Abstract
Pancreatic cancer, a highly aggressive tumour type with uniformly poor prognosis, exemplifies the classically held view of stepwise cancer development1. The current model of tumorigenesis, based on analyses of precursor lesions, termed pancreatic intraepithelial neoplasm (PanINs) lesions, makes two predictions: first, that pancreatic cancer develops through a particular sequence of genetic alterations2,3,4,5 (KRAS, followed by CDKN2A, then TP53 and SMAD4); and second, that the evolutionary trajectory of pancreatic cancer progression is gradual because each alteration is acquired independently. A shortcoming of this model is that clonally expanded precursor lesions do not always belong to the tumour lineage2,5,6,7,8,9, indicating that the evolutionary trajectory of the tumour lineage and precursor lesions can be divergent. This prevailing model of tumorigenesis has contributed to the clinical notion that pancreatic cancer evolves slowly and presents at a late stage10. However, the propensity for this disease to rapidly metastasize and the inability to improve patient outcomes, despite efforts aimed at early detection11, suggest that pancreatic cancer progression is not gradual. Here, using newly developed informatics tools, we tracked changes in DNA copy number and their associated rearrangements in tumour-enriched genomes and found that pancreatic cancer tumorigenesis is neither gradual nor follows the accepted mutation order. Two-thirds of tumours harbour complex rearrangement patterns associated with mitotic errors, consistent with punctuated equilibrium as the principal evolutionary trajectory12. In a subset of cases, the consequence of such errors is the simultaneous, rather than sequential, knockout of canonical preneoplastic genetic drivers that are likely to set-off invasive cancer growth. These findings challenge the current progression model of pancreatic cancer and provide insights into the mutational processes that give rise to these aggressive tumours.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hruban, R. H., Goggins, M., Parsons, J. & Kern, S. E. Progression model for pancreatic cancer. Cancer Res. 6, 2969–2972 (2000)
Moskaluk, C. A., Hruban, R. H. & Kern, S. E. p16 and K-RAS gene mutations in the intraductal precursors of human pancreatic adenocarcinoma. Cancer Res. 57, 2140–2143 (1997)
Wilentz, R. E. et al. Inactivation of the p16 (INK4A) tumor-suppressor gene in pancreatic duct lesions: loss of intranuclear expression. Cancer Res. 58, 4740–4744 (1998)
Wilentz, R. E. et al. Loss of expression of Dpc4 in pancreatic intraepithelial neoplasia: evidence that DPC4 inactivation occurs late in neoplastic progression. Cancer Res. 60, 2002–2006 (2000)
Lüttges, J. et al. Allelic loss is often the first hit in the biallelic inactivation of the p53 and DPC4 genes during pancreatic carcinogenesis. Am. J. Pathol. 158, 1677–1683 (2001)
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015)
Cooper, C. S. et al. Analysis of the genetic phylogeny of multifocal prostate cancer identifies multiple independent clonal expansions in neoplastic and morphologically normal prostate tissue. Nat. Genet. 47, 367–372 (2015)
Ross-Innes, C. S. et al. Whole-genome sequencing provides new insights into the clonal architecture of Barrett’s esophagus and esophageal adenocarcinoma. Nat. Genet. 47, 1038–1046 (2015)
Stachler, M. D. et al. Paired exome analysis of Barrett’s esophagus and adenocarcinoma. Nat. Genet. 47, 1047–1055 (2015)
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010)
Chari, S. T. et al. Early detection of sporadic pancreatic cancer: summative review. Pancreas 44, 693–712 (2015)
Eldredge, N. & Gould, S. J. in Models in Paleobiology (ed. Schopf, T. J. M. ) 82–115 (Freeman, Cooper and Company, 1972)
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015)
Campbell, P. J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010)
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011)
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013)
Rausch, T. et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148, 59–71 (2012)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Real, F. X. A. A “catastrophic hypothesis” for pancreas cancer progression. Gastroenterology 124, 1958–1964 (2003)
Zhang, C.-Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015)
Maciejowski, J., Li, Y., Bosco, N., Campbell, P. J. & de Lange, T. Chromothripsis and kataegis induced by telomere crisis. Cell 163, 1641–1654 (2015)
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015)
Li, Y. et al. Constitutional and somatic rearrangement of chromosome 21 in acute lymphoblastic leukaemia. Nature 508, 98–102 (2014)
Murphy, S. J. et al. Genetic alterations associated with progression from pancreatic intraepithelial neoplasia to invasive pancreatic tumor. Gastroenterology 145, 1098–1109.e1 (2013)
Özdemir, B. C. et al. Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell 25, 719–734 (2014)
Rhim, A. D. et al. Stromal elements act to restrain, rather than support, pancreatic ductal adenocarcinoma. Cancer Cell 25, 735–747 (2014)
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005)
Haeno, H. et al. Computational modeling of pancreatic cancer reveals kinetics of metastasis suggesting optimum treatment strategies. Cell 148, 362–375 (2012)
Rhim, A. D. et al. EMT and dissemination precede pancreatic tumor formation. Cell 148, 349–361 (2012)
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013)
Acknowledgements
We would like to thank N. Simard, S. Zhao and members of the SickKids-UHN Flow facility for technical support. Funding sources for this study include grants to the Pancreatic Cancer Sequencing Initiative program from the Ontario Institute for Cancer Research (OICR), through support from the Ontario Ministry of Research and Innovation, the Canada Foundation for Innovation; research award to F.N. from the OICR and the Canadian Institutes for Health Research (CIHR); Canadian Friends of the Hebrew University, the SMGS Family Foundation, NCI grant P50 CA102701 (Mayo Clinic SPORE in Pancreatic Cancer) and NCI grant R01 CA97075 (Pancreatic Cancer Genetic Epidemiology Consortium). F.N. is supported by a fellowship award from CIHR and is a recipient of a scholar’s research award from the Ontario Institute of Cancer Research (OICR), through support from the Ontario Ministry of Research and Innovation. G.Z. is a Clinician–Scientist of the Fonds de la Recherche en Sante du Quebec. P.J.C. is a Wellcome Trust Senior Clinical Fellow. T.J.H., L.D.S., J.D.M. and S.G. are recipients of Senior or Clinician–Scientist Awards from the Ontario Institute for Cancer Research.
Author information
Authors and Affiliations
Contributions
Data analysis and interpretation was performed by F.N., M.L., Y.L., M.C.-S.-Y., G.W.W., A.A.C., F.X.R., P.J.C., S.G. and T.J.H.; tumour enrichment by S.-B.L., I.L. and F.N.; pathological assessment by T.W., M.-S.T., J.M.S.B., M.H.R. and S.H.-B.; genomics by R.E.D., A.M.K.B., K.N., J.C.K., L.T., N.B., D.P., L.H., E.I., G.H.J., J.J., L.G.T.J., J.D.M., L.D.S., L.I.S., L.H., J.E.D., C.K.Y., T.B. and L.B.A.; FISH by O.L.; CELLULOID analysis by M.L. and single-cell analysis by G.W.W., J.T.S. and F.N. Sample acquisition, annotation and collection from institutes external to University Health Network. was performed by G.M.P., M.A.H., G.Z. and C.L. Sample acquisition, annotation and collection from the University Health Network was performed by J.M.W., A.B., S.G. and S.P.C. The study was designed by F.N., T.J.H. and S.G.; F.N. prepared and wrote the manuscript; the manuscript was edited by M.L., F.X.R., J.E.D., P.J.C., T.J.H. and S.G.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Chanock, M. Rossi and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
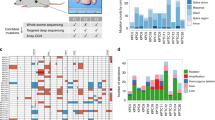
Extended Data Figure 1 Tumour enrichment and overview of somatic alterations in the cohort used in this study.
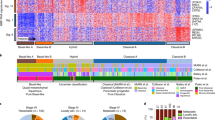
a, Flow cytometry profiles of EpCAM and CD45 from 2 (of 21) representative cases of pancreas ductal adenocarcinoma (PDA) (i,ii). On the right, post-sort analysis of EpCAM+ cells (Tu) and CD45+ lymphocytes (Ly) demonstrates the high level of purity obtained from flow-sorting. b, Immunohistochemical analysis of formalin-fixed tumours using the EpCAM clone for flow sorting in a (H&E, haematoxylin and eosin). Two independent cases are shown (i, ii). c, Profiles of haematoxylin-stained sections of PDA before and after LCM from two representative cases (of 86) (i, ii). d, Box and whisker plots represent median and 10–90 percentile ranges of tumour cellularity of flow-sorted (n = 21), LCM (n = 86) and the total cohort (n = 107) of tumours. Dashed line depicts cellularity of bulk tumours that have not undergone enrichment. e, Overview of somatic alterations of the cohort used in the study. f, X-chromosome mutation ratio in diploid pancreatic cancer genomes showing hypermutation on this chromosome in females. Males were corrected for single copy of the X chromosome by doubling the raw value. P values were derived from t-tests. A more detailed description of these data is provided in Supplementary Results.
Extended Data Figure 2 CELLULOID validation.
The copy number for common alterations (TP53, SMAD4; shown by black arrow) was derived from ploidy estimates generated by CELLULOID. Six diploid and five polyploid tumours were analysed by FISH (shown on the right of each contour plot). In all cases, the copy number from CELLULOID ploidy estimates was confirmed. In Pcsi_0084 (diploid), CELLULOID predicted zero copies of SMAD4. The allelic ratio in this region was 50% (heterozygous) as only reads from normal cells spanned this region. In Ashpc_0027, both CELLULOID and FISH indicate that this tumour is polyploid. The CELLULOID plot demonstrates that there is a further subclonal amplification in TP53 from polyploid clone (copy state = 3.2 derived from one allele). FISH analysis shows tumour cells with two or three copies of TP53 supporting this is subclonal. Copy number by FISH for SMAD4 and TP53 is indicated in red at the top right of each plot.
Extended Data Figure 3 Tumour ploidy and genetic instability in pancreatic cancer.
a, Tumour ploidy and sample cellularity estimates are interconnected: although the ploidy of a tumour can always be doubled and still provide copy number segments at integer levels (albeit only at even values), the estimate of cellularity would have to decrease. To maintain an allelic ratio at a given value, the proportion of tumour cells has to be reduced to compensate for the higher copy numbers in them (from a cellularity value t to a value t/(2 − t) in the case of a doubling of the ploidy). A test can thus be designed to verify that ploidy estimates have not been systematically over- or underestimated, simply by comparing the distribution of cellularity estimates stratified by ploidy. P value was derived using Kruskal–Wallis test. b, Deviation from baseline ploidy in diploids, tetraploids and hexaploids indicates a marked loss of genomic material in polyploids. c, Box and whisker plots (showing the median and 10th–90th percentile ranges) of the total copy number alterations in polyploid and diploid tumours. d, Mutational signatures of the 107 genomes used in this study. The signatures were derived using the trinucleotide mutation context as previously published18. The proportion of individual signature operative in each tumour is shown in the bar plot. The overall classification of each case is indicated below. Signatures of polyploidy tumours is shown on the left, diploids is shown on the right. ND, not done; n = 1 polyploid and 4 diploid patient samples. Detailed analysis of mutational signatures in PDA is covered elsewhere (Connor et al., manuscript under review) e, Percentage of copy number losses (left) and gains (right) that occurred before (yellow) or after (blue) genome duplication for each polyploid tumour. Box and whisker plots depict median ± 10th–90th percentile range. P values were derived using a t-test.
Extended Data Figure 4 Characterization of chromothripsis events in pancreatic cancer.
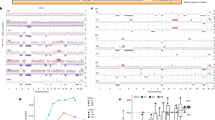
a, The distribution of chromothripsis events across the genome (single-chromosome, white; multi-chromosome, black). **P < 0.001 (Monte Carlo sampling, Supplementary Methods). b, The specific effects of chromothripsis on the copy number of chromosome 18 (top, n = 22), chromosome 12 (middle, n = 15), and chromosome 19 (bottom, n = 5). Statistical differences in copy number between the groups were performed using Wilcoxon test using 10-kb bins that covered GATA6 (chromosome 18), KRAS (chromosome 12) and PAK4 (chromosome 19) genes (description of PAK4 event is covered in supplementary results). Copy number profiles of polyploids were adjusted according to tumour ploidy to allow comparison against diploids (referred to as ‘Normalized copy number’ on the y axis). Interquartile ranges for chromothripsis cases are indicated in pale red and for non-chromothripsis cases in pale blue. c, Two cases of chromothripsis resulting in the amplification of the mutant KRAS allele. In Pcsi_0290, the mutant allele was amplified as part of a multi-chromosomal event involving chromothripsis and BFB with chromosome 18 (top). In Pcsi_0356, the chromothripsis event was co-opted with cycles of BFB to knock out the wild-type allele (bottom). The absolute copy number of the locus encompassing KRAS and mutation is shown for each case. d, Cumulative incidence of chromothripsis events in polyploid and diploid tumours (P = 0.013, Fisher’s exact test).
Extended Data Figure 5 Most copy number alterations arise from individual chromothripsis events.
a, In Pcsi_0082, five distinct chromothripsis events on chromosome 15 (top, 1), chromosome 18 (top, 2), chromosome 8 (top, 3), chromosome 13 (bottom, 4), and chromosome 16 (bottom, 5) are displayed. Copy number steps on chromosome 15 (1), chromosome 8 (2) are 2 or greater indicating that these events occurred before polyploidization. Single copy number steps on chromosome 18 (2), chromosome 13 (4) and chromosome 16 (5) indicate that these events were sustained after polyploidization. The single rearrangement between chromosome 15 and chromosome 18 appears to be independent from the chromothripsis on chromosome 18. Pie charts depict the proportion of copy number alterations derived from each chromothripsis event. b, Distribution of copy number alterations due to chromothripsis for all cases where such an event was detected by ChromAL. c, In Ashpc_0008, two multi-chromosomal chromothripsis events, joining chromosome 14, chromosome 6, chromosome 18 (top, 1), and chromosome 3, chromosome 20 (bottom, 2), are shown (discussed in Supplementary Results).
Extended Data Figure 6 Characterization of chromothripsis and polyploidy in metastases.
a, CELLULOID plots illustrating polyploidy in metastases. In Pcsi_0380, the primary tumour was directly available for analysis. Similarly to Pcsi_0378, multiple metastases were polyploid suggesting the primary tumour was also polyploid. The primary tumour was unavailable for sequencing in this case. b, A case (Pcsi_0407) with discordant ploidy amongst different metastases. c, Percentage of diploid mutations from liver metastases that are shared (white) or unique (black) when compared to the primary tumour or the lymph node metastasis. d, Plots of chromothripsis events in metastases. ln, lymph node; lv, liver; pa, primary tumour.
Extended Data Figure 7 Chromothripsis and polyploidy in Pcsi_0410.
This figure accompanies Fig. 2. a, CELLULOID (left panel) and chromothripsis plots (middle and right) of the different metastases from a patients with fulminant metastatic progression. b, Copy number and LOH from chromosome 8 (left) and chromosome 6 (right) chromothripsis events indicate that these events were sustained before polyploidization.
Extended Data Figure 8 Case of a simultaneous loss of CDKN2A and SMAD4 due to a chromothripsis event.
a, Rearrangement and copy number profile of a multi-chromosome chromothripsis event between chromosome 9 and chromosome 18 (Pcsi_0171). b, Detailed view of the two inversions (one in the head-to-head orientation (HH), the other in tail-to-tail orientation (TT) for more detail, see Methods) in the chromothripsis event that resulted in the concurrent loss of CDKN2A and SMAD4. c, Schematic depiction of the temporal order of events derived from the rearrangement profile shown in a. d, Summary of tumour evolution in Pcsi_0171. A more detailed description of Pcsi_0171 is provided in Supplementary Results.
Extended Data Figure 9 Single-cell sequencing reconstruction of the evolutionary events when rearrangements did not span the classical pancreatic cancer drivers.
a, A fresh tumour specimen (Ashpc_0008) was dissociated and single tumour cells were deposited using flow sorting. The whole genomes of 96 single cells were amplified using REPLI-g and paired-end whole-genome sequencing was performed using an Illumina HiSeq 2500 system. Single cells were sequenced to a median whole-genome depth of 3.9× (Supplementary Fig. 18). Only cells with enough whole-genome coverage (n = 70) were used in the analysis. This sequencing depth allowed us to track heterozygous SNPs across the whole genome in single cells. Using this methodology, we were able to follow LOH events across the whole genome in single cells that show high concordance with bulk tumour tissue (Supplementary Fig. 18). Hierarchical clustering based on LOH events across the whole genome was performed and found four independent cell clusters. b, Specific LOH events on chromosome 3, chromosome 9, chromosome 17 and chromosome 18 are shown from single cells in a. The chromothripsis event on chromosome 3 is shown in greater detail in Extended Data Fig. 5c. A summary of the sequence of allelic losses is shown below. Supportive data that allelic losses precede mutational inactivation is shown in Supplementary Figs 13, 14. c, Plot of the shared chromosomal break point on chromosome 18 on the bulk (top), preneoplastic single cell (middle) and tumour single cell (bottom). d, The classical model of pancreatic tumour progression.
Extended Data Figure 10 Theoretical model of pancreatic cancer tumour progression.
Shown is the classical model of tumour evolution driven at a gradual pace (grey) and an alternate model driven at punctuated equilibrium (red). In the classical model, there is a period of latency between the driver mutations that lead to tumour development and the multiple, independent, transforming events are required for tumour development (top, grey dashed line; bottom-left schematic). In the punctuated equilibrium model, tumour development can be divided into two major events, the cancer-initiating event and cancer-transforming event (top, red dashed line; bottom-right schematic). Under this model, most mutations (indicated by ×) would accrue in an extended phase of preneoplastic tumour development. Transformation, probably due to genetic instability from copy number changes (arrow heads) ensuing from a cataclysmic event, would rapidly lead to invasive cancer and metastases. Classical drivers (KRAS, CDKN2A, TP53, SMAD4) from the PanIN progression model are overlaid onto these models. Theoretical PanIN stages are shown as P1–P3.
Supplementary information
Supplementary Information
This file contains Supplementary Results, Supplementary Methods, Supplementary References, Supplementary Figures 1-18, Supplementary Tables 1-2 and Celluloid and Chrom-AL solutions data. (PDF 16401 kb)
Rights and permissions
About this article
Cite this article
Notta, F., Chan-Seng-Yue, M., Lemire, M. et al. A renewed model of pancreatic cancer evolution based on genomic rearrangement patterns. Nature 538, 378–382 (2016). https://doi.org/10.1038/nature19823
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19823
This article is cited by
-
Pancreatic cancer acquires resistance to MAPK pathway inhibition by clonal expansion and adaptive DNA hypermethylation
Clinical Epigenetics (2024)
-
Small extrachromosomal circular DNA harboring targeted tumor suppressor gene mutations supports intratumor heterogeneity in mouse liver cancer induced by multiplexed CRISPR/Cas9
Genome Medicine (2023)
-
Splicing alterations in pancreatic ductal adenocarcinoma: a new molecular landscape with translational potential
Journal of Experimental & Clinical Cancer Research (2023)
-
ACT-Discover: identifying karyotype heterogeneity in pancreatic cancer evolution using ctDNA
Genome Medicine (2023)
-
RBFOX2 deregulation promotes pancreatic cancer progression and metastasis through alternative splicing
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.