Abstract
The basal ganglia, a group of subcortical nuclei, play a crucial role in decision-making by selecting actions and evaluating their outcomes1,2. While much is known about the function of the basal ganglia circuitry in selection1,3,4, how these nuclei contribute to outcome evaluation is less clear. Here we show that neurons in the habenula-projecting globus pallidus (GPh) in mice are essential for evaluating action outcomes and are regulated by a specific set of inputs from the basal ganglia. We find in a classical conditioning task that individual mouse GPh neurons bidirectionally encode whether an outcome is better or worse than expected. Mimicking these evaluation signals with optogenetic inhibition or excitation is sufficient to reinforce or discourage actions in a decision-making task. Moreover, cell-type-specific synaptic manipulations reveal that the inhibitory and excitatory inputs to the GPh are necessary for mice to appropriately evaluate positive and negative feedback, respectively. Finally, using rabies-virus-assisted monosynaptic tracing5, we show that the GPh is embedded in a basal ganglia circuit wherein it receives inhibitory input from both striosomal and matrix compartments of the striatum, and excitatory input from the ‘limbic’ regions of the subthalamic nucleus. Our results provide evidence that information about the selection and evaluation of actions is channelled through distinct sets of basal ganglia circuits, with the GPh representing a key locus in which information of opposing valence is integrated to determine whether action outcomes are better or worse than expected.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nelson, A. B. & Kreitzer, A. C. Reassessing models of basal ganglia function and dysfunction. Annu. Rev. Neurosci. 37, 117–135 (2014)
Amemori, K., Gibb, L. G. & Graybiel, A. M. Shifting responsibly: the importance of striatal modularity to reinforcement learning in uncertain environments. Front. Hum. Neurosci. 5, 47 (2011)
Hikosaka, O. Basal ganglia mechanisms of reward-oriented eye movement. Ann. NY Acad. Sci. 1104, 229–249 (2007)
Alexander, G. E. & Crutcher, M. D. Functional architecture of basal ganglia circuits: neural substrates of parallel processing. Trends Neurosci. 13, 266–271 (1990)
Callaway, E. M. & Luo, L. Monosynaptic circuit tracing with glycoprotein-deleted rabies viruses. J. Neurosci. 35, 8979–8985 (2015)
Stephenson-Jones, M., Kardamakis, A. A., Robertson, B. & Grillner, S. Independent circuits in the basal ganglia for the evaluation and selection of actions. Proc. Natl Acad. Sci. USA 110, E3670–E3679 (2013)
Hong, S. & Hikosaka, O. The globus pallidus sends reward-related signals to the lateral habenula. Neuron 60, 720–729 (2008)
Shabel, S. J., Proulx, C. D., Trias, A., Murphy, R. T. & Malinow, R. Input to the lateral habenula from the basal ganglia is excitatory, aversive, and suppressed by serotonin. Neuron 74, 475–481 (2012)
Matsumoto, M. & Hikosaka, O. Lateral habenula as a source of negative reward signals in dopamine neurons. Nature 447, 1111–1115 (2007)
Hong, S., Jhou, T. C., Smith, M., Saleem, K. S. & Hikosaka, O. Negative reward signals from the lateral habenula to dopamine neurons are mediated by rostromedial tegmental nucleus in primates. J. Neurosci. 31, 11457–11471 (2011)
Stamatakis, A. M. & Stuber, G. D. Activation of lateral habenula inputs to the ventral midbrain promotes behavioral avoidance. Nat. Neurosci. 15, 1105–1107 (2012)
Rajakumar, N., Elisevich, K. & Flumerfelt, B. A. Compartmental origin of the striato-entopeduncular projection in the rat. J. Comp. Neurol. 331, 286–296 (1993)
Parent, M., Lévesque, M. & Parent, A. Two types of projection neurons in the internal pallidum of primates: single-axon tracing and three-dimensional reconstruction. J. Comp. Neurol. 439, 162–175 (2001)
Vincent, S. R. & Brown, J. C. Somatostatin immunoreactivity in the entopeduncular projection to the lateral habenula in the rat. Neurosci. Lett. 68, 160–164 (1986)
Cohen, J. Y., Haesler, S., Vong, L., Lowell, B. B. & Uchida, N. Neuron-type-specific signals for reward and punishment in the ventral tegmental area. Nature 482, 85–88 (2012)
DeLong, M. R., Crutcher, M. D. & Georgopoulos, A. P. Primate globus pallidus and subthalamic nucleus: functional organization. J. Neurophysiol. 53, 530–543 (1985)
Bromberg-Martin, E. S., Matsumoto, M., Hong, S. & Hikosaka, O. A pallidus-habenula-dopamine pathway signals inferred stimulus values. J. Neurophysiol. 104, 1068–1076 (2010)
Pan, W. X., Schmidt, R., Wickens, J. R. & Hyland, B. I. Dopamine cells respond to predicted events during classical conditioning: evidence for eligibility traces in the reward-learning network. J. Neurosci. 25, 6235–6242 (2005)
Schultz, W. Dopamine reward prediction-error signalling: a two-component response. Nat. Rev. Neurosci. 17, 183–195 (2016)
Bru, T., Salinas, S. & Kremer, E. J. An update on canine adenovirus type 2 and its vectors. Viruses 2, 2134–2153 (2010)
Tai, L. H., Lee, A. M., Benavidez, N., Bonci, A. & Wilbrecht, L. Transient stimulation of distinct subpopulations of striatal neurons mimics changes in action value. Nat. Neurosci. 15, 1281–1289 (2012)
Ahrens, S. et al. ErbB4 regulation of a thalamic reticular nucleus circuit for sensory selection. Nat. Neurosci. 18, 104–111 (2015)
Wulff, P. et al. From synapse to behavior: rapid modulation of defined neuronal types with engineered GABAA receptors. Nat. Neurosci. 10, 923–929 (2007)
Fujiyama, F. et al. Exclusive and common targets of neostriatofugal projections of rat striosome neurons: a single neuron-tracing study using a viral vector. Eur. J. Neurosci. 33, 668–677 (2011)
Kita, H. & Kitai, S. T. Efferent projections of the subthalamic nucleus in the rat: light and electron microscopic analysis with the PHA-L method. J. Comp. Neurol. 260, 435–452 (1987)
Mastro, K. J., Bouchard, R. S., Holt, H. A. & Gittis, A. H. Transgenic mouse lines subdivide external segment of the globus pallidus (GPe) neurons and reveal distinct GPe output pathways. J. Neurosci. 34, 2087–2099 (2014)
Hamani, C., Saint-Cyr, J. A., Fraser, J., Kaplitt, M. & Lozano, A. M. The subthalamic nucleus in the context of movement disorders. Brain 127, 4–20 (2004)
Breysse, E., Pelloux, Y. & Baunez, C. The good and bad differentially encoded within the subthalamic nucleus in rats(1,2,3). eNeuro 2, ENEURO.0014-15.2015 (2015)
Jhou, T. C., Fields, H. L., Baxter, M. G., Saper, C. B. & Holland, P. C. The rostromedial tegmental nucleus (RMTg), a GABAergic afferent to midbrain dopamine neurons, encodes aversive stimuli and inhibits motor responses. Neuron 61, 786–800 (2009)
Tian, J. & Uchida, N. Habenula lesions reveal that multiple mechanisms underlie dopamine prediction errors. Neuron 87, 1304–1316 (2015)
He, M. et al. Cell-type-based analysis of microRNA profiles in the mouse brain. Neuron 73, 35–48 (2012)
Penzo, M. A. et al. The paraventricular thalamus controls a central amygdala fear circuit. Nature 519, 455–459 (2015)
Li, L. et al. Visualizing the distribution of synapses from individual neurons in the mouse brain. PLoS One 5, e11503 (2010)
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A. D. Quantitative measures of cluster quality for use in extracellular recordings. Neuroscience 131, 1–11 (2005)
Courtin, J. et al. Prefrontal parvalbumin interneurons shape neuronal activity to drive fear expression. Nature 505, 92–96 (2014)
Lau, B. & Glimcher, P. W. Dynamic response-by-response models of matching behavior in rhesus monkeys. J. Exp. Anal. Behav. 84, 555–579 (2005)
Acknowledgements
We thank A. Cutrone, D. Li, and G.-R. Hwang for technical assistance, V. Rao and N. Uchida for sharing the Matlab code for the ROC and clustering analysis, S. D. Shea and S. H. Ebbesen for critical reading of the manuscript, Z. J. Huang for providing mouse strains, and members of the Li laboratory for helpful discussions. This work was supported by grants from the National Institutes of Health (NIH) (R01MH108924 to B.L.), the Dana Foundation (to B.L.), NARSAD (to B.L., M.S. and S.A.), Louis Feil Trust (to B.L.), the Stanley Family Foundation (to B.L.), Simons Foundation (to B.L.), Wodecroft Foundation (to B.L.), and an EMBO Long-Term Fellowship Award (to M.S.).
Author information
Authors and Affiliations
Contributions
M.S. and B.L. designed the study. M.S. conducted experiments and analysed the data. K.Y. assisted with analysis and implementation of the in vivo recording experiments. S.A. and M.P. performed the patch clamp recording experiments. A.N.H. assisted with behavioural training and tracing experiments. J.T. designed and produced the starter virus for the rabies tracing and assisted all the rabies tracing experiments. L.M. assisted with the in vitro electrophysiology experiments. L.H.T. and L.W. assisted with the analysis of behaviour in the probabilistic switching task. M.S. and B.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks G. Stuber and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
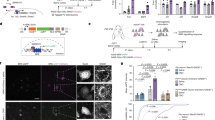
Extended Data Figure 1 Vglut2 and Somatostatin are markers for GPh neurons.
a, Image showing the projection patterns of non-specifically labelled neurons (green, infected with adeno-associated virus (AAV) expressing GCaMP6 (AAV1-Syn-GCAMP6f.WPRE.SV40); signal was enhanced by anti-GFP antibody; see Methods) and Vglut2+ neurons (red, infected with AAV expressing mCherry in a Cre-dependent manner (AAV8-hSyn-DIO-mCherry); signal was enhanced by anti-mCherry antibody; see Methods) in the EP of a Vglut2-Cre mouse. b, Confocal images of the LHb, ventrolateral thalamus (VL) and ventromedial thalamus (VM), showing fibres originating from the non-specifically labelled neurons (green) and Vglut2+ neurons (red) in the EP. c, Quantification of the GFP and mCherry fluorescence intensity in the projection targets of the EP neurons. d, Upper panel, representative image showing retrograde labelling of GPh neurons by injection of the cholera toxin subunit B conjugated to Alexa Fluor 594 (CTB-594) into the LHb (inset) of Vglut2-Cre;Rosa26-stopflox-H2b-GFP mice, in which Vglut2+ cells can be identified based on their expression of nuclear GFP. Lower panels, high-magnification pictures of the boxed area in the EP in the upper panel, showing the co-labelling of GPh neurons by CTB-594 and Vglut2 (arrowheads). The vast majority of CTB-labelled neurons expressed Vglut2 (95.45 ± 1.2% (mean ± s.e.m.), n = 6 mice). e, Upper panel, a representative image showing retrograde labelling of VM-projecting EP neurons by injection of CTB-594 into the VM (inset) of Vglut2-Cre;Rosa26-stopflox-H2b-GFP mice. Lower panels, high-magnification pictures of the boxed area in the EP in the upper panel, showing the segregation of the EP neurons labelled by CTB-594 and those labelled by Vglut2 (arrowheads). Very few CTB-labelled neurons expressed Vglut2 (0.51 ± 0.45%, n = 6 mice). f, Upper panel, a representative image showing retrograde labelling of VM-projecting EP neurons by injection of CTB-594 into the VM (inset). Lower panels, high-magnification pictures of the boxed area in the EP in the upper panel, showing the segregation of the EP neurons labelled by CTB-594 and those labelled by anti-Som antibody (arrowheads). Very few CTB-labelled cells expressed Som (0.88 ± 0.72%, n = 5 mice). g, Upper panel, a representative image showing antibody labelling of Som in the EP of Vglut2-Cre;Rosa26-stopflox-H2b-GFP mice. Lower panels, high-magnification pictures of the boxed area in the upper panel, showing the co-labelling of EP neurons by Som and Vglut2 (arrowheads). The vast majority of Vglut2 neurons expressed Som (90.87 ± 0.79%, n = 6 mice). h, A cartoon showing the only projection target of GPh neurons (red) and the different projection targets of classic GPi neurons (blue). Diagram in h was modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
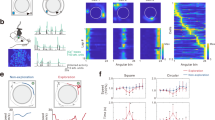
Extended Data Figure 2 Classification of EP neurons on the basis of their distinct response profiles.
a, Schematic of the experimental approach used for in vivo recording and optogenetic tagging. b, Photomicrograph showing a DiI-labelled recording site. c, Schematics showing the locations of the recording sites (n = 15 mice). d, Responses of three example neurons in the classic conditioning task. e, Left, auROC plots of the responses of all neurons during large reward trials. Red, increase from baseline; blue, decrease from baseline; each row represents one neuron. Green bars indicate the neurons that were optogenetically tagged (n = 11 neurons). The three main clusters are arranged in order to match the neurons presented in d. Right, first three principle components and hierarchical clustering dendrogram showing the relationship of each neuron within the three clusters. f, Average firing rates of the three types of neurons (n = 86 neurons from 9 mice). g, Plots of peristimulus time histogram (PSTH) showing inhibition for type I (top, n = 7 neurons from 4 mice), but no change for type II (middle, n = 9 neurons from 4 mice) or type III (bottom, n = 10 neurons from 4 mice) neurons in response to green light pulses (green bars, 200 ms; 100 trials per neuron, 0.3 Hz). Only type II and type III neurons that were recorded in the same sessions and animals as those of the light-responsive type I neurons represented in g are shown. h, auROC plots of the responses of all 38 neurons (n = 9 mice) recorded during large punishment trials. i, j, Average firing rates of type II (n = 11 neurons from 9 mice) (i) and type III (n = 11 neurons from 9 mice) (j) neurons during punishment trials. Diagrams in a and c were modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
Extended Data Figure 3 Response profiles of putative GPh neurons during different CS–US contingencies.
a, Graphs showing hierarchical clustering used to identify additional putative GPh neurons used in the analysis for this figure. All data shown (b–i) are from type I neurons only. Left, auROC plots of the responses of all additional neurons recorded. Red, increase from baseline; blue, decrease from baseline. Each row represents one neuron. Green bars indicate the neurons that were optogenetically tagged. Right, first three principle components and hierarchical clustering dendrogram showing the relationship of each neuron within the three clusters. b, auROC plots showing the firing rate changes in response to CS (top) and reward (bottom) before behavioural training. c, auROC plots showing the firing rate changes in response to CS (top) and air puff (bottom) before behavioural training. d, auROC plots showing the firing rate changes in response to an expected (top) or unexpected (bottom) reward. e, auROC plots showing the firing rate changes in response to an expected (top) or unexpected (bottom) air puff. f, auROC plots showing the firing rate changes in response to receiving an expected air puff (left) or having an expected air puff omitted (right). g, Histogram of difference in firing rate between air puff omission and air puff (filled bars, P < 0.05, t-test). Values are represented using auROC. h, auROC plots showing the firing rate changes in response to receiving an expected reward (left) or having an expected reward omitted (right). i, Histogram of difference in firing rate between reward omission and reward (filled bars, P < 0.05, t-test). Values are represented using auROC.
Extended Data Figure 4 Optic fibre implantation locations.
a, A schematic of the experimental approach used for Arch-mediated inhibition of GPh neurons. b, A photomicrograph showing the location of optic fibre placement and ArchT–GFP+ GPh neurons within the EP. c, Schematics showing the location of the optic fibre placements (n = 5). d, A schematic of the experimental approach used for Arch-mediated inhibition of the GPh–LHb projection. e, A photomicrograph showing the location of optic fibre placement and ArchT–GFP+ axon fibres within the LHb. f, Schematics showing the location of the optic fibre placements (n = 7). g, A schematic of the experimental approach used for Arch-mediated inhibition of the GPh, which was targeted retrogradely by injection of the LHb with CAV2–Cre. h, A photomicrograph showing the location of the optic fibre placement and ArchT–GFP+ neurons in the EP. i, Schematics showing the location of the optic fibre placements (n = 5). j, A schematic of the experimental approach used for ChR2-mediated excitation of the GPh, which was targeted retrogradely by injection of the LHb with CAV2–Cre. k, A photomicrograph showing the location of the optic fibre placement and ChR2–GFP+ neurons in the EP. l, Schematics showing the location of the optic fibre placements (n = 5). m, Schematic of the experimental approach used for ChR2-mediated activation of the GPh–LHb projection. n, A photomicrograph showing the optic fibre placement and ChR2–YFP+ axon fibres in the LHb. o, Schematics showing the location of the optic fibre placements (n = 6). Diagrams in a, c, d, f, g, i, j, l, m and o were modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
Extended Data Figure 5 Optogenetic inhibition of the GPh drives reward-related behaviours.
a, Confocal images from a Som-Cre;Ai14 mouse, showing the overlap in expression of ChR2–YFP and tdTomato (indicating Som+ neurons) in GPh neurons. b, Quantification of the percentage of ChR2–YFP+ neurons that expressed tdTomato (n = 2). c, Confocal images from a Som-Cre;Ai14 mouse, showing the overlap in expression of ArchT–YFP and tdTomato in GPh neurons. d, Quantification of the percentage of ArchT–YFP+ neurons that expressed tdTomato (n = 2). e, Schematic of the experimental approach used for ArchT-mediated inhibition of GPh neurons. f, Heat maps for the activity of a representative mouse at baseline (top), or during optogenetic inhibition of the GPh in either the left (middle) or right (bottom) chamber. g, GPhArch mice (n = 5), but not GPheYFP mice (n = 5), showed a significant place preference for the chamber paired with laser stimulation in the GPh (F(5,29) = 14.95, P < 0.0001, ***P < 0.001, **P < 0.01, two-way ANOVA followed by Tukey’s test). h, Schematic of the experimental approach used for ArchT-mediated inhibition of GPh axon terminals in the LHb. i, GPhArchT mice (n = 7), but not GPheYFP mice (n = 5), showed a significant place preference for the chamber paired with laser stimulation in the GPh (F(5,35) = 52.22, P < 0.0001, ***P < 0.001, **P < 0.01, two-way ANOVA followed by Tukey’s test). j, GPhArch mice (n = 5) made significantly more nose pokes than GPheYFP mice (n = 5) to obtain laser stimulation in the GPh (t(8) = 2.61, *P < 0.05, t-test). k, Schematic of the retrograde labelling approach used to target the GPh for ArchT-mediated optical inhibition (top). GPhCAV-Cre/Arch mice (n = 5), but not GPheYFP mice (n = 5), showed a significant place preference for the chamber paired with laser stimulation in the GPh (bottom) (F(5,29) = 5.98, P < 0.01, *P < 0.05, two-way ANOVA followed by Tukey’s test). l, Schematic of the retrograde labelling approach used to target the GPh for ChR2-mediated optical excitation (top). GPhCAV-Cre/ChR2 mice (n = 5), but not GPheYFP mice (n = 5), showed a significant place aversion for the chamber paired with laser stimulation in the GPh (bottom) (F(5,29) = 26.50, P < 0.0001; ***P < 0.001, **P < 0.01, two-way ANOVA followed by Tukey’s test). m, Heat maps for the activity of a representative mouse at baseline (top), or during optogenetic excitation of the GPh in either the left (middle) or right (bottom) chamber. n, Mice did not move faster (left) or further (right) during the Arch stimulation sessions when compared to their baseline activity (t(32) = 0.15, P > 0.05; t(32) = 0.16, P > 0.05; t-test, n = 17). o, Mice did not move faster (left) or further (right) during the ChR2 stimulation sessions when compared to their baseline activity (t(8) = 0.12, P > 0.05; t(8) = 0.26, P > 0.05; t-test, n = 5). All data are presented as mean ± s.e.m. Diagrams in e, h, k and i were modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
Extended Data Figure 6 A probabilistic switching task for studying action evaluation.
a, Schematic of the task. b, The probability of choosing the left port by one mouse for reward history in which consecutive choices to either the right or the left port were made during the previous two trials. c, The contribution of rewarded and unrewarded outcomes in the previous five trials—represented by regression coefficients βReward and βNo Reward, respectively—to choices in the current trial (n = 10 mice, 4,685 ± 786 trials per mouse). d, The fraction of left port choice for ten mice plotted against the relative action value (sum of the regression coefficients from the previous two trials). Data from each mouse was grouped into ten bins and represented by a distinct colour. e, The actual probability of choosing the left port plotted against the probability of choosing the left port predicted by the logistic regression model. f, Example data from one session showing 12 trial blocks. Blue bars represent left reward blocks (top); orange bars indicate right reward blocks (bottom). Green, orange, and red ticks represent whether a particular trial was a correct rewarded trial, a correct unrewarded trial, or an incorrect trial, respectively. The grey dashed line represents a four-trial running average of the mouse’s probability of choosing the left port, and the black line indicates the probability of choosing the left port predicted by the logistic regression model. g, h, Change in chosen value one to three trials after optogenetic inhibition of the GPh (g), or activation of the GPh–LHb pathway (h). i, Changes in chosen value one trial after optogenetic activation or inhibition at the left or right reward port. g, h, ****P < 0.0001, t-test. b, c, g–i, Data are represented as mean ± s.e.m.
Extended Data Figure 7 Optogenetic inhibition or activation of the GPh–LHb pathway does not influence action selection.
a, A schematic of optogenetic inhibition of the GPh at the point of action selection. b, Data points indicate the probability of left port choice as a function of action value for the trials in which the photo-stimulation was delivered at the centre port (stim) or was not delivered (no stim). Lines indicate the fit by the logistic regression model on the pooled data for each of the two conditions (n = 5 mice, 15,411 trials, 3,082 ± 1,063 trials per mouse). c, Similar to b, except that control mice with eYFP-expressing GPh neurons were used (n = 6 mice, 56,241 trials, 9,373 ± 596 trials per mouse). d, e, Similar to a and b, except that optogenetic activation of the GPh–LHb projection was applied at the point of action selection (n = 6 mice, 41,557 trials, 8,311 ± 2,565 trials per mouse). f, Similar to e, except that control mice with eYFP-expressing GPh neurons were used (n = 6 mice, 72,423 trials, 12,070 ± 1,673 trials per mouse). g, h, The changes in action value in response to optogenetic stimulation of the GPh–LHb pathway one to three trials after the photostimulation, for mice in which GPh neurons expressed Arch (n = 5) or eYFP (n = 6) (g), or ChR2 (n = 6) or eYFP (n = 6) (h). In b, c, e and f, P values reported for t tests: H0: βstim = 0. i–l, Graphs showing the average withdrawal (calculated as the time from centre port entry to exit) and movement (calculated as the time from centre port exit to the poke at the chosen port) time for trials with or without light stimulation. Both withdrawal time and movement time were shorter when the action value associated with the chosen action was higher. Neither activation of GPh neurons with ChR2 (n = 6 mice) (i, j) (movement time for leftward choices, ChR2 stimulated trials (ChR2) versus unstimulated trials (no stim), F(1,8) = 0.174, P > 0.05; rightward choices, ChR2 versus no stim, F(1,8) = 1.352, P > 0.05; withdrawal time preceding leftward choices, ChR2 versus no stim, F(1,8) = 0.667, P > 0.05; preceding rightward choices, ChR2 versus no stim, F(1,8) = 0.599, P > 0.05; two-way ANOVA), nor inhibition of these neurons with Arch (n = 5 mice) (k, l) (movement time for leftward choices, Arch stimulated trails (Arch) versus unstimulated trials (no stim), F(1,8) = 0.105, P > 0.05; rightward choices, Arch versus no stim, F(1,8) = 0.023, P > 0.05; withdrawal time preceding leftward choices, Arch versus no stim, F(1,8) = 0.821, P > 0.05; preceding rightward choices, Arch versus no stim, F(1,8) = 0.459, P > 0.05; two-way ANOVA) had any significant effect on the ongoing behaviour. Data in g–l are presented as mean ± s.e.m.
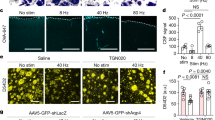
Extended Data Figure 8 Weakening of excitatory or inhibitory synapses onto GPh neurons and its effects on the sensitivity to negative or positive feedback.
a, Confocal images from a Som-Cre;Ai14 mouse, showing the overlap in expression of GluA4-ct–GFP (delivered by injecting the EP with the AAV-DIO-GluA4-ct-GFP) and tdTomato (indicating the expression of Som) in GPh neurons. 97.86 ± 2.9% of GluA4-ct–GFP+ neurons expressed tdTomato (n = 2 mice). b, Schematics of the experimental approach. CTB-594 was injected into the LHb to label GPh neurons in the EP. On the right is an enlarged graph of the boxed area in the cartoon on the left. Inset is a photomicrograph showing simultaneous recording of a CTB+/GluA4-ct+ GPh neuron and a nearby CTB+/GluA4-ct– GPh neuron. c, EPSC traces recorded from the two neurons shown in b. d, Quantification of the ratio between AMPA receptor-mediated EPSC amplitude and NMDA receptor-mediated EPSC amplitude (AMPA/NMDA ratio) for the two populations of GPh neurons (CTB+/GluA4-ct+, n = 6 cells; CTB+/GluA4-ct–, n = 8 cells; n = 3 mice; t(12) = −1.89, *P < 0.05, t-test). e, A representative image showing the expression of GluA4-ct–GFP (delivered by injecting the EP of a Vglut2-Cre mouse with the AAV-DIO-GluA4-ct-GFP) in GPh neurons (left) and a schematic of the approach (right). f, The win–stay percentage in these mice (GPhGluA4-ct, 94.17 ± 1.02%; GPheYFP, 95.82 ± 0.51%; P > 0.05, t test). g, For animals (n = 10 mice) used in Fig. 4a–e, the number of GPh neurons that were infected with the GluA4-ct–GFP virus correlated with the change in animal behaviour in the switching task, measured as an increase in action value following two consecutive unrewarded trials (R2 = 0.72, P < 0.05 by a linear regression). h, Contributions of rewarded outcomes over the past five trials, as reflected by their regression coefficients, to the current choice. GPhGluA4-ct mice were not significantly different from control mice or their pre-surgery condition (first two trials back × groups, F(3,33) = 0.5412, P > 0.05; two-way ANOVA, n = 10 GPhGluA4-ct mice and n = 7 control mice). i, The action value following two sequentially rewarded trials was not significantly different between GPhGluA4-ct mice and GPheYFP mice (P > 0.05, t-test). j, Confocal images from a Som-flp mouse, showing the overlap in expression of Cre–GFP (delivered by injecting the EP with the AAV-FSF-GFP-Cre) and somatostatin, recognized through antibody labelling. 96.25 ± 2.3% of Cre–GFP+ neurons expressed somatostatin (n = 2 mice). k, Schematics of the experimental approach. CTB-594 was injected into the LHb to label GPh neurons in the EP. On the right is an enlarged graph of the boxed area in the cartoon on the left. l, Sample miniature IPSC (mIPSC) traces recorded from a GPh neuron that expressed Cre–GFP (and thus had γ2 ablated (γ2-KO)) and a control GPh neuron that did not express the Cre–GFP (γ2-WT). m, Quantification of the frequency (left) and amplitude (right) of mIPSCs recorded from the two groups of GPh neurons (γ2-KO, n = 7 cells; γ2-WT, n = 10 cells; n = 3 mice; frequency, t(15) = 5.51, ****P < 0.0001; amplitude, t(15) = 8.19, ****P < 0.0001; t-test). n, A representative image showing the expression of Cre–GFP (delivered by injecting the EP of a Som-Flp;Gabrg2flox mouse with the AAV-FSF-GFP-Cre) in GPh neurons (left) and a schematic of the approach (right). o, The lose–switch percentage in these mice (P > 0.05, t-test). p, For animals (n = 9) used in Fig. 4f–j, the number of GPh neurons that were infected with the Cre–GFP virus correlated with the change in animal behaviour in the switching task, measured as a reduction in action value following two consecutive rewarded trials (R2 = 0.53, P < 0.05 by a linear regression). q, The negative regression coefficients associated with the past five trials were not significantly different between GPhγ2-KO mice and control mice either before or after surgery (first two trials back × groups, F(3,35) = 0.9072, P > 0.05, n = 9 GPhγ2-KO mice and n = 9 control mice). r, The action value following two sequentially unrewarded trials was not significantly different between GPhγ2-KO mice and GPhmCherry mice (P > 0.05, t-test). All data are represented as mean ± s.e.m. Diagrams in b, e, k and n were modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
Extended Data Figure 9 Monosynaptic inputs onto the GPh and a schematic of the circuitry for reinforcement learning.
a, Schematics of experimental design. The GPh neurons in the EP were targeted using either Vglut2-Cre;Rosa26-stopflox-tTA mice or by injecting the LHb of Rosa26-stopflox-tTA mice with the retrograde CAV2–Cre. b, Images showing the starter cell location in the EP. c, Relationship between the number of starter and input neurons. d, Graph showing the fraction of monosynaptically labelled neurons in each brain region that projects to the GPh (n = 9 mice) e, Confocal images of the rabies virus and parvalbumin (PV) labelled neurons in the GPe. Only a small fraction of the virally labelled GPe cells expressed PV (arrows). On the right is a high-magnification image of the boxed area in the GPe. f, Quantification of the fraction of rabies virus-labelled GPe neurons that expressed PV (n = 3 mice). g, Center of mass analysis for all GPe labelled neurons (n = 9 mice). h, A confocal image of the parasubthalamic nucleus (pSTN) showing monosynaptically labelled neurons. i, Center of mass analysis for all pSTN labelled neurons (n = 9 mice). j, A schematic showing the proposed selection and evaluation circuits within the basal ganglia. Question marks indicate elements of the proposed circuit that remain to be tested experimentally. Diagrams in a, g and i were modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
Extended Data Figure 10 The proposed function of the basal ganglia and midbrain evaluation circuits.
a, Schematic showing the activity of GPh neurons and the downstream circuitry controlling the midbrain dopaminergic system. b, Proposed sequence of events by which GPh activity may influence the firing rate in downstream structures. Upward arrows indicate an increase in firing; downward arrows indicate a decrease in firing. RMTg, Rostromedial tegmental nucleus; SNc, Substantia nigra pars compacta; VTA, ventral tegmental area; DA, dopamine. DR, dorsal raphe; MR, median raphe. A question mark (?) indicates that alternative circuits downstream of the LHb, including the serotonergic raphe nuclei, may constitute other key pathways that also process the GPh–LHb prediction error signals that we demonstrate in this study. Diagram in a was modified from the Allen Mouse Brain Atlas, Allen Institute for Brain Science; available from http://mouse.brain-map.org/.
Supplementary information
Supplementary Tables
Supplementary Table 1 contains equations for alternative models to describe the behavioural effect of optogenetic manipulations of the GPh. (PDF 215 kb)
Arch-mediated inhibition of the GPh reinforces actions
This video shows how inhibition of the GPh at the time of outcome evaluation biases the mouse to repeat the same choice. For the purpose of the video optical inhibition was delivered on a number of consecutive trials instead of the randomly selected 10% of trials, as was the case for the data shown in Fig. 3. (MP4 25822 kb)
ChR2-mediated excitation of the GPh discourages actions
This video shows how excitation of the GPh-LHb axon terminals in the LHb at the time of outcome evaluation biases the mouse to choose an alternative action following the stimulation. (MP4 6397 kb)
Rights and permissions
About this article
Cite this article
Stephenson-Jones, M., Yu, K., Ahrens, S. et al. A basal ganglia circuit for evaluating action outcomes. Nature 539, 289–293 (2016). https://doi.org/10.1038/nature19845
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19845
This article is cited by
-
Optogenetic recruitment of hypothalamic corticotrophin-releasing-hormone (CRH) neurons reduces motivational drive
Translational Psychiatry (2024)
-
A neural mechanism for conserved value computations integrating information and rewards
Nature Neuroscience (2024)
-
Plasticity of neuronal dynamics in the lateral habenula for cue-punishment associative learning
Molecular Psychiatry (2023)
-
Distributed processing for value-based choice by prelimbic circuits targeting anterior-posterior dorsal striatal subregions in male mice
Nature Communications (2023)
-
Plastic and stimulus-specific coding of salient events in the central amygdala
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.