Abstract
Cultured pluripotent stem cells are a cornerstone of regenerative medicine owing to their ability to give rise to all cell types of the body. Although pluripotent stem cells can be propagated indefinitely in vitro, pluripotency is paradoxically a transient state in vivo, lasting 2–3 days around the time of blastocyst implantation1. The exception to this rule is embryonic diapause, a reversible state of suspended development triggered by unfavourable conditions2. Diapause is a physiological reproductive strategy widely employed across the animal kingdom, including in mammals, but its regulation remains poorly understood. Here we report that the partial inhibition of mechanistic target of rapamycin (mTOR), a major nutrient sensor and promoter of growth3, induces reversible pausing of mouse blastocyst development and allows their prolonged culture ex vivo. Paused blastocysts remain pluripotent and competent—able to give rise to embryonic stem (ES) cells and live, fertile mice. We show that both naturally diapaused blastocysts in vivo and paused blastocysts ex vivo display pronounced reductions in mTOR activity, translation, histone modifications associated with gene activity and transcription. Pausing can be induced directly in cultured ES cells and sustained for weeks without appreciable cell death or deviations from cell cycle distributions. We show that paused ES cells display a remarkable global suppression of transcription, maintain a gene expression signature of diapaused blastocysts and remain pluripotent. These results uncover a new pluripotent stem cell state corresponding to the epiblast of the diapaused blastocyst and indicate that mTOR regulates developmental timing at the peri-implantation stage. Our findings have implications in the fields of assisted reproduction, regenerative medicine, cancer, metabolic disorders and ageing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009)
Fenelon, J. C., Banerjee, A. & Murphy, B. D. Embryonic diapause: development on hold. Int. J. Dev. Biol. 58, 163–174 (2014)
Zoncu, R., Efeyan, A. & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nat. Rev. Mol. Cell Biol. 12, 21–35 (2011)
Naeslund, G. The effect of glucose-, arginine- and leucine-deprivation on mouse blastocyst outgrowth in vitro. Ups. J. Med. Sci. 84, 9–20 (1979)
Gwatkin, R. B. L. Amino acid requirements for attachment and outgrowth of the mouse blastocyst in vitro. J. Cell. Physiol. 68, 335–343 (1966)
Scognamiglio, R. et al. Myc depletion induces a pluripotent dormant state mimicking diapause. Cell 164, 668–680 (2016)
Hsieh, A. C. et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 485, 55–61 (2012)
Rodrik-Outmezguine, V. S. et al. Overcoming mTOR resistance mutations with a new-generation mTOR inhibitor. Nature 534, 272–276 (2016)
MacLean Hunter, S. & Evans, M. Non-surgical method for the induction of delayed implantation and recovery of viable blastocysts in rats and mice by the use of tamoxifen and Depo-Provera. Mol. Reprod. Dev. 52, 29–32 (1999)
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981)
Fu, Z. et al. Integral proteomic analysis of blastocysts reveals key molecular machinery governing embryonic diapause and reactivation for implantation in mice. Biol. Reprod. 90, 52–52 (2014)
Chan, E. Y. mTORC1 phosphorylates the ULK1-mAtg13-FIP200 autophagy regulatory complex. Sci. Signal. 2, pe51–pe51 (2009)
Tsukamoto, S. et al. Autophagy is essential for preimplantation development of mouse embryos. Science 321, 117–120 (2008)
Lee, J.-E. et al. Autophagy regulates embryonic survival during delayed implantation. Endocrinology 152, 2067–2075 (2011)
Nie, Z. et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 151, 68–79 (2012)
Lin, C. Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012)
Martin, P. M. & Sutherland, A. E. Exogenous amino acids regulate trophectoderm differentiation in the mouse blastocyst through an mTOR-dependent pathway. Dev. Biol. 240, 182–193 (2001)
Blaschke, K. et al. Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 500, 222–226 (2013)
Ying, Q.-L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008)
Boroviak, T. et al. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis. Dev. Cell 35, 366–382 (2015)
Murakami, M. et al. mTOR is essential for growth and proliferation in early mouse embryos and embryonic stem cells. Mol. Cell. Biol. 24, 6710–6718 (2004)
Guertin, D. A. et al. Ablation in mice of the mTORC components raptor, rictor, or mLST8 reveals that mTORC2 is required for signaling to Akt-FOXO and PKCalpha, but not S6K1. Dev. Cell 11, 859–871 (2006)
Gangloff, Y.-G. et al. Disruption of the mouse mTOR gene leads to early postimplantation lethality and prohibits embryonic stem cell development. Mol. Cell. Biol. 24, 9508–9516 (2004)
Schiesari, L. & O’Connor, M. B. Diapause: delaying the developmental clock in response to a changing environment. Curr. Top. Dev. Biol. 105, 213–246 (2013)
Yeom, Y. I. et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 122, 881–894 (1996)
Behringer, R., Gertsenstein, M., Nagy, K. V. & Nagy, A. Manipulating the Mouse Embryo. (2013)
Lee, K.-H., Chuang, C.-K., Guo, S.-F. & Tu, C.-F. Simple and efficient derivation of mouse embryonic stem cell lines using differentiation inhibitors or proliferation stimulators. Stem Cells Dev. 21, 373–383 (2012)
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006)
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl Acad. Sci. USA 90, 8424–8428 (1993)
George, S. H. L. et al. Developmental and adult phenotyping directly from mutant embryonic stem cells. Proc. Natl Acad. Sci. USA 104, 4455–4460 (2007)
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013)
External RNA Controls Consortium. Proposed methods for testing and selecting the ERCC external RNA controls. BMC Genomics 6, 150 (2005)
Lazar, C. et al. Batch effect removal methods for microarray gene expression data integration: a survey. Brief. Bioinform. 14, 469–490 (2013)
Gene Ontology Consortium. Gene Ontology Consortium: going forward. Nucleic Acids Res. 43, D1049–D1056 (2015)
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000)
Climent, M. et al. Functional analysis of Rex1 during preimplantation development. Stem Cells Dev. 22, 459–472 (2013)
Aziz, M. & Alexandre, H. The origin of the nascent blastocoele in preimplantation mouse embryos ultrastructural cytochemistry and effect of chloroquine. Rouxs Arch. Dev. Biol. 200, 77–85 (1991)
Acknowledgements
We are grateful to K. Shokat for the kind gift of RapaLink-1. We thank R. Blelloch, B. Bruneau, M. Conti, S. Fisher, D. Ruggero, and members of the Santos Laboratory for critical reading of the manuscript. This research was supported by grants NIH 5P30CA082103 to the UCSF Center for Advanced Technology, NIH P30DK063720 to the UCSF Flow Cytometry Core, NSF 1442504 and NIH R01CA163336 to J.S.S., and NIH R01OD012204 and R01GM113014 to M.R.-S.
Author information
Authors and Affiliations
Contributions
A.B.-K., S.B. and M.R.-S. conceived of the project. A.B.-K. and S.B. isolated embryos. A.B.-K. performed most paused embryo and ES cell cultures, embryo stainings and quantifications, and ES cell experiments with the following exceptions: T.A.M. performed HPG, EU and cell cycle analyses on ES cells. S.B. generated diapaused embryos, performed all NSET embryo transfers, and parturitions when necessary. H.J., and M.H. analysed RNA-seq data under the supervision of J.S.S. M.G. performed parallel embryo culture for surgical transfers and ES cell culture for aggregations, supported by staff of TCP Model Production Core. M.R.-S. supervised the project. A.B.-K. and M.R.-S. wrote the manuscript with feedback from all authors.
Corresponding author
Additional information
Reviewer InformationNature thanks K. Kaji, T. Rodríguez and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Catalytic mTOR inhibitors can induce blastocyst pausing.
a, b, Kaplan–Meier survival curves of blastocysts cultured with two catalytic mTor inhibitors RapaLink-1 and AZD2014 (a) and two allosteric mTOR inhibitors rapamycin and everolimus (b) at different concentrations. c, Kaplan–Meier survival curves of blastocysts cultured with all 5 mTOR inhibitors used in this study. The concentrations yielding the best survival outcome are shown. d, Kaplan–Meier survival curves of blastocysts cultured with the Myc-inhibitor 10058-F4 at 55 μM, as in ref. 6. Dotted lines indicate the time of blastocyst flushing (E3.5).
Extended Data Figure 2 The ex vivo paused epiblast retains naive pluripotency markers and is devoid of apoptosis.
It is possible that the signs of apoptosis in the trophoectoderm contribute to the eventual demise of mTOR-inhibited blastocysts after prolonged culture. a–d, Immunofluorescence images of untreated versus paused blastocysts for Nanog (a), Rex1 (b), cleaved caspase 3 (c) and cleaved Parp1 (d). Oct4 staining is shown in all cases as a marker of the ICM. Note that the ubiquitous staining pattern for Rex1 is as expected36. Scale bar, 50 μm. The number of embryos analysed (n) is indicated.
Extended Data Figure 3 Only blastocyst-stage embryos can be sustainably paused and they can give rise to live, fertile mice.
a, Detailed information on the generation of live mice from paused blastocysts by transfer into pseudo-pregnant surrogate females. Conditions used and live birth events are indicated. NSET, non-surgical embryo transfer; ET, embryo transfer (surgical). b, Live pups born as a result of mating mice generated from paused blastocysts with wild-type ICR mice. All tested mice (5 out of 5, from a total of 15 mice generated from paused blastocysts) proved to be fertile. c, Representative images of ex vivo cultured 2- or 8-cell embryos with or without the mTOR inhibitor. Cleavage-stage embryos proceed to generate blastocysts even in presence of mTOR inhibitor. n represents number of blastocysts developed per number of tested cleavage stage embryos.
Extended Data Figure 4 Paused blastocysts show signs of autophagy.
a, b, Immunofluorescence images of phosphorylated Ulk1 in untreated versus paused (a), or normal versus diapaused (b) embryos. Oct4 staining is shown as a marker of the ICM. Scale bar, 50 μm. The number of embryos analysed (n) is indicated. Error bars indicate standard deviation. P values are from two-tailed unpaired Student’s t-test. c, Images showing embryos retrieved at different developmental stages incubated with the autophagy/lysosomal inhibitor chloroquine (CQ) for 24 h. The number of embryos analysed (n) is indicated. d, Kaplan–Meier survival curves of blastocysts cultured with chloroquine in the presence or absence of the mTOR-inhibitor INK128. CQ treatment leads to blastocysts collapse, as previously reported37. The experiment was terminated at EDG8.5. e, Kaplan–Meier survival curves of blastocysts cultured with the autophagy/Ulk1 inhibitor SBI-0206965 in the presence or absence of the mTOR inhibitor INK128. Inhibition of Ulk1 leads to decreased survival of paused blastocysts. **P < 0.01 from log-rank (Mantel–Cox) test. Dotted lines indicate the time of blastocyst flushing (E3.5).
Extended Data Figure 5 In vivo diapaused blastocysts have reduced histone H4 acetylation.
a–c, Immunofluorescence images and quantification at the single ICM cell level of normal versus in vivo diapaused blastocysts for H4K5/8/12ac (a), H3K4me3 (b) and H3K9me3 (c) levels. Oct4 staining is shown in all cases as a marker of the ICM. Scale bar, 50 μm. The number of embryos analysed (n) is indicated. Graphs show data pooled from all embryos. Error bars indicate standard deviation. P values are from two-tailed unpaired Student’s t-test.
Extended Data Figure 6 Partial inhibition of mTOR activity results in ES cell pausing.
a, Representative image showing morphology of ES cells paused in 2i medium, with removal of the mTOR inhibitor at passaging. Under these conditions, ES cells can be discontinuously paused in 2i medium. b, Western blot showing moderately reduced 4EBP1 and S6K1 phosphorylation, which are mediated by mTORC1, and abolished Akt phosphorylation, which is mediated by mTORC2, in paused ES cells. For gel source data, see Supplementary Fig. 1. c, Flow cytometry analysis of nascent translation in the three states measured by HPG incorporation in E14 cells, in triplicates. d, Analysis of cell diameter in the three states, with each data point representing a population average of at least 300 cells per measurement. Values are represented as mean ± s.d. P values are from two-tailed unpaired Student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001, NS, not significant. e, Live pups born as a result of mating highly chimeric males generated using paused G4 ES cells with wild-type CD-1(ICR) female mice.
Extended Data Figure 7 ERCC sequence abundances followed a highly linear trend in all pairs of samples across at least five orders of magnitude.
The absolute abundance of mRNA transcripts was estimated using the ERCC92 RNA spike-ins32. Sequencing reads were aligned to the 92 reference spike-in sequences and the abundance of these sequences between different samples was compared. ERCC sequence abundances followed a highly linear trend in all pairs of samples across at least 5 orders of magnitude (Pearson correlation coefficient larger than 99.7%). Diagonal entries in the figure matrix show the density of read counts. ERCC RNAs are consistently detected at higher abundance in the paused state compared to serum and 2i.
Extended Data Figure 8 Distinct functional annotations are associated with different states of ES cells.
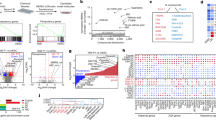
a, Clustering of dynamically expressed genes. Heat map shows 3,864 dynamically expressed genes (differentially expressed between any two of the 2i, serum, and paused states and mean FPKM >10 in at least one state). The FPKM value of each gene was standardized across the nine samples by subtracting the mean and then dividing by the standard deviation. Hierarchical clustering was performed using the standardized expression values using Euclidean metric and average linkage. b–d, Selected GO terms enriched in the annotations of genes upregulated in serum (b), 2i (c) or paused (d) ES cells. See Supplementary Table 3 for complete list of significant gene ontology terms associated with each ES cell state.
Extended Data Figure 9 Analysis of RNA-seq data indicates that paused ES cells mimic embryonic diapause.
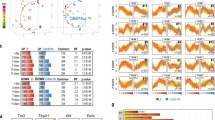
a, b, Scatter plots showing gene expression (9,418 genes that have FPKM >10 in any of our or Boroviak’s samples are shown) (a) and pathway expression (b) comparing paused ES cells to different developmental stages. Spearman correlation coefficients and P values indicate a significant similarity of paused ES cells to the diapaused epiblast. y axes represent log2 fold change in gene expression (a) or pathway expression (b) in paused versus serum ES cells. x axes represent log2 fold changes in gene expression (a) or pathway expression (b) in the different developmental stages (indicated above plots) versus E4.5 epiblast. Red boxes indicate the developmental stage that paused ES cells are closest to in each analysis. c, Scatter plot showing pathway expression in paused/serum ES cells (y axis) versus diapaused epiblast/E4.5 epiblast (x axis) as in (b). Pathways coordinately upregulated (red) or downregulated (blue) in paused ES cells and diapaused epiblast are indicated next to the scatter plot.
Extended Data Figure 10 Analysis of RNA-seq data indicates that Myc double knockout (DKO) cells have similarities to embryonic diapause at the GO term and pathway levels, but to a lesser extent than paused ES cells.
a–c, Scatter plots showing gene ontology term expression (a), gene expression (b), and pathway expression (c) comparing Myc DKO ES cells6 to different developmental stages, similarly to Fig. 4f and Extended Data Fig. 9a, b. Red boxes indicate the developmental stage that Myc DKO ES cells are closest to in each analysis. d, Scatter plots showing ‘diapause-driver’ gene expression (1,324 genes that are differentially expressed between the diapaused epiblast and E4.5 epiblast) in paused ES cells (left panel) and Myc DKO cells (right panel) compared to the diapaused epiblast. Pseudocount 1 was used when calculating the log2 fold changes. Spearman correlation coefficient and P value indicates a statistically significant similarity of paused ES cells, but not Myc DKO cells, to the diapaused epiblast. e, Scatter plot showing pathway expression in paused ES cells versus the diapaused epiblast, as in Extended Data Fig. 9c. Colour-coded expression levels of these pathways in Myc DKO ES cells are superimposed onto this graph. Numerical values for selected pathways are indicated. Upregulated pathways are in general concordant in paused and Myc DKO ES cells, whereas downregulated pathways are mostly discordant.
Supplementary information
Supplementary Figure 1
This file contains the uncropped western blot images with size marker indications. (PDF 299 kb)
Supplementary Table 1
This file shows the survival data for Extended Data Figure 1c. (XLSX 38 kb)
Supplementary Table 2
This file shows the Gene expression values from RNA-seq of serum, 2i and paused ES cells (this study), Boroviak et al. samples and Scognamiglio et al. samples. (XLSX 10693 kb)
Supplementary Table 3
This file contains the Gene ontology analysis of genes upregulated in serum, 2i or paused ES cells. (XLSX 169 kb)
Rights and permissions
About this article
Cite this article
Bulut-Karslioglu, A., Biechele, S., Jin, H. et al. Inhibition of mTOR induces a paused pluripotent state. Nature 540, 119–123 (2016). https://doi.org/10.1038/nature20578
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20578
This article is cited by
-
A transcriptomic examination of encased rotifer embryos reveals the developmental trajectory leading to long-term dormancy; are they “animal seeds”?
BMC Genomics (2024)
-
FOXO1-mediated lipid metabolism maintains mammalian embryos in dormancy
Nature Cell Biology (2024)
-
Quiescence enables unrestricted cell fate in naive embryonic stem cells
Nature Communications (2024)
-
Temperature-induced embryonic diapause in chickens is mediated by PKC-NF-κB-IRF1 signaling
BMC Biology (2023)
-
Ageing as a software design flaw
Genome Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.