Abstract
The heterotrimeric influenza polymerase (FluPol), comprising subunits PA, PB1 and PB2, binds to the conserved 5′ and 3′ termini (the ‘promoter’) of each of the eight single-stranded viral RNA (vRNA) genome segments and performs both transcription and replication of vRNA in the infected cell nucleus1,2,3. To transcribe viral mRNAs, FluPol associates with cellular RNA polymerase II (Pol II)4,5,6,7, which enables it to take 5′-capped primers from nascent Pol II transcripts8,9. Here we present a co-crystal structure of bat influenza A polymerase bound to a Pol II C-terminal domain (CTD) peptide mimic, which shows two distinct phosphoserine-5 (SeP5)-binding sites in the polymerase PA subunit, accommodating four CTD heptad repeats overall. Mutagenesis of the SeP5-contacting basic residues (PA K289, R454, K635 and R638) weakens CTD repeat binding in vitro without affecting the intrinsic cap-primed (transcription) or unprimed (replication) RNA synthesis activity of recombinant polymerase, whereas in cell-based minigenome assays the same mutations substantially reduce overall polymerase activity. Only recombinant viruses with a single mutation in one of the SeP5-binding sites can be rescued, but these viruses are severely attenuated and genetically unstable. Several previously described mutants that modulate virulence can be rationalized by our results, including a second site mutation (PA(C453R)) that enables the highly attenuated mutant virus (PA(R638A)) to revert to near wild-type infectivity10. We conclude that direct binding of FluPol to the SeP5 Pol II CTD is fine-tuned to allow efficient viral transcription and propose that the CTD-binding site on FluPol could be targeted for antiviral drug development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Sequence Read Archive
References
Eisfeld, A. J., Neumann, G. & Kawaoka, Y. At the centre: influenza A virus ribonucleoproteins. Nat. Rev. Microbiol. 13, 28–41 (2015)
Ortín, J. & Martín-Benito, J. The RNA synthesis machinery of negative-stranded RNA viruses. Virology 479-480, 532–544 (2015)
Te Velthuis, A. J. & Fodor, E. Influenza virus RNA polymerase: insights into the mechanisms of viral RNA synthesis. Nat. Rev. Microbiol. 14, 479–493 (2016)
Mahy, B. W., Hastie, N. D. & Armstrong, S. J. Inhibition of influenza virus replication by alpha-amanitin: mode of action. Proc. Natl Acad. Sci. USA 69, 1421–1424 (1972)
Engelhardt, O. G., Smith, M. & Fodor, E. Association of the influenza A virus RNA-dependent RNA polymerase with cellular RNA polymerase II. J. Virol. 79, 5812–5818 (2005)
Loucaides, E. M. et al. Nuclear dynamics of influenza A virus ribonucleoproteins revealed by live-cell imaging studies. Virology 394, 154–163 (2009)
Chan, A. Y., Vreede, F. T., Smith, M., Engelhardt, O. G. & Fodor, E. Influenza virus inhibits RNA polymerase II elongation. Virology 351, 210–217 (2006)
Plotch, S. J., Bouloy, M., Ulmanen, I. & Krug, R. M. A unique cap(m7GpppXm)-dependent influenza virion endonuclease cleaves capped RNAs to generate the primers that initiate viral RNA transcription. Cell 23, 847–858 (1981)
Reich, S. et al. Structural insight into cap-snatching and RNA synthesis by influenza polymerase. Nature 516, 361–366 (2014)
Fodor, E., Mingay, L. J., Crow, M., Deng, T. & Brownlee, G. G. A single amino acid mutation in the PA subunit of the influenza virus RNA polymerase promotes the generation of defective interfering RNAs. J. Virol. 77, 5017–5020 (2003)
Gu, W. et al. Influenza A virus preferentially snatches noncoding RNA caps. RNA 21, 2067–2075 (2015)
Palancade, B. & Bensaude, O. Investigating RNA polymerase II carboxyl-terminal domain (CTD) phosphorylation. Eur. J. Biochem . 270, 3859–3870 (2003)
Lidschreiber, M., Leike, K. & Cramer, P. Cap completion and C-terminal repeat domain kinase recruitment underlie the initiation-elongation transition of RNA polymerase II. Mol. Cell. Biol. 33, 3805–3816 (2013)
Hsin, J. P., Xiang, K. & Manley, J. L. Function and control of RNA polymerase II C-terminal domain phosphorylation in vertebrate transcription and RNA processing. Mol. Cell. Biol. 34, 2488–2498 (2014)
Martinez-Rucobo, F. W. et al. Molecular basis of transcription-coupled pre-mRNA capping. Mol. Cell 58, 1079–1089 (2015)
Rodriguez, A., Pérez-González, A. & Nieto, A. Influenza virus infection causes specific degradation of the largest subunit of cellular RNA polymerase II. J. Virol. 81, 5315–5324 (2007)
Vreede, F. T., Chan, A. Y., Sharps, J. & Fodor, E. Mechanisms and functional implications of the degradation of host RNA polymerase II in influenza virus infected cells. Virology 396, 125–134 (2010)
Pflug, A., Guilligay, D., Reich, S. & Cusack, S. Structure of influenza A polymerase bound to the viral RNA promoter. Nature 516, 355–360 (2014)
Meinhart, A., Kamenski, T., Hoeppner, S., Baumli, S. & Cramer, P. A structural perspective of CTD function. Genes Dev. 19, 1401–1415 (2005)
Martínez-Alonso, M., Hengrung, N. & Fodor, E. RNA-free and ribonucleoprotein-associated influenza virus polymerases directly bind the serine-5-phosphorylated carboxyl-terminal domain of host RNA polymerase II. J. Virol. 90, 6014–6021 (2016)
Robb, N. C., Smith, M., Vreede, F. T. & Fodor, E. NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome. J. Gen. Virol . 90, 1398–1407 (2009)
Kawakami, E. et al. Strand-specific real-time RT–PCR for distinguishing influenza vRNA, cRNA, and mRNA. J. Virol. Methods 173, 1–6 (2011)
Fodor, E. et al. A single amino acid mutation in the PA subunit of the influenza virus RNA polymerase inhibits endonucleolytic cleavage of capped RNAs. J. Virol. 76, 8989–9001 (2002)
Llompart, C. M., Nieto, A. & Rodriguez-Frandsen, A. Specific residues of PB2 and PA influenza virus polymerase subunits confer the ability for RNA polymerase II degradation and virus pathogenicity in mice. J. Virol. 88, 3455–3463 (2014)
Rolling, T. et al. Adaptive mutations resulting in enhanced polymerase activity contribute to high virulence of influenza A virus in mice. J. Virol. 83, 6673–6680 (2009)
Mehle, A., Dugan, V. G., Taubenberger, J. K. & Doudna, J. A. Reassortment and mutation of the avian influenza virus polymerase PA subunit overcome species barriers. J. Virol. 86, 1750–1757 (2012)
Wu, N. C. et al. Functional constraint profiling of a viral protein reveals discordance of evolutionary conservation and functionality. PLoS Genet. 11, e1005310 (2015)
Darnell, J. E., Jr. Reflections on the history of pre-mRNA processing and highlights of current knowledge: a unified picture. RNA 19, 443–460 (2013)
Vreede, F. T. & Fodor, E. The role of the influenza virus RNA polymerase in host shut-off. Virulence 1, 436–439 (2010)
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D 66, 133–144 (2010)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012)
DeLano, W. L. PyMOL Molecular Graphics System. http://www.pymol.sourceforge.net (2002)
Fodor, E. et al. Rescue of influenza A virus from recombinant DNA. J. Virol . 73, 9679–9682 (1999)
Dos Santos Afonso, E., Escriou, N., Leclercq, I., van der Werf, S. & Naffakh, N. The generation of recombinant influenza A viruses expressing a PB2 fusion protein requires the conservation of a packaging signal overlapping the coding and noncoding regions at the 5′ end of the PB2 segment. Virology 341, 34–46 (2005)
Matrosovich, M., Matrosovich, T., Garten, W. & Klenk, H. D. New low-viscosity overlay medium for viral plaque assays. Virol. J. 3, 63 (2006)
Watson, S. J. et al. Viral population analysis and minority-variant detection using short read next-generation sequencing. Phil. Trans. R. Soc. Lond. B 368, 20120205 (2013)
Gabriel, G. et al. The viral polymerase mediates adaptation of an avian influenza virus to a mammalian host. Proc. Natl Acad. Sci. USA 102, 18590–18595 (2005)
Acknowledgements
We thank ESRF for access to X-ray beamlines, the EMBL eukaryotic expression and crystallisation facilities and the biophysical platform within the Partnership for Structural Biology (PSB). D. Guilligay, M. Lethier and S. Gaudon helped with protein expression and crystallization. J. Ortin and T. Wolff supplied plasmids and members of the R. Pillai group (EMBL) provided advice for the minigenome assays. We thank V. Enouf and S. Leandri (Institut Pasteur, Pasteur International Bioresources network, Plateforme de Microbiologie Mutualisée) for the next-generation sequencing analysis and H. Varet (Institut Pasteur) for help with the statistical analysis. We acknowledge A. Politi, N. Daigle and J. Ellenberg for fluorescent microscopy experiments and discussions. This work was supported by ERC Advanced Grant V-RNA (322586) to S.C. and by the Integrative Biology of Emerging Infectious Diseases Laboratory of Excellence to N.N. The Institut Carnot Pasteur Maladies Infectieuses and the EU PREDEMICS project (278433) supported G.F.
Author information
Authors and Affiliations
Contributions
M.L. performed protein expression, purification and crystallization with the help of A.P. and S.R. X-ray data collection and crystallographic analysis was performed by A.P., S.C. and M.L. Virus rescue experiments and RNA quantification were performed by G.F. under the supervision of N.N. M.L. performed peptide binding and polymerase activity assays, designed by S.R., who also helped with data analysis. P.R.-I. performed activity assays and primer extension assays. M.L. performed the minigenome experiments with the help of P.R-.I. S.C. and N.N. designed and supervised the project. S.C. and M.L wrote the manuscript with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks K. Murakami, S. Shuman and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
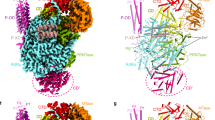
Extended Data Figure 2 Conformations of CTD peptides bound to bat FluA polymerase compared to other Pol-II-interacting factors.
Structures of proteins bound to SeP5 or SeP2-SeP5 CTD peptides are shown in grey cartoon, the peptides (stick representation) are coloured to highlight their position and zoomed in next to their corresponding structure. a, b, Binding site 1 (a) and site 2 (b) of bat FluA polymerase, with the corresponding parts of the CTD SeP5 peptides. c, SeP2–SeP5 CTD bound to the peptidyl-prolyl isomerase Pin1 (PDB: 1F8A). d, SeP5 CTD bound to Candida albicans capping-enzyme Cgt1 (PDB: 1P16). e, SeP2–SeP5 CTD bound to mammalian capping-enzyme Mce1 (PDB: 3RTX). f, SeP2–SeP5 CTD bound to Schizosaccharomyces pombe capping-enzyme Pce1 (PDB: 4PZ6). g, SeP5 CTD bound to human serine phosphatase Ssu72 (PDB: 3O2Q).
Extended Data Figure 3 Sequence alignment of the PA subunit of various influenza strains (bat A, human A, avian A, B, C, D).
Only the PA-C region starting from residue 220 is shown. Absolutely conserved residues are white on a red background and highly conserved residues are red letters boxed in blue. The amino acid numbering and secondary structure correspond to bat FluA polymerase. CTD binding site 1 residues are indicated with a cyan triangle (conserved in FluA and FluB strains) and site 2 with a yellow triangle (key residues are only conserved in FluA strains). Residues referred to in the discussion (site 1, C448 (H1N1, C453), site 2, I545 (H1N1, L550), T547 (H1N1, S552)) are shown with a black triangle.
Extended Data Figure 4 Fluorescence anisotropy data for bat FluA polymerase.
a, Displacement of a fluorescent SeP5 CTD peptide (four-repeat) bound to Pol–vRNA complex with non-labelled SeP5 peptide. The apparent Kd (Kd′) is shown. b, Binding of SeP5 peptide to FluA polymerase in the absence of the vRNA promotor. c, Interaction with non-phosphorylated four-repeat CTD peptide (Y1S2P3T4S5P6S7). The binding curve can be extrapolated to an estimated Kd of >10 μM. Error bars represent s.d. of three independent experiments, Kd values are shown ± the error of fit.
Extended Data Figure 5 In vitro data for CTD-binding-deficient mutants.
Binding data of SeP5 peptide to recombinant mutant proteins. a, Bat FluA site 2 double mutant K289A/R449A (left), site 1 double mutant K630A/R633A (right)). b, Influenza B site 1 double mutant K631A/R634A. Error bars represent s.d. of three independent experiments, Kd values are shown ± the error of the fit. c, Time courses of cap-primed transcription (left panel) and unprimed replication reactions in vitro with vRNA (middle panel) and cRNA (right panel) as template, comparing the site 1 and 2 double mutants with the wild-type bat FluA. Error bars show the s.d. from three different reactions. The tables show the measured rate constants ± the error of the fit.
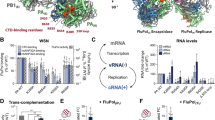
Extended Data Figure 6 Minigenome assays for CTD-binding mutants of influenza A/WSN/33 in absence of NS2.
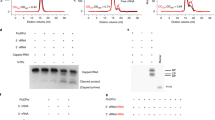
a, Overall RNP activity measured by a luciferase reporter assay (upper panel). Expression of PA, PB2 and nucleoprotein subunits is monitored by western blot (lower panel). b, Primer extension assay showing the different RNA levels produced by wild-type and mutant polymerases in the minigenome assay. Products of primer extension reactions were loaded on a 6% urea polyacrylamide gel, with their expected size indicated on the right and molecular weight markers on the left. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 7 RNA quantification of the A/WSN/33 minigenome assay with NS2.
Normalized copy numbers of the different RNAs produced by wild-type and mutant polymerases, measured by quantitative RT–PCR. Error bars represent the s.d. of three independent experiments. Values lower than 105 are below the detection limit of the assay (indicated by a dashed line).
Extended Data Figure 8 Mutant A/WSN/33 virus rescue experiments.
The titers and plaque phenotypes of the recombinant A/WSN/33 viruses were determined on reverse genetics supernatants (left) or upon a single plaque purification followed by viral amplification (right), using a standard plaque assay on MDCK cells. Crystal violet staining of cell monolayers infected with the indicated viral dilutions is shown. WT, control performed with wild-type PA. For each recombinant virus, two independent viral stocks (denoted as P1 and P1′) derived from the same purified plaque were subjected to vRNA extraction, RT–PCR of the eight genomic segments and next-generation sequencing. The non-synonymous mutations detected in >30% of the reads upon alignment with the reference sequences for the wild-type virus are indicated. *Number of purified plaques leading to successful viral amplification per total number tested. †P1′ did not yield any detectable infectious virus for the R454A mutant. ‡P1 (for R454A and wild type) and P1′ (for K289A, K635A and R638A) were assayed in parallel.
Extended Data Figure 9 Overlap between a functionally important region of FluA PA defined by fitness profiling and the CTD-binding site.
a, b, A putative functional subdomain (a) identified in figure 5 of ref. 27 partially overlaps with the two CTD-binding sites (b). Amongst the residues shown to be functionally important in ref. 27, K559, R454 and K635 (K554, R449 and K630 bat FluA numbering) are directly involved in the interaction with the CTD peptide (see Extended Data Fig. 3). The authors also showed that R454 and K635 mutations essentially eliminated polymerase activity in a minigenome assay, consistent with our results.
Supplementary information
Supplementary Figures
This file contains the uncropped gels with molecular markers for Figure 3b,d and Extended Data Figure 7a,b. (PDF 927 kb)
Rights and permissions
About this article
Cite this article
Lukarska, M., Fournier, G., Pflug, A. et al. Structural basis of an essential interaction between influenza polymerase and Pol II CTD. Nature 541, 117–121 (2017). https://doi.org/10.1038/nature20594
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20594
This article is cited by
-
The host RNA polymerase II C-terminal domain is the anchor for replication of the influenza virus genome
Nature Communications (2024)
-
The ubiquitination landscape of the influenza A virus polymerase
Nature Communications (2023)
-
Phosphorylation of the PA subunit of influenza polymerase at Y393 prevents binding of the 5′-termini of RNA and polymerase function
Scientific Reports (2023)
-
An Influenza A virus can evolve to use human ANP32E through altering polymerase dimerization
Nature Communications (2023)
-
An intermediate state allows influenza polymerase to switch smoothly between transcription and replication cycles
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.