded fibrils, and selection of ssNMR measurements. W.Q., J.-X.L., and R.T. prepared fibril samples and acquired TEM images and ssNMR data. W.-M.Y. synthesized isotopically labelled peptides and performed ELISA measurements. W.Q. and R.T. analysed ssNMR data. J.C. and R.T. wrote the manuscript, with contributions from all other authors.
Corresponding authors
Extended data figures and tables
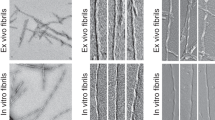
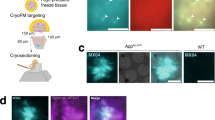
Extended Data Figure 1 Additional TEM images of brain-seeded fibrils.
a, TEM grids were prepared 4 h after the addition of solubilized Aβ40 or Aβ42 to sonicated brain extract and were negatively stained with uranyl acetate. Collagen fibrils in the extract (40–100 nm width, with characteristic transverse bands) appear in some images. Material with an amorphous appearance are non-fibrillar, non-Aβ components of the brain extract. Yellow arrows indicate Aβ40 fibrils with an apparent width modulation, attributable to an approximately periodic twisting of the fibril structure about the fibril growth direction. TEM images of all 37 brain-seeded Aβ40 and all 33 Aβ42 fibril samples are available at http://dx.doi.org/10.17632/whgp9r7tkd.1. b, Histogram of distances between width minima for Aβ40 fibrils with apparent width modulation. The Gaussian fit to this histogram (red curve) has a mean value of 107.2 nm (n = 65) and a full-width-at-half-maximum of 46.1 nm.
Extended Data Figure 2 2D ssNMR spectra of brain-seeded Aβ40 fibrils.
a, 2D 13C–13C spectra of fibrils seeded with extract from t-AD, PCA-AD, r-AD, or non-dementia (ND) tissue. Aliphatic regions are shown, with 15 contour levels (increasing by successive factors of 1.3, and with the highest contour at the maximum signal in each 2D spectrum). b, 2D 15N–13C spectra of fibrils seeded with extract from t-AD, PCA-AD, r-AD, or non-dementia tissue. Regions containing intra-residue 15N–13Cα cross-peaks are shown, with 11 contour levels (increasing by successive factors of 1.3, with the highest contour at the maximum signal in each spectrum). 15N–13Cβ cross-peaks from L34 appear in some spectra. Positions of cross-peaks from the predominant Aβ40 fibril structure are indicated by colour-coded circles (F19, blue; V24, cyan; G25, pink; S26, orange; A30, purple; I31, red; L34, green; M35, magenta). Only 2D spectra that were included in the analyses in Figs. 3 and 4 are shown. The full set of 42 2D 13C–13C spectra and 40 2D 15N–13C spectra, including those with lower signal-to-noise ratios, controls, and technical replicates, is available on-line at http://dx.doi.org/10.17632/tbp45pm92x.1.
Extended Data Figure 3 2D ssNMR spectra of brain-seeded Aβ42 fibrils.
a, 2D 13C–13C spectra of fibrils seeded with extract from t-AD, PCA-AD, r-AD, or non-dementia (ND) tissue. Aliphatic regions are shown, with 15 contour levels (increasing by successive factors of 1.3, and with the highest contour at the maximum signal in each spectrum). b, 2D 15N–13C spectra of fibrils seeded with extract from t-AD, PCA-AD, r-AD, or non-dementia tissue. Regions containing intra-residue 15N–13Cα cross-peaks are shown, with 11 contour levels (increasing by successive factors of 1.3, with the highest contour at the maximum signal in each spectrum). Only 2D spectra that were included in the analyses in Figs. 3 and 4 are shown. The full set of 33 2D 13C–13C spectra and 23 2D 15N–13C spectra, including those with lower signal-to-noise, controls, and technical replicates, is available at http://dx.doi.org/10.17632/tbp45pm92x.1.
Extended Data Figure 4 Control experiments using cortical tissue without Aβ deposits.
a, Comparison of TEM images of control tissue extract and r-AD2p′ extract after incubation for 4 h with solubilized Aβ40, under conditions identical to those that led to fibrils shown in Extended Data Fig. 1a. Fibrils associated with brain material were abundant on the TEM grid of the r-AD2p′-seeded sample, but were not observed in an extensive search over the TEM grid of the control sample. b, TEM images of control tissue extract and r-AD2p′ extract after incubation for 4 h with solubilized Aβ42, under conditions identical to those that led to fibrils shown in Extended Data Fig. 1b. Fibrils associated with brain material were abundant on the TEM grid of the r-AD2p′-seeded sample, but were not observed in an extensive search over the TEM grid of the control sample. c, 2D 13C–13C and 15N–13C spectra of Aβ40 fibrils (blue) and Aβ42 fibrils (red) that developed in control samples after 168 h or 48 h incubation, respectively, followed by 24 h intermittent sonication (see Supplementary Methods) and a further 72 h of additional incubation. Contour levels increase by successive factors of 1.4. d, r.m.s.d. values between 2D spectra of control fibrils and 2D spectra of AD brain-seeded fibrils, with dashed lines at values corresponding to white shades in Fig. 3. Occipital tissue of a female who died from cardiac arrest at age 86 was used as a control.
Extended Data Figure 5 Principal component analyses of 2D 13C–13C and 15N–13C ssNMR spectra of brain-seeded Aβ40 fibrils.
a, The first five principal components (PC1–PC5) of the 32 experimental 2D 13C–13C spectra shown as contour plots, with positive contours in blue and negative contours in red. Principal component spectra were obtained by singular-value decomposition of the experimental spectra, considering only the aliphatic region and excluding points within 5 p.p.m. of the diagonal. Contour levels increase (or decrease, in the case of negative contours) by successive factors of 1.5. b, Experimental 2D 13C–13C spectrum of t-AD4f Aβ40 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right, with coefficients of PC1–PC5 shown in parentheses). c, Experimental 2D 13C–13C spectrum of r-AD1f Aβ40 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right). d, The first five principal components of the 29 experimental 2D 15N–13C spectra. e, Experimental 2D 15N–13C spectrum of PCA2p Aβ40 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right). f, Experimental 2D 15N–13C spectrum of r-AD2o Aβ40 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right).
Extended Data Figure 6 Principal component analyses of 2D 13C–13C and 15N–13C ssNMR spectra of brain-seeded Aβ42 fibrils.
a, The first five principal components (PC1–PC5) of the 17 experimental 2D 13C–13C spectra, plotted as in Extended Data Fig. 5. b, Experimental 2D 13C–13C spectrum of t-AD1p Aβ42 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right, with coefficients of PC1–PC5 shown in parentheses). c, Experimental 2D 13C–13C spectrum of r-AD2f Aβ42 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right). d, The first five principal components of the 15 experimental 2D 15N–13C spectra. e, Experimental 2D 15N–13C spectrum of t-AD3f Aβ42 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right). f, Experimental 2D 15N–13C spectrum of r-AD2p Aβ42 fibrils (left) and 2D spectrum constructed as a linear combination of PC1–PC5 (right).
Extended Data Figure 7 Analysis of 2D 15N–13C ssNMR spectra of brain-seeded fibrils by fitting with cross-peaks at fixed chemical-shift positions.
a, Examples of 2D spectra (of the 29 Aβ40 and 15 Aβ42 spectra with adequate signal-to-noise ratios presented in Table 1), with fitted cross-peak positions indicated by crosses. Red and blue crosses indicate cross-peaks for chemical shift sets ‘a’ and ‘b’, respectively (see Supplementary Methods, Supplementary Discussion and Extended Data Fig. 8). Cyan crosses indicate additional cross-peaks. Contour levels increase by successive factors of 1.4. b, Pairwise differences among fitted cross-peak volumes for spectra of Aβ40 fibrils (left) and Aβ42 fibrils (right), with colour scales representing r.m.s.d. values. Total cross-peak volumes in each spectrum were normalized before calculation of r.m.s.d. values. Results from this cross-peak-fitting analysis are similar to those in Fig. 3, in which the same experimental data were analysed by direct comparisons of signal amplitudes in 2D spectra without fitting the signals with cross-peaks at specific positions. c, Fractions of the total fitted cross-peak volumes at ‘a’ and ‘b’ chemical shifts, with mean values indicated by horizontal bars. For Aβ40, mean values of ‘a’ volumes in spectra of t-AD (n = 12) or PCA-AD (n = 6) samples are significantly greater than the mean value (n = 10) in spectra of r-AD samples (P < 0.02, Welch’s t-test; P < 0.02, Mann–Whitney–Wilcoxon test).
Extended Data Figure 8 Comparisons of ssNMR chemical shifts of brain-seeded Aβ40 and Aβ42 fibrils with previously reported chemical shifts.
a, 15N and 13C chemical shifts (p.p.m.) from spectra of brain-seeded samples in Table 1 (grouped into sets ‘a’, ‘b’, and so on, based on correlations of the corresponding signal amplitudes over multiple 2D spectra) are compared with chemical shifts from previous ssNMR studies of Aβ40 and Aβ42 fibrils, as deposited in the Biological Magnetic Resonance Bank (http://www.bmrb.wisc.edu/) with the indicated BMRB accession numbers. b, Differences in chemical shift after adjustments of chemical shift referencing in each set to make the average 13Cα shifts and the average 15N shifts equal in all sets.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, a Supplementary Discussion and Supplementary References. The Supplementary Methods describe the selection of brain tissue samples, preparation of brain-seeded fibrils, conditions for ssNMR and TEM measurements, determination of Aβ40/Aβ42 ratios, RMSD, principal component, and crosspeak fitting calculations, statistical tests, and code availability and the Supplementary Discussion describes control experiments, chemical shift comparisons, ssNMR linewidths, and consistency among samples. (PDF 361 kb)
Rights and permissions
About this article
Cite this article
Qiang, W., Yau, WM., Lu, JX. et al. Structural variation in amyloid-β fibrils from Alzheimer's disease clinical subtypes. Nature 541, 217–221 (2017). https://doi.org/10.1038/nature20814
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20814
This article is cited by
-
Misfolded protein oligomers: mechanisms of formation, cytotoxic effects, and pharmacological approaches against protein misfolding diseases
Molecular Neurodegeneration (2024)
-
A one-two punch targeting reactive oxygen species and fibril for rescuing Alzheimer’s disease
Nature Communications (2024)
-
Uncovering supramolecular chirality codes for the design of tunable biomaterials
Nature Communications (2024)
-
Iatrogenic Alzheimer’s disease in recipients of cadaveric pituitary-derived growth hormone
Nature Medicine (2024)
-
Exploring Heparan Sulfate Proteoglycans as Mediators of Human Mesenchymal Stem Cell Neurogenesis
Cellular and Molecular Neurobiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.