Abstract
Ageing is driven by a loss of transcriptional and protein homeostasis1,2,3 and is the key risk factor for multiple chronic diseases. Interventions that attenuate or reverse systemic dysfunction associated with age therefore have the potential to reduce overall disease risk in the elderly. Precursor mRNA (pre-mRNA) splicing is a fundamental link between gene expression and the proteome, and deregulation of the splicing machinery is linked to several age-related chronic illnesses4,5. However, the role of splicing homeostasis in healthy ageing remains unclear. Here we demonstrate that pre-mRNA splicing homeostasis is a biomarker and predictor of life expectancy in Caenorhabditis elegans. Using transcriptomics and in-depth splicing analysis in young and old animals fed ad libitum or subjected to dietary restriction, we find defects in global pre-mRNA splicing with age that are reduced by dietary restriction via splicing factor 1 (SFA-1; the C. elegans homologue of SF1, also known as branchpoint binding protein, BBP). We show that SFA-1 is specifically required for lifespan extension by dietary restriction and by modulation of the TORC1 pathway components AMPK, RAGA-1 and RSKS-1/S6 kinase. We also demonstrate that overexpression of SFA-1 is sufficient to extend lifespan. Together, these data demonstrate a role for RNA splicing homeostasis in dietary restriction longevity and suggest that modulation of specific spliceosome components may prolong healthy ageing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kenyon, C. J. The genetics of ageing. Nature 464, 504–512 (2010)
Taylor, R. C. & Dillin, A. Aging as an event of proteostasis collapse. Cold Spring Harb. Perspect. Biol . 3, a004440 (2011)
Vermulst, M. et al. Transcription errors induce proteotoxic stress and shorten cellular lifespan. Nat. Commun. 6, 8065 (2015)
Wang, G.-S. & Cooper, T. A. Splicing in disease: disruption of the splicing code and the decoding machinery. Nat. Rev. Genet. 8, 749–761 (2007)
Singh, R. K. & Cooper, T. A. Pre-mRNA splicing in disease and therapeutics. Trends Mol. Med. 18, 472–482 (2012)
Lee, B. P. et al. Changes in the expression of splicing factor transcripts and variations in alternative splicing are associated with lifespan in mice and humans. Aging Cell 15, 903–913 (2016)
Seo, M., Park, S., Nam, H. & Lee, S.-J. RNA helicase SACY-1 is required for longevity caused by various genetic perturbations in Caenorhabditis elegans . Cell Cycle 15, 1821–1829 (2016)
Rodríguez, S. A. et al. Global genome splicing analysis reveals an increased number of alternatively spliced genes with aging. Aging Cell 15, 267–278 (2016)
Curran, S. P. & Ruvkun, G. Lifespan regulation by evolutionarily conserved genes essential for viability. PLoS Genet. 3, e56 (2007)
Gao, X. et al. The survival motor neuron gene smn-1 interacts with the U2AF large subunit gene uaf-1 to regulate Caenorhabditis elegans lifespan and motor functions. RNA Biol. 11, 1148–1160 (2014)
Zhang, T., Hwang, H.-Y., Hao, H., Talbot, C., Jr & Wang, J. Caenorhabditis elegans RNA-processing protein TDP-1 regulates protein homeostasis and life span. J. Biol. Chem. 287, 8371–8382 (2012)
Kuroyanagi, H., Watanabe, Y., Suzuki, Y. & Hagiwara, M. Position-dependent and neuron-specific splicing regulation by the CELF family RNA-binding protein UNC-75 in Caenorhabditis elegans . Nucleic Acids Res. 41, 4015–4025 (2013)
Zahler, A. Pre-mRNA splicing and its regulation in Caenorhabditis elegans . Wormbook http://dx.doi.org/10.1895/wormbook.1.31.2 (2012)
Ramani, A. K. et al. Genome-wide analysis of alternative splicing in Caenorhabditis elegans . Genome Res. 21, 342–348 (2011)
Herndon, L. A. et al. Stochastic and genetic factors influence tissue-specific decline in ageing C. elegans . Nature 419, 808–814 (2002)
Ching, T.-T. T. & Hsu, A.-L. L. Solid plate-based dietary restriction in Caenorhabditis elegans . J. Vis. Exp. (51):2701 (2011)
Lakowski, B. & Hekimi, S. The genetics of caloric restriction in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 95, 13091–13096 (1998)
Krämer, A. Purification of splicing factor SF1, a heat-stable protein that functions in the assembly of a presplicing complex. Mol. Cell. Biol. 12, 4545–4552 (1992)
Ma, L. & Horvitz, H. R. Mutations in the Caenorhabditis elegans U2AF large subunit UAF-1 alter the choice of a 3′ splice site in vivo . PLoS Genet. 5, e1000708 (2009)
Ma, L., Tan, Z., Teng, Y., Hoersch, S. & Horvitz, H. R. In vivo effects on intron retention and exon skipping by the U2AF large subunit and SF1/BBP in the nematode Caenorhabditis elegans. RNA 17, 2201–2211 (2011)
Fontana, L., Partridge, L. & Longo, V. D. Extending healthy life span--from yeast to humans. Science 328, 321–326 (2010)
Johnson, S. C., Rabinovitch, P. S. & Kaeberlein, M. mTOR is a key modulator of ageing and age-related disease. Nature 493, 338–345 (2013)
Stamm, S. Regulation of alternative splicing by reversible protein phosphorylation. J. Biol. Chem. 283, 1223–1227 (2008)
Yu, Y. et al. Phosphoproteomic analysis identifies Grb10 as an mTORC1 substrate that negatively regulates insulin signaling. Science 332, 1322–1326 (2011)
Hsu, P. P. et al. The mTOR-regulated phosphoproteome reveals a mechanism of mTORC1-mediated inhibition of growth factor signaling. Science 332, 1317–1322 (2011)
Wang, X. et al. Phosphorylation of splicing factor SF1 on Ser20 by cGMP-dependent protein kinase regulates spliceosome assembly. EMBO J. 18, 4549–4559 (1999)
Tanackovic, G. & Krämer, A. Human splicing factor SF3a, but not SF1, is essential for pre-mRNA splicing in vivo . Mol. Biol. Cell 16, 1366–1377 (2005)
Zhang, D., Paley, A. J. & Childs, G. The transcriptional repressor ZFM1 interacts with and modulates the ability of EWS to activate transcription. J. Biol. Chem. 273, 18086–18091 (1998)
Mair, W. et al. Lifespan extension induced by AMPK and calcineurin is mediated by CRTC-1 and CREB. Nature 470, 404–408 (2011)
Burkewitz, K. et al. Neuronal CRTC-1 governs systemic mitochondrial metabolism and lifespan via a catecholamine signal. Cell 160, 842–855 (2015)
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Anders, S., Pyl, P. T. & Huber, W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Cunningham, F. et al. Ensembl 2015. Nucleic Acids Res. 43, D662–D669 (2015)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Hansen, K. D., Irizarry, R. A. & Wu, Z. Removing technical variability in RNA-seq data using conditional quantile normalization. Biostatistics 13, 204–216 (2012)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010)
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinformatics 10, 161 (2009)
Mazin, P. et al. Widespread splicing changes in human brain development and aging. Mol. Syst. Biol. 9, 633–633 (2013)
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012)
Acknowledgements
C.H. is supported by the Swiss National Science Foundation (P2ZHP3_151609) and the Fonds National de la Recherche Luxembourg (AFR7883116). W.B.M. is funded by The Lawrence Ellison Medical Foundation (U54CA155626), The Glenn Foundation for Medical Research and the National Institutes of Health (NIH, 1R01AG044346). C.E. is supported by the Ligue Nationale contre le Cancer. G.H.B and T.K.D. were supported through a grant to B.S.A. from The Novo Nordisk Foundation (NNF13OC0007939). We are grateful to H. Kuroyanagi for providing the splicing reporter strain and corresponding plasmids. We thank K. Blackwell for providing the glp-4 mutant strain and RNAi constructs. We also thank the Caenorhabditis Genetics Center for providing worm strains. We also thank the Mair laboratory members for comments and discussion on the project and manuscript.
Author information
Authors and Affiliations
Contributions
C.H. and W.B.M. designed the study, C.H. performed the majority of the experiments and analysed results, A.L. performed experiments and edited the manuscript. T.K.D. analysed all the RNA-seq data. C.C.E., C.G.S.-G. and S.D. performed lifespan repeats and C. elegans sample collections, I.M. helped with cloning and crosses. H.J.W. performed the oxygen consumption experiments. G.H.B. and B.S.A. designed and performed the HeLa cells experiments including the sequencing run. Y.Z., G.H. and B.D.M. designed and performed the MEF and C. elegans immunoblotting experiments. C.H. and W.B.M. wrote the manuscript implementing comments and edits from all authors.
Corresponding author
Additional information
Reviewer Information Nature thanks M. Kaeberlein, N. Tavernarakis and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Heterogeneous splicing patterns in response to knockdown of conserved splicing factors.
a, Inverted fluorophore splicing reporter. b, Simplified diagram of C. elegans intron splicing showing representative splicing factors investigated herein c, C. elegans splicing factors and their mammalian homologues. d–l, Knockdown of hrp-2 in splicing reporter (d) and hrp-2 in inverted reporter (e), uaf-2 (f), snr-1 (g), prp-38 (h), rsp-2 (i), prp-8 (j), unc-75 (k) and uaf-1 (l) by RNAi at day 1 of adulthood. m, hrp-1 depletion at day 4 of adulthood. n, o, Representative images of worms with phi-9 (n) and hrpf-1 (o) knockdown in day 1 adults with reduced exon inclusion.
Extended Data Figure 2 Effects of splicing factor knockdown on splicing homeostasis and dietary-restriction-mediated longevity.
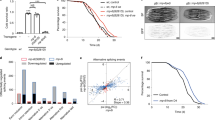
a, RNA-seq coverage tracks for endogenous ret-1 splicing in hrp-2 knockdown samples. Sequencing reads tracks generated by Splicing Java Coverage Viewer as part of SAJR40. Height of red lines represents RNA coverage of splice junctions, dark grey boxes represent exonic sequence, light grey box denotes alternative exon sequence. b, Endogenous ret-1 splicing exon 5 skipping in wild-type and hrp-2 RNAi worms by RT–PCR (three biological replicates). c, d, Intronic reads (c, P = 0.0042) and unannotated junctions reads (d, P = 0.056) as hallmarks of deregulated splicing with hrp-2 knockdown. Three biological replicates for control and with hrp-2 RNAi. Mean ± s.e.m., per cent of total reads shown. P values calculated with unpaired, two-tailed t-test after probit transformation. e, Differentially regulated alternative splicing events induced by hrp-2 depletion (****P < 0.0001 for exon inclusion, intron retention P < 0.0001, Pearson’s χ2 test). f, Proportion plot of all exon skipping events with proportions of novel and known exons up- or downregulated in hrp-2 knockdown samples (*P = 0.0157, differences in proportions of novel exons in up- and downregulated events were tested with Pearson’s χ2 test, deviations from an even proportion of up- and downregulated splicing events were tested with binomial test). g, EGFP and mCherry mRNA levels up to day 8 of adulthood by qRT–PCR (mean + s.d., technical replicates shown, 1 of 2 biological replicates). h, sDR robustly extends the lifespan of C. elegans fed ad libitum16 (P < 0.0001). i, Fluorescence quantification of splice isoforms in 7-day-old ad libitum and sDR animals (****P < 0.0001, unpaired two-tailed t-test, mean + s.d., n = 8 worms per condition, 6 biological replicates). j, Age-matched, ad libitum-fed worm populations separated at day 6 according to group A (increased exon 5 skipping) and group B (increased exon 5 inclusion) (mean + s.d., n = 6, 1 of 3 biological replicates shown). k, Effect of hrpf-1 RNAi on wild-type and eat-2(ad1116) lifespan (wild-type versus eat-2(ad1116) with hrpf-1 RNAi, P < 0.0001). l, Comparison of survival rates of wild-type and eat-2(ad1116) with snr-1 RNAi (P = 0.5147). m, n, Survival analysis of hrp-2 (m) and uaf-2 (n) downregulation by RNAi (wild-type with hrp-2 RNAi versus eat-2(ad1116) with hrp-2 RNAi, P = 0.1617, 2 replicates; wild-type with uaf-2 RNAi versus eat-2(ad1116) with uaf-2 RNAi, P = 0.001, 3 replicates). o, Effect of snr-2 knockdown on wild-type and dietary-restricted lifespan (wild-type with snr-2 RNAi versus eat-2(ad1116) with snr-2 RNAi, P = 0.3456, 2 replicates). p, Wild-type and dietary-restricted lifespan curves with rsp-2 knockdown (wild-type with rsp-2 RNAi versus eat-2(ad1116) with rsp-2 RNAi, P < 0.0001, 3 replicates). Lifespan experiments done with FUDR as indicated in Extended Data Table 1 and Supplementary Table 10. n = 100 worms per condition for lifespan analysis. P values of survival analysis calculated with log-rank test.
Extended Data Figure 3 Effects of sfa-1 downregulation on splicing.
a, Expression of uaf-2 is not affected by reduced sfa-1 levels in wild-type worms at day 1 of adulthood (mean + s.d., technical replicates shown). b, c, Effect of reduced sfa-1 expression on pumping rates in wild-type and a genetic dietary restriction model (eat-2(ad1116)) at day 1 (b) and 4 of adulthood (c) (mean + s.d., ns, P > 0.05, unpaired, two-tailed t-test, n = 10 worms per condition, 1 of 2 replicates shown). d, Splicing reporter pattern with sfa-1 knockdown from egg hatch, day 1, 3 and 7 adults. e, Endogenous ret-1 exon 5 splicing pattern with age and sfa-1 RNAi in wild-type and dietary-restricted worms by RT–PCR (day 3 versus 15, fifth replicate sample set).
Extended Data Figure 4 RT–PCR validation of alternative splicing events in ageing and with sfa-1 knockdown.
a, Sequencing reads coverage for tos-1 b, Age-associated isoform ratio change of a target of SFA-1, tos-1, in wild-type worms at day 3 and 15 of adulthood with or without sfa-1 RNAi by RT–PCR (two biological replicates, distinct from those used in Fig. 1i, shown). c, Sequencing read coverage map for ret-1 shows increased exon 5 skipping with age and with sfa-1 RNAi. d, Endogenous ret-1 exon 5 splicing pattern with age and sfa-1 RNAi in wild-type and dietary-restricted worms by RT–PCR (day 3 versus 15, two biological replicates shown). e, Sequencing tracks for lipl-7 pre-mRNA. f, Monitoring of intron retention between exons 4 and 5 at day 15 versus day 3 of adulthood in wild-type and dietary-restricted worms with or without sfa-1 RNAi. g, Sequencing reads tracks for slo-2 pre-mRNA. h, slo-2 alternative exon skipping in day 3 and day 15 wild-type and dietary-restricted worms with or without sfa-1 RNAi. i, Sequencing reads tracks for lea-1 pre-mRNA j, Alternative exon skipping in lea-1 with age and sfa-1 knockdown in wild-type and dietary-restricted animals. Two biological replicates shown for all RT–PCR analyses. Sequencing reads tracks generated by Splicing Java Coverage Viewer as part of SAJR40; height of red lines represent RNA coverage of splice junctions, dark grey boxes represent exonic sequence, light grey boxes denote the alternative exon sequence.
Extended Data Figure 5 RNA-seq expression data validation by quantitative RT–PCR.
a–j, Monitoring of gene expression levels by quantitative RT–PCR for RNA-seq data validation of acs-2 (a), rsr-2 (b), fat-6 (c), fat-5 (d), fat-7 (e), acs-17 (f), acdh-2 (g), cpr-1 (h), lips-17 (i) and gst-4 (j) in 6 biological replicates for 3-day-old wild-type worms and 5 biological replicates for all other samples at day 15 (****P ≤ 0.0001, ***P ≤ 0.001, **P ≤ 0.01, *P ≤ 0.05; ns, P >0.05; RNA-seq data are mean + s.e.m. of normalized read counts of 4 biological replicates. qRT–PCR data are mean + s.e.m. P values calculated with unpaired, two-tailed t-test).
Extended Data Figure 6 Genome-wide effects of dietary restriction and SFA-1 depletion on pre-mRNA splicing and metabolism.
a, Differentially regulated splicing events (exons, introns, alternative 5′ and 3′ splice sites) in dietary restriction at day 15 compared to feeding ad libitum (*P = 0.0156, Wilcoxon signed rank test). b, Multidimensional scaling plot of significantly different splicing patterns using inclusion ratio estimates between day 3 and day 15 worms (changes in all significant pre-mRNA segments, for example, exons, introns, alternative splice sites, considered). c, Venn diagram representing significantly up- or downregulated novel splicing events at day 15 in worms fed ad libitum, worms on dietary restriction and dietary restriction with sfa-1 RNAi (subset of unannotated splice junctions). d, Differentially regulated splicing events (exons, introns, alternative 5′ and 3′ splice sites) with sfa-1 knockdown (ad libitum with sfa-1 RNAi versus dietary restriction with sfa-1 RNAi, P = 0.7999, Wilcoxon signed rank test). e, KEGG pathways significantly upregulated in dietary-restricted worm populations at day 15 compared to wild-type worm populations of the same chronological age with false discovery rate (FDR) of 10%. f, KEGG pathways significantly upregulated in dietary-restricted worm populations with sfa-1 knockdown at day 15 compared to worm populations fed ad libitum with FDR 10%. g, h, Basal respiration (g) and reserve capacity (h) in wild-type and dietary-restricted animals (mean + s.e.m., ***P ≤ 0.001, *P ≤ 0.05, unpaired two-tailed t-test). Results shown are oxygen consumption rates of 15-day-old worms normalized to 4-day-old populations, n = 100 worms per condition (1 of 3 replicate experiments). i, Transcriptional induction of acs-2p::GFP in fed control and sfa-1 knockdown worms (representative image of two repeat experiments shown, n worms: empty vector, fed, n = 83; empty vector, fasted, n = 77; sfa-1 RNAi, fed, n = 76; sfa-1 RNAi, fasted, n = 84). j, Quantification of acs-2p::GFP after 23 h of fasting and sfa-1 knockdown (mean + s.e.m. of two replicate experiments, n as in i, P < 0.0001, unpaired two-tailed t-test).
Extended Data Figure 7 Effects of SF1 knockdown in HeLa cells.
a, Effect of SF1 knockdown in HeLa cells on unannotated junction reads (mean ± s.e.m., ***P = 0.0006) and reads in introns (mean ± s.e.m., *P = 0.0242, unpaired, two-tailed t-test after probit transformation) Three biological replicate samples. b, Differentially regulated alternative splicing events (exon skipping, intron retention, alternative 5′ and 3′ splice sites) with SF1 knockdown. Exons are primarily downregulated, whereas introns are significantly upregulated (**** P < 0.0001, Pearson χ2 test). c, KEGG pathway analysis of gene expression changes upon SF1 knockdown. Pathways with P ≤ 0.05 considered significant, P values derived by gage39. q values are P values adjusted for multiple testing using the Benjamini–Hochberg method37.
Extended Data Figure 8 SFA-1 and the mTORC1 pathway.
a, b, Quantification of ret-1 minigene exon inclusion (GFP intensity) in aak-2(ok524) (a; ad libitum versus dietary restriction, P = 0.5488, unpaired two-tailed t-test, mean + s.d., n = 8, 3 replicates), and in daf-16(mu86) (b; ad libitum versus dietary restriction, P = 0.1835, unpaired two-tailed t-test, mean + s.d., n = 8, 2 replicates) c, Quantification of GFP in raga-1(ok386) mutants at day 8 in ad libitum and dietary restriction conditions (****P < 0.0001, *P < 0.05, n = 8, unpaired two-tailed t-test, mean + s.d.). One of three replicate imaging experiments shown. d, Splicing reporter expression in day 1 wild-type and raga-1(ok386) animals. e, Immunoblot of proteins from wild-type and raga-1(ok386) animals assaying S6K phosphorylation state (*P = 0.017, unpaired two-tailed t-test, mean + s.e.m., 3 biological replicates). f, tos-1 isoform ratios in raga-1(ok386) mutants at day 3 and 15 of adulthood with sfa-1 RNAi (2 biological replicates shown). g, Survival analysis of sfa-1 RNAi in constitutively active (CA) AMPK mutant (P = 0.4844, wild-type with sfa-1 RNAi versus CA AMPK with sfa-1 RNAi). h, Survival analysis of SFA-1 knockdown in insulin/IGF signalling-mediated longevity (daf-2(e1370), P < 0.0001, wild-type with sfa-1 RNAi versus daf-2(e1370) with sfa-1 RNAi. i, Immunoblots of proteins from wild-type mouse embryonic fibroblasts. p-S6K(T389) and p-S6(S240/S244) (assaying phosphorylation at sites T389 (S6K) and S240/S244 (S6), markers of mTORC1 activation), total S6K, total S6, non-targeting control short interfering RNAs (siCt), SF1 siRNA, and β-actin (loading control) are shown. j, Immunoblots of proteins from wild-type mouse embryonic fibroblasts treated for 16 h. Biological duplicates are shown except for the siRNA-treated lanes. n = 100 worms per condition for lifespan analysis. P values for survival analysis calculated with log-rank test.
Extended Data Figure 9 Differential effects of sfa-1 and repo-1 knockdown in multiple longevity pathways.
a, Effect of raga-1 RNAi in smg-1(cc546) mutant worms, which are defective in nonsense-mediated decay (P = 0.041 between smg-1(cc546) with and without raga-1 RNAi, lifespan at 24 °C, Gehan–Breslow–Wilcoxon test, 3 replicates). b, Survival of wild-type and (eat-2(ad1116)) worms with repo-1 RNAi treatment (P < 0.0001 between eat-2(ad1116) with and without repo-1 RNAi, 4 replicates). c, sfa-1 RNAi blocks RAGA-1 mediated longevity (P = 0.2181, wild-type with sfa-1 RNAi versus raga-1(ok386) with sfa-1 RNAi, Gehan–Breslow–Wilcoxon test, 7 replicates). d, repo-1 RNAi has no effect on raga-1(ok386) longevity (P < 0.0001, wild-type with repo-1 RNAi versus raga-1(ok386) with repo-1 RNAi, 3 replicates). e, Effect of sfa-1 RNAi on mitochondrial ETC mutant isp-1(qm150)-mediated longevity (P = 0.004, wild-type with sfa-1 RNAi versus isp-1(qm150) with sfa-1 RNAi, 2 replicates). f, repo-1 RNAi shortens isp-1(qm150)-mediated longevity to wild-type levels (P = 0.4951, wild-type with repo-1 RNAi versus isp-1(qm150) with repo-1 RNAi, 2 replicates). g, Effect of SFA-1 overexpression on wild-type lifespan on OP50-1 bacteria (P < 0.0001 both lines versus wild type, 3 replicates). h, Survival analysis of wild-type and SFA-1 overexpression lines on sfa-1 RNAi (P = 0.0042, wild-type with sfa-1 RNAi versus sfa-1 OE line 1 with sfa-1 RNAi, 2 replicates, P > 0.05, wild-type empty vector versus sfa-1 OE line 1 with sfa-1 RNAi, 2 replicates). i, Top, monitoring of sfa-1 levels by qRT–PCR (C, injection marker control line; 1, 2, SFA-1 overexpression lines; error bars, mean + s.d. of two biological replicates for strains grown on HT115 bacteria (left); error bars, mean + s.d. of two technical replicates for strains grown on OP50-1 bacteria (right)). Bottom, tos-1 isoform ratios with SFA-1 overexpression in day 1 adults. n = 100 worms per condition for lifespan analysis. P values of survival analysis calculated with log-rank test.
Supplementary information
Supplementary Information
This file contains Supplementary Data 1 - all primers with sequence information used for PCR and qRT-PC, Supplementary Data 2 - all RNAi clone sequences used in this work and Supplementary Figure 1 - uncropped agarose gels and immunoblots. Please note, Supplementary Figure 1 has incorrect labelling, see the Corrigendum associated with this Letter for the corrected Supplementary Figure 1. (PDF 548 kb)
Supplementary Tables
This zipped file contains Supplementary Tables 1-10 and a Supplementary Table guide. (ZIP 10967 kb)
Source data
Rights and permissions
About this article
Cite this article
Heintz, C., Doktor, T., Lanjuin, A. et al. Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans. Nature 541, 102–106 (2017). https://doi.org/10.1038/nature20789
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20789
This article is cited by
-
The CRTC-1 transcriptional domain is required for COMPASS complex-mediated longevity in C. elegans
Nature Aging (2023)
-
Genome-wide CRISPR screens identify ILF3 as a mediator of mTORC1-dependent amino acid sensing
Nature Cell Biology (2023)
-
Spatial and single-cell profiling of the metabolome, transcriptome and epigenome of the aging mouse liver
Nature Aging (2023)
-
Ageing-associated changes in transcriptional elongation influence longevity
Nature (2023)
-
ATF-4 and hydrogen sulfide signalling mediate longevity in response to inhibition of translation or mTORC1
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.