Abstract
Intermediary metabolism generates substrates for chromatin modification, enabling the potential coupling of metabolic and epigenetic states. Here we identify a network linking metabolic and epigenetic alterations that is central to oncogenic transformation downstream of the liver kinase B1 (LKB1, also known as STK11) tumour suppressor, an integrator of nutrient availability, metabolism and growth. By developing genetically engineered mouse models and primary pancreatic epithelial cells, and employing transcriptional, proteomics, and metabolic analyses, we find that oncogenic cooperation between LKB1 loss and KRAS activation is fuelled by pronounced mTOR-dependent induction of the serine–glycine–one-carbon pathway coupled to S-adenosylmethionine generation. At the same time, DNA methyltransferases are upregulated, leading to elevation in DNA methylation with particular enrichment at retrotransposon elements associated with their transcriptional silencing. Correspondingly, LKB1 deficiency sensitizes cells and tumours to inhibition of serine biosynthesis and DNA methylation. Thus, we define a hypermetabolic state that incites changes in the epigenetic landscape to support tumorigenic growth of LKB1-mutant cells, while resulting in potential therapeutic vulnerabilities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gut, P. & Verdin, E. The nexus of chromatin regulation and intermediary metabolism. Nature 502, 489–498 (2013)
Etchegaray, J.-P. & Mostoslavsky, R. Interplay between metabolism and epigenetics: a nuclear adaptation to environmental changes. Mol. Cell 62, 695–711 (2016)
Carey, B. W., Finley, L. W. S., Cross, J. R., Allis, C. D. & Thompson, C. B. Intracellular α-ketoglutarate maintains the pluripotency of embryonic stem cells. Nature 518, 413–416 (2015)
Shyh-Chang, N. et al. Influence of threonine metabolism on S-adenosylmethionine and histone methylation. Science 339, 222–226 (2013)
Losman, J.-A. & Kaelin, W. G. Jr. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev . 27, 836–852 (2013)
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015)
Witkiewicz, A. K. et al. Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat. Commun. 6, 6744 (2015)
Su, G. H. et al. Germline and somatic mutations of the STK11/LKB1 Peutz-Jeghers gene in pancreatic and biliary cancers. Am. J. Pathol. 154, 1835–1840 (1999)
Giardiello, F. M. et al. Very high risk of cancer in familial Peutz-Jeghers syndrome. Gastroenterology 119, 1447–1453 (2000)
Korsse, S. E. et al. Pancreatic cancer risk in Peutz-Jeghers syndrome patients: a large cohort study and implications for surveillance. J. Med. Genet . 50, 59–64 (2013)
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007)
Chen, Z. et al. A murine lung cancer co-clinical trial identifies genetic modifiers of therapeutic response. Nature 483, 613–617 (2012)
Wingo, S. N. et al. Somatic LKB1 mutations promote cervical cancer progression. PLoS One 4, e5137 (2009)
Liu, W. et al. LKB1/STK11 inactivation leads to expansion of a prometastatic tumor subpopulation in melanoma. Cancer Cell 21, 751–764 (2012)
Shackelford, D. B. & Shaw, R. J. The LKB1-AMPK pathway: metabolism and growth control in tumour suppression. Nat. Rev. Cancer 9, 563–575 (2009)
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014)
Ting, L., Rad, R., Gygi, S. P. & Haas, W. MS3 eliminates ratio distortion in isobaric multiplexed quantitative proteomics. Nat. Methods 8, 937–940 (2011)
Mehrmohamadi, M., Liu, X., Shestov, A. A. & Locasale, J. W. Characterization of the usage of the serine metabolic network in human cancer. Cell Reports 9, 1507–1519 (2014)
Bardeesy, N. et al. Both p16(Ink4a) and the p19(Arf)-p53 pathway constrain progression of pancreatic adenocarcinoma in the mouse. Proc. Natl Acad. Sci. USA 103, 5947–5952 (2006)
Locasale, J. W. Serine, glycine and one-carbon units: cancer metabolism in full circle. Nat. Rev. Cancer 13, 572–583 (2013)
Mentch, S. J. et al. Histone methylation dynamics and gene regulation occur through the sensing of one-carbon metabolism. Cell Metab . 22, 861–873 (2015).
Maddocks, O. D. K., Labuschagne, C. F., Adams, P. D. & Vousden, K. H. Serine metabolism supports the methionine cycle and DNA/RNA methylation through de novo ATP synthesis in cancer cells. Mol. Cell 61, 210–221 (2016)
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011)
Schübeler, D. Function and information content of DNA methylation. Nature 517, 321–326 (2015)
Elbarbary, R. A., Lucas, B. A. & Maquat, L. E. Retrotransposons as regulators of gene expression. Science http://dx.doi.org/10.1126/science.aac7247 (2016)
Kulis, M., Queirós, A. C., Beekman, R. & Martín-Subero, J. I. Intragenic DNA methylation in transcriptional regulation, normal differentiation and cancer. Biochim. Biophys. Acta 1829, 1161–1174 (2013)
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013)
Lee, J. V. et al. Akt-dependent metabolic reprogramming regulates tumor cell histone acetylation. Cell Metab . 20, 306–319 (2014)
Shim, E.-H. et al. L-2-Hydroxyglutarate: an epigenetic modifier and putative oncometabolite in renal cancer. Cancer Discov . 4, 1290–1298 (2014)
Intlekofer, A. M. et al. Hypoxia induces production of L-2-hydroxyglutarate. Cell Metab . 22, 304–311 (2015)
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961–973 (2015)
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015)
Treppendahl, M. B., Kristensen, L. S. & Grønbæk, K. Predicting response to epigenetic therapy. J. Clin. Invest. 124, 47–55 (2014)
Locasale, J. W. et al. Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nat. Genet . 43, 869–874 (2011)
DeNicola, G. M. et al. NRF2 regulates serine biosynthesis in non-small cell lung cancer. Nat. Genet . 47, 1475–1481 (2015)
Sun, L. et al. cMyc-mediated activation of serine biosynthesis pathway is critical for cancer progression under nutrient deprivation conditions. Cell Res . 25, 429–444 (2015)
Ye, J. et al. Serine catabolism regulates mitochondrial redox control during hypoxia. Cancer Discov . 4, 1406–1417 (2014)
Zhang, W. C. et al. Glycine decarboxylase activity drives non-small cell lung cancer tumor-initiating cells and tumorigenesis. Cell 148, 259–272 (2012)
Ben-Sahra, I., Howell, J. J., Asara, J. M. & Manning, B. D. Stimulation of de novo pyrimidine synthesis by growth signaling through mTOR and S6K1. Science 339, 1323–1328 (2013)
Acknowledgements
We thank A. Kimmelman, K. Patra, L. J. Etchegaray, and R. Mostoslavsky for comments on the manuscript, and P. Foltopoulou, B. Martinez and Bardeesy laboratory members for advice. N.B. holds the Gallagher Endowed Chair in Gastrointestinal Cancer Research and received support from the Granara-Skerry Trust, the Linda J. Verville Foundation and the Begg Family, and grants from the NIH (P01 CA117969-07, R01 CA133557-05). F.K. is supported by a Hirshberg Foundation Career Development Award. F.K. and N.B. were supported by NIH grant P50CA1270003 and are members of the Andrew Warshaw Institute.
Author information
Authors and Affiliations
Contributions
F.K. and N.B. conceived and designed the study. F.K., A.R. and J.M.N. performed cell-based and mouse experiments. T.C. assisted with mouse experiments. F.K. and B.N.N. performed and interpreted the tracing experiments. M.B and W.H. performed proteomics. F.K. and M.L. performed the OCR measurements. M.C.H. and D.N.H. provided essential samples and data analysis. Y.Y.L. performed computational analysis. H.G. prepared WGBS libraries. R.K. and A.M. analysed and interpreted the WGBS data. P.S.H., K.K.W., O.S.S. and N.J.D. assisted with data interpretation. F.K. and N.B. wrote the manuscript with feedback from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks M. Rehli and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 LKB1 suppresses KRASG12D-driven tumorigenesis and limits glycolysis in primary pancreatic ductal epithelial cells.
a, Schematic of GEM models. Sox9-CreER, LKB1L/L, and LSL-KRASG12D/+ mice were crossed to generate four cohorts: WT (Sox9-CreER), L (Sox9-CreER;LKB1L/L), K (Sox9-CreER;LSL-KRASG12D/+) and KL (Sox9-CreER;LSL-KRASG12D/+;LKB1L/L). Genetic lesions were induced by intraperitoneal injections of tamoxifen at 6 weeks of age, after which mice were observed for signs of disease and killed when KL animals were moribund (20–25 weeks of age). The WT, K, and L mice had no signs of illness or other abnormalities at this time point. b, Haematoxylin and eosin-stained sections of representative pancreata from WT and L mice (n = 4 mice per group). Scale bars, 50 μm. c, Schematic of primary pancreatic ductal epithelial cell system. Pancreatic ductal epithelial cells were isolated from LSL-KRASG12D/+ and LSL-KRASG12D/+;LKB1L/L mice and infected with Adeno-Cre to generate K and KL cells. For studies comparing K, L, KL and WT genotypes, cells were isolated from FSF-KRASG12D/+;LKB1L/L mice and infected with Adeno-Flipase and/or Adeno-Cre, or neither. d, Volume and e, weight of subcutaneous tumours derived from ductal cells of the indicated genotypes (n = 4 tumours per group). Error bars show s.e.m. For source data on tumour volume, see Supplementary Data Table 4. f, Weight of subcutaneous tumours from K (n = 6), KL (n = 8) and KL cells transduced with retrovirus expressing LKB1 cDNA (rescue, n = 8). g, Number of colonies formed in soft agar by K (none detected) or KL cells (n = 6 independent biological replicates). Error bars show s.e.m. h, Proliferation of K and KL cells in nutrient-replete medium (n = 4). i, Proliferation of KL cells transduced with retroviruses expressing empty vector or LKB1 (rescue; n = 3). j, Proliferation of wild-type (WT), KRASG12D/+ (K), LKB1−/− (L) and KRASG12D/+;LKB1−/− (KL) cells (n = 6). k–t, In vitro studies of K and KL cells. k, Detection of GLUT1 (SLC2A1) by immunofluorescence (scale bar, 20 μm) or immunoblot. 2-(4-amidinophenyl)-1H-indole-6-carboxamidine (DAPI) was used to visualize nuclei. Actin was used as the loading control. For gel source images see Supplementary Data Fig. 1. l, Steady-state ATP levels under nutrient-replete conditions measured by CellTiterGlo (Promega), normalized to cell number and expressed as relative to ATP levels in K cells (n = 4). m, Intracellular levels of pyruvate, lactate, and TCA cycle metabolites as detected by GC–MS. Values are normalized to cell number. Data are expressed as relative to the levels in K cells (n = 6 biological replicates). n, Three-day proliferation of cells in 25 mM glucose or acutely switched to media with the indicated reduced glucose concentrations (n = 6). Data are expressed as relative to day 0. o–q, Proliferation of cells treated with 5 mM 2-deoxyglucose (2DG) (o), 5 mM dichloroacetate (DCA) (p) or 20 μM galloflavin (gallo) (q). Values are expressed as percentage of normal growth (2DG n = 6, DCA n = 8, gallo n = 6). r–t, WT, K, L and KL cells were measured for glucose uptake using 2NBDG (r, n = 6), lactate release into the medium (s, n = 4), and expression of glycolytic genes (t, n = 6). Data are pooled from two (j, l, n–r) or three (t) experiments or representative of two (h, i, s) experiments. For all panels, error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 2 LKB1 loss induces the serine–glycine–one-carbon network.
a, GSEA showing enrichment of proteins involved in the serine–glycine–one-carbon network18 in KL cells compared to K cells using global proteomics (n = 4 samples per group). b, Plot of isotopomer abundance of U[13C]glucose-derived intracellular M+3 lactate over time (n = 3 independent biological replicates). c, Plot of isotopomer abundance of U[13C]glucose-derived intracellular M+2 glycine over time (n = 3 independent biological replicates). d, Intracellular levels of serine, glycine and glutamine as detected by GC–MS. Values normalized to cell number. Data are expressed as relative to the levels in K cells (n = 6 biological replicates). e, PSAT1 and GLDC expression determined by immunoblot in K and KL cells and in KL cells transduced with LKB1 cDNA. Actin was used as loading control. For gel source images see Supplementary Data Fig. 1. f, GSEA of RNA-seq data showing suppression of genes involved in glycolysis, serine biosynthesis, folate cycle and the serine–glycine–one-carbon network upon re-expression of LKB1 cDNA in KL cells (rescue, n = 2 samples) compared to parental KL cells (n = 4 samples). g, Plot of isotopomer abundance of U[13C]glucose-derived intracellular M+3 serine 6 h after addition of U[13C]glucose (n = 3 independent biological replicates). For all panels, error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 3 The de novo serine biosynthesis pathway is required for KL but not KPC tumour growth.
a, Proliferation of KL cells transduced with vector or LKB1 cDNA and cultured in the presence or absence of 0.4 mM serine. Growth is expressed as relative to day 0 (n = 3). LKB1 re-expression slows growth and results in sensitivity to serine deprivation. b, qRT–PCR showing effective knockdown of PSAT1 in K, KL, KPC and KIC cells transduced with shControl or two different shRNAs against PSAT1 (n = 2). Data are expressed as relative to shControl for each cell line. 18S rRNA was used for normalization. c, Number of colonies formed in soft agar by KL cells transduced with shControl or two different shRNAs against PSAT1 (n = 6). d, Proliferation of K or KL cells transduced with shControl or two independent shRNAs against PSAT1 in the absence of serine. Data are expressed as percentage of growth in the presence of 0.4 mM serine (n = 6). e, qRT–PCR showing effective knockdown of endogenous mouse PSAT1 (mPSAT1, endogenous) and forced expression of human PSAT1 (hPSAT1, exogenous) in KL cells transduced with shControl, shPSAT1-1, shPSAT1-2, vector or hPSAT1 cDNA. Data are normalized to 18s rRNA (n = 2). f, Weight at the time of harvesting of subcutaneous tumours from KL cells transduced with shControl (n = 8), shPSAT1-1 (n = 12), or shSPAT1-2 (n = 12). Error bars show s.e.m. g, Haematoxylin and eosin-stained sections and immunofluorescence analysis of representative tumours derived from subcutaneous injections of KL cells transduced with shControl, shPSAT1-1, or shPSAT1-2. Note that PSAT1 knockdown tumours have a reduction in malignant glands (arrows) relative to the fibrotic stroma. Lower panels: anti-CK19 (green) was used to visualize the neoplastic epithelium and anti-PCNA (red) was used to mark proliferation. DAPI was used to stain nuclei (blue). h, Proliferation of KPC and KIC pancreatic cancer cells transduced with shControl or two independent shRNAs against PSAT1. Growth is expressed as relative to day 0 (n = 6 for KPC and n = 4 for KIC). i–k, KPC cells were transduced with shControl (n = 6), shPSAT1-1 (n = 6) or shSPAT1-2 (n = 4) and injected into SCID mice. i, Volume (left) and weight (right) of subcutaneous tumours. Error bars show s.e.m. For source data on tumour volume, see Supplementary Data Table 4. j, k, Tumours in i were stained using anti-CK19 antibody (green) to visualize the neoplastic epithelium and anti-PCNA staining (red) to mark proliferating cells. DAPI was used to stain nuclei (blue). The proportion of stained CK19+ cells is quantified in j (n = 4, representative tumours) and CK19+ cells with nuclear PCNA staining are quantified in j. There are no significant effects on any of these parameters. k, Haematoxylin and eosin-stained sections (top) and immunofluorescence analysis (bottom) of representative tumours. Scale bars, 100 μm. Insets show threefold magnification. Data pooled from two (c, d) or representative of two (a, b, e) or three (h) experiments. For all panels, error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 4 Characterization of serine–glycine–one-carbon pathway in KL cells.
a, Detailed graph of SGOC network. Enzymatic inhibitors used in this study are marked in red. b, ROS in K or KL cells transduced with shControl, shPSAT1-1 or shPSAT1-2 measured by DCFDA (left) and CellRox staining (right). Data are normalized to cell number (n = 4). c, Six-day proliferation assay of KL cells transduced with shControl, shPSAT1-1 or shPSAT1-2, showing lack of growth rescue by N-acetylcysteine (NAC). Data are expressed as relative to day 0 (n = 6). d, Three-day growth assay of KL cells transduced with shControl, shPSAT1-1 or shPSAT1-2, showing the lack of rescue by excess nucleosides (adenosine, guanosine, thymidine, uridine, cytidine; 1 mM each). Data are presented as percentage of the growth of shControl cells (n = 16). e, Proliferation of K or KL cells treated with aminooxyacetate (AOA). Data are expressed as relative to day 0 (n = 8). f, Five-day proliferation of K or KL cells treated with AOA and/or NAC. Data are expressed as relative to day 0 (n = 6). g, Data from RNA-seq (left) and quantitative proteomics (right) showing levels of genes involved in the production of SAM in K and KL. The data plotted are expressed as mean-centred values. h, Proliferation of K or KL cells treated with 2 mM cycloleucine. Data are expressed as percentage of the growth of vehicle treated cells (n = 12). Data are pooled from two (b–f, h) experiments. For all panels, error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 5 Deletion of LKB1 induces DNMT1 and DNMT3A expression and increases global DNA methylation.
a, Heat map of RNA-seq data showing levels of the differentially regulated SAM-using enzymes. Plotted data are expressed as mean centred values. b, Expression of DNMT1 and DNMT3A in K, KL, and KL cells transduced with LKB1 cDNA (rescue) was measured by qRT–PCR. Levels were normalized to 18S rRNA. Data are expressed as relative to K cells (n = 4, representative of two experiments). c, Immunoblots of lysates from K, KL or rescue cells were probed for DNMT1 or DNMT3A. Actin was used as loading control. For gel source images see Supplementary Data Fig. 1. d, Measurement of SAM in K and KL cells treated with 5-aza-2-deoxycytidine (decitabine) or RG108 for 3 days. In each case, data are expressed as relative to the amount of SAM in vehicle-treated K cells, which was arbitrarily set to 1 (n = 6 independent replicates). e, Immunofluorescence staining and quantitation of 5mC in K or KL cells (77–130 cells). Scale bar, 25 μm. f, Dot blot of DNA isolated from K or KL cells probed with anti-5mC antibody. Quantified signal was normalized to total DNA as measured by methylene blue staining (n = 4, independent replicates). g, Dot blot of DNA isolated from KL cells transduced with empty vector or LKB1 cDNA probed with anti-5mC antibody. Quantified signal was normalized to total DNA as measured by methylene blue staining (n = 3, independent replicates). h, Immunoblot analysis of histone 3 (H3) methyl marks from K or KL cells. Data are normalized to total H3 (K4me3, n = 2; K27me3, n = 5; K36me3, n = 5, independent replicates). For gel source images see Supplementary Data Fig. 1. i, Dot blot of DNA isolated from K or KL cells probed with anti-5hmC antibody. Quantified signal was normalized to total DNA as measured by methylene blue staining (n = 4, independent replicates). For all panels, error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 6 Serine pathway activity sustains DNA methylation in KL cells.
a, Dot blot of DNA isolated from K or KL cells transduced with shControl, shPSAT1-1 or shPSAT1-2 probed with anti-5mC antibody. Total DNA was visualized by methylene blue staining. Graph shows quantified signal normalized to total DNA as measured by methylene blue staining (K cells, n = 4; KL cells, n = 8, independent replicates). b, Dot blot of DNA isolated from KPC cells transduced with shControl, shPSAT1-1 or shPSAT1-2 probed with anti-5mC antibody. The graph shows quantified signal normalized to total DNA (n = 4, independent replicates). c, Dot blot of DNA probed with anti-5mC antibody. DNA was isolated from KL cells first transduced with vector or human PSAT1 then transduced with shControl, shPSAT1-1 or shPSAT-2. Graph shows quantified signal normalized to total DNA as measured by methylene blue staining (n = 3, independent replicates). d, Expression of serine pathway genes and DNMT genes in WT, K, L or KL cells by qRT–PCR. Data are normalized to 18S and expressed as relative to K cells (n = 6, pooled data from two experiments). e, Plots of isotopomer abundance of U[13C]glucose-derived M+3 serine and M+2 glycine, 6 h after addition of U[13C]glucose (n = 6 independent biological replicates). f, Quantified DNA dot blot signal of DNA isolated from WT, K, L or KL cells probed with anti-5mC antibody normalized to total DNA as measured by methylene blue staining (n = 4, independent replicates). g, Five-day growth of WT, K, L or KL cells transduced with shControl or two shRNAs against PSAT1. Data are expressed as relative to day 0 (n = 4, pooled from two experiments). For all panels, error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 7 LKB1-mediated regulation of glycolysis–SGOC–DNMT pathway involves the AMKP–mTOR axis.
a–e, KL cells were transduced with vector, wild-type LKB1 or kinase-dead LKB1. a, Proliferation, expressed as relative to day 0 (n = 4). b, Glucose uptake measured using 2NBDG followed by fluorimetry (data normalized to cell number and expressed as relative to KL-vector cells (n = 8)). c, Lactate levels measured by fluorimetry 3 h after medium change, normalized to cell number (n = 8). d, Oxygen consumption rates measured in normal duct medium, followed by injections with 4 μM oligomycin (O), 4 μM FCCP (F), or 4 μM antimycin A (A) (n = 3). e, Expression of the indicated genes measured by qRT–PCR (n = 4), with levels normalized to 18S rRNA. Data are expressed as relative to KL-empty vector (EV) cells. f, Immunoblot of K, KL or KL cells transduced with LKB1 cDNA (rescue). For gel source images see Supplementary Data Fig. 1. g, Immunoblot of K cells treated overnight with 10 μΜ Compound C. For gel source images see Supplementary Data Fig. 1. h, i, Glucose uptake (h) and 3 h lactate production (i) in the same cells as in h. Data are normalized to cell number and expressed as relative to control K cells (glucose uptake, n = 3; lactate, n = 4). j, Glucose uptake in K or KL cells treated with vehicle or 10 μΜ Compound C. Data are normalized to cell number and expressed as relative to control K cells (n = 4). Note the blunted response of KL cells. k, qRT–PCR analysis of the indicated genes in the same cells as in h. Levels are normalized to 18S rRNA. Data are expressed as relative to control K cells (n = 6). l, Immunostaining for 5-mC (left) and quantification of staining (right) in K cells treated with vehicle or Compound C for 4 days (158–163 cells). m–q, K cells were transduced with shControl or shRNAs against AMPKa1 or AMPKa2. m, Glucose uptake. Data are normalized to cell number and expressed as relative to shControl-treated cells (n = 4). n, Steady-state ATP levels measured with CellTiterGlo (Promega), normalized to cell number and expressed as relative to ATP levels in shControl-treated cells (n = 4). o, Lactate levels measured by fluorimetry 3 h after medium change. Data are normalized to cell number (n = 3). p, Immunostaining for 5-mC (left) and quantification of staining (right) (159–296 cells). q, Expression of the indicated genes determined by qRT–PCR. Levels are normalized to 18S rRNA. Data are expressed as relative to shControl cells (n = 4). r–u, KL cells were transduced with vector or an LKB1 cDNA and shControl or shRNAs against AMPKa1 or AMPKa2. r, Glucose uptake. Data are normalized to cell number and expressed as relative to vector-shControl cells (n = 4). s, Steady-state ATP levels normalized to cell number and expressed as relative to ATP levels in EV-shControl cells (n = 4). t, Lactate levels 3 h after medium change. Data are normalized to cell number (n = 4). u, Proliferation expressed as relative to day 0 (n = 3). v, Impact of Torin 1 treatment on growth of K and KL cells (n = 4). w, Immunoblot of K and KL cells treated with vehicle or 25 nM Torin 1. For gel source images see Supplementary Data Fig. 1. x, Glucose uptake and 3 h lactate production in K or KL cells treated overnight with 25 nM Torin 1. Data are normalized to cell number (n = 4). y, Isotopomer abundance of U[13C]glucose-derived serine and glycine in the same cells as in r (n = 3 independent replicates). Cells were labelled with U[13C]glucose for 6 h. z, Expression of the indicated genes in Torin-treated K (left) or KL cells (right) determined by qRT–PCR. Levels are normalized to 18S rRNA. Data are expressed as relative to control-treated cells (n = 4). Data are pooled from two (b, c, e, i–n, p–t, v, x, z) or representative of two (a, d, h, o) or three (u) experiments. Error bars show s.e.m. in d, s.d. in all other panels. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 8 Loss of LKB1 increases DNA methylation in retrotransposon elements.
a, Hierarchical clustering of K and KL samples based on methylation levels at 100-bp autosomal genomic tiles as measured by whole-genome bisulfite sequencing. b, Two-dimensional density plot for methylation at non-repetitive 100-bp autosomal genomic tiles in KL versus K cells. The yellow line shows the mean methylation of tiles. The dots represent differentially methylated tiles (FDR q-value < 0.05, methylation change > 0.1). There are 3,395 hypermethylated tiles and 1,270 hypomethylated tiles in KL cells versus K cells. c, Overlap of genes associated with differentially methylated regions and differentially regulated genes (hyper DMRs, hypermethylated regions; hypo DMRs, hypomethylated regions; DRGs, differentially regulated genes). d–j, Distribution of methylation density within the low-CpG-promoters (LCPs), high-CpG-promoters (HCPs), islands, shores, exons, introns and LTRs. Numbers reflect median values. k, Average percentile change in methylation in the same elements as in d–j as well as LINEs and LTRs. l, Number of genes containing retrotransposon repeat elements (sum of LINEs, SINEs, LTRs) among the set of differentially regulated genes (DRGs) in K versus KL cells and among all genes. Note that the DRGs are enriched for the presence of retrotransposons in their gene bodies (top), but not in their promoters (middle) or 3′UTRs (bottom). m, Specific enrichment of LINE, SINE and LTR elements in the gene bodies of DRGs when compared to all genes in the genome.
Extended Data Figure 9 LKB1 deficiency confers hypersensitivity to inhibitors of DNA methylation.
a–c, Proliferation of K (a), KL (b) and KPC (c) cells transduced with shControl or two shRNAs against each of DNMT1 (D1) or DNMT3A (D3A). Data are expressed as relative to day 0 (K and KL n = 6, KPC n = 4). d–f, Volume of subcutaneous tumours derived from KPC cells transduced with doxcycline (Dox)-inducible shRNAs against DNMT1 or DNMT3A (n = 4). Doxcycline was introduced to the drinking water when tumours reached 125 mm3. Error bars show s.e.m. For source data on tumour volume, see Supplementary Data Table 4. g, Apoptosis measured by caspase 3/7 activity in K or KL cells treated with decitabine for 48 h. Values are normalized to cell number (n = 3). h, i, Proliferation of KL cells transduced with empty vector (h) or LKB1 cDNA (i) treated with decitabine. Data are expressed as relative to day 0 (n = 3). j, Proliferation of KL cells transduced with empty vector or LKB1 cDNA treated with RG108. Data are expressed as percentage of growth of untreated cells (n = 6). k–m, Proliferation of K or KL cells treated with RG108 (n = 6) (k), EGCG (n = 12) (l) or SGI1027 (n = 12) (m). Data are expressed as percentage of growth of untreated cells. n–q, Mice bearing subcutaneous KPC tumours were treated with decitabine (n = 12) or vehicle (n = 12) when tumours reached 125 mm3. n, o, Tumour volume (n) and final tumour weight (o). Error bars show s.e.m. For source data on tumour volume, see Supplementary Data Table 4. p, Haematoxylin and eosin-stained slides from representative tumours (top). Bottom, anti-CK19 (green) was used to visualize the neoplastic epithelium and anti-PCNA (red) was used to mark proliferating cells. DAPI was used to stain nuclei (blue). q, Quantification of the CK19+ neoplastic epithelial compartment (%CK19+ cells/total cells; top). Quantification of CK19+ cells with nuclear PCNA staining (bottom; n = 6). Scale bars, 100 μm. Insets are threefold magnification. Data pooled from two (a–c, j) or four (k–m) experiments or representative of two (h) or three (g) experiments. Error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 10 Vulnerabilities of human LKB1 mutant pancreatic cancer cell lines.
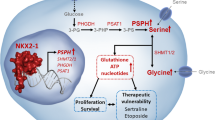
a, Three-day growth of LKB1 wild-type (black) or LKB1 mutant (red) human pancreatic cancer cells. Data are expressed as relative to shControl-transduced cells, which is arbitrarily set to 100 (n = 3). b, Quantification of tumour volume of subcutaneous tumours derived from implantation of the indicated cells transduced with shControl or two shRNAs against PSAT1 (n = 4). Error bars show s.e.m. For source data on tumour volume, see Supplementary Data Table 4. c, Immunofluorescence staining and quantification of 5mC in COLO357, PANC1 and PATU-8988T cells transduced with shControl or two shRNAs against PSAT1 (607–760 cells for COLO357, 305–342 cells for PANC1 and 623–889 cells for PATU-8988T). Scale bar, 25 μm. Error bars show s.e.m. Data are expressed as fluorescence per nucleus. d, Detection of 5mC by immunofluorescence in COLO357 (LKB1-deficient) cells transduced with vector or wild-type LKB1. Quantification is presented as fluorescence per nucleus (438–618 cells). Scale bar, 25 μm. Error bars show s.e.m. e, Three-day growth of LKB1 wild-type (black) or LKB1 mutant (red) human pancreatic cancer cells treated with 10 μM 3-deazaadenosine (3DZA) (top) or 2 mM cycloleucine (CycloLeu) (bottom). f, Metabolic and epigenetic changes promoting transformation upon deletion of the tumour suppressor LKB1. Data pooled from (c, d) or representative of (a) two experiments. Error bars show s.d. unless otherwise stated; *P < 0.05, **P < 0.01, ***P < 0.001.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1, western blot source data and Supplementary Methods. (PDF 846 kb)
Supplementary Data
This file contains Supplementary Table 1, Metabolic signatures (KEGG) regulated by LKB1. Metabolic KEGG genesets that are significantly enriched upon LKB1 deletion. GSEA was performed using both RNA-sequencing and proteomics data. (XLSX 12 kb)
Supplementary Data
This file contains Supplementary Table 2, a curated list of S-adenosyl-methionine utilizing enzymes. A List of 183 SAM-utilizing methyltransferases with expression values for K, KL and rescue cells (RNA) or K and KL cells (protein). (XLSX 89 kb)
Supplementary Data
This file contains Supplementary Table 3. Bisulfite conversion rates and sequencing statistics for WGBS. (XLSX 37 kb)
Supplementary Data
This file contains Supplementary Table 4, source data for tumour volumes in Extended Data Figures 1d, 3i, 9d, 9e, 9f, 9n, 10b. (XLSX 46 kb)
Rights and permissions
About this article
Cite this article
Kottakis, F., Nicolay, B., Roumane, A. et al. LKB1 loss links serine metabolism to DNA methylation and tumorigenesis. Nature 539, 390–395 (2016). https://doi.org/10.1038/nature20132
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20132
This article is cited by
-
Targeting serine/glycine metabolism improves radiotherapy response in non-small cell lung cancer
British Journal of Cancer (2024)
-
Decoding cancer insights: recent progress and strategies in proteomics for biomarker discovery
Journal of Proteins and Proteomics (2024)
-
Serine metabolism in macrophage polarization
Inflammation Research (2024)
-
The activation of mTOR signalling modulates DNA methylation by enhancing DNMT1 translation in hepatocellular carcinoma
Journal of Translational Medicine (2023)
-
Transcription factor NKX2–1 drives serine and glycine synthesis addiction in cancer
British Journal of Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.