Abstract
In plants and algae, light serves both as the energy source for photosynthesis and a biological signal that triggers cellular responses via specific sensory photoreceptors. Red light is perceived by bilin-containing phytochromes and blue light by the flavin-containing cryptochromes and/or phototropins (PHOTs)1, the latter containing two photosensory light, oxygen, or voltage (LOV) domains2. Photoperception spans several orders of light intensity3, ranging from far below the threshold for photosynthesis to values beyond the capacity of photosynthetic CO2 assimilation. Excess light may cause oxidative damage and cell death, processes prevented by enhanced thermal dissipation via high-energy quenching (qE), a key photoprotective response4. Here we show the existence of a molecular link between photoreception, photosynthesis, and photoprotection in the green alga Chlamydomonas reinhardtii. We show that PHOT controls qE by inducing the expression of the qE effector protein LHCSR3 (light-harvesting complex stress-related protein 3) in high light intensities. This control requires blue-light perception by LOV domains on PHOT, LHCSR3 induction through PHOT kinase, and light dissipation in photosystem II via LHCSR3. Mutants deficient in the PHOT gene display severely reduced fitness under excessive light conditions, indicating that the sensing, utilization, and dissipation of light is a concerted process that plays a vital role in microalgal acclimation to environments of variable light intensities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jiao, Y., Lau, O. S. & Deng, X. W. Light-regulated transcriptional networks in higher plants. Nat. Rev. Genet. 8, 217–230 (2007)
Christie, J. M. Phototropin blue-light receptors. Annu. Rev. Plant Biol. 58, 21–45 (2007)
Briggs, W. R. Phototropism: some history, some puzzles, and a look ahead. Plant Physiol. 164, 13–23 (2014)
Li, Z., Wakao, S., Fischer, B. B. & Niyogi, K. K. Sensing and responding to excess light. Annu. Rev. Plant Biol. 60, 239–260 (2009)
Eberhard, S., Finazzi, G. & Wollman, F. A. The dynamics of photosynthesis. Annu. Rev. Genet. 42, 463–515 (2008)
Horton, P., Ruban, A. V. & Walters, R. G. Regulation of light harvesting in green plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 655–684 (1996)
Niyogi, K. K. & Truong, T. B. Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photosynthesis. Curr. Opin. Plant Biol. 16, 307–314 (2013)
Peers, G. et al. An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462, 518–521 (2009)
Tokutsu, R. & Minagawa, J. Energy-dissipative supercomplex of photosystem II associated with LHCSR3 in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 110, 10016–10021 (2013)
Ruban, A. V. et al. Identification of a mechanism of photoprotective energy dissipation in higher plants. Nature 450, 575–578 (2007)
Ahn, T. K. et al. Architecture of a charge-transfer state regulating light harvesting in a plant antenna protein. Science 320, 794–797 (2008)
Finazzi, G. & Minagawa, J. in Non-PHOTochemical Quenching and Energy Dissipation in Plants, Algae and Cyanobacteria (eds Demmig-Adams, B., Garab, G., Adams, W. III & Govindjee ) Ch 21, 445–469 (Springer Netherlands, 2014)
Petroutsos, D. et al. The chloroplast calcium sensor CAS is required for photoacclimation in Chlamydomonas reinhardtii. Plant Cell 23, 2950–2963 (2011)
Maruyama, S., Tokutsu, R. & Minagawa, J. Transcriptional regulation of the stress-responsive light harvesting complex genes in Chlamydomonas reinhardtii. Plant Cell Physiol. 55, 1304–1310 (2014)
Watanabe, M. et al. Design and performance of the Okazaki large spectrograph for photobiological research. Photochem. photobiol. 36, 491–498 (1982)
Beel, B. et al. A flavin binding cryptochrome photoreceptor responds to both blue and red light in Chlamydomonas reinhardtii. Plant Cell 24, 2992–3008 (2012)
Zorin, B., Lu, Y., Sizova, I. & Hegemann, P. Nuclear gene targeting in Chlamydomonas as exemplified by disruption of the PHOT gene. Gene 432, 91–96 (2009)
Merchant, S. S. et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318, 245–250 (2007)
Trippens, J. et al. Phototropin influence on eyespot development and regulation of phototactic behavior in Chlamydomonas reinhardtii. Plant Cell 24, 4687–4702 (2012)
Kong, S.-G. et al. The C-terminal kinase fragment of Arabidopsis phototropin 2 triggers constitutive phototropin responses. Plant J. 51, 862–873 (2007)
Harada, A., Sakai, T. & Okada, K. phot1 and phot2 mediate blue light-induced transient increases in cytosolic Ca2+ differently in Arabidopsis leaves. Proc. Natl Acad. Sci. USA 100, 8583–8588 (2003)
Babourina, O., Newman, I. & Shabala, S. Blue light-induced kinetics of H+ and Ca2+ fluxes in etiolated wild-type and phototropin-mutant Arabidopsis seedlings. Proc. Natl Acad. Sci. USA 99, 2433–2438 (2002)
Folta, K. M. & Kaufman, L. S. Phototropin 1 is required for high-fluence blue-light-mediated mRNA destabilization. Plant Mol. Biol. 51, 609–618 (2003)
Kirilovsky, D. & Kerfeld, C. A. The orange carotenoid protein: a blue-green light photoactive protein. Photochem. photobiol. Sci. 12, 1135–1143 (2013)
Schellenberger Costa, B. et al. Blue light is essential for high light acclimation and photoprotection in the diatom Phaeodactylum tricornutum. J. Exp. Bot. 64, 483–493 (2013)
Cazzaniga, S., Dall’ Osto, L., Kong, S. G., Wada, M. & Bassi, R. Interaction between avoidance of photon absorption, excess energy dissipation and zeaxanthin synthesis against photooxidative stress in Arabidopsis. Plant J. 76, 568–579 (2013)
Galván-Ampudia, C. S. & Offringa, R. Plant evolution: AGC kinases tell the auxin tale. Trends Plant Sci. 12, 541–547 (2007)
Huang, K. & Beck, C. F. Phototropin is the blue-light receptor that controls multiple steps in the sexual life cycle of the green alga Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 100, 6269–6274 (2003)
Im, C. S., Eberhard, S., Huang, K., Beck, C. F. & Grossman, A. R. Phototropin involvement in the expression of genes encoding chlorophyll and carotenoid biosynthesis enzymes and LHC apoproteins in Chlamydomonas reinhardtii. Plant J. 48, 1–16 (2006)
Gorman, D. S. & Levine, R. P. Cytochrome f and plastocyanin: their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardi. Proc. Natl Acad. Sci. USA 54, 1665–1669 (1965)
Sueoka, N. Mitotic replication of deoxyribonucleic acid in Chlamydomonas reinhardii. Proc. Natl Acad. Sci. USA 46, 83–91 (1960)
Allorent, G. et al. A dual strategy to cope with high light in Chlamydomonas reinhardtii. Plant Cell 25, 545–557 (2013)
Schloss, J. A. A Chlamydomonas gene encodes a G protein β subunit-like polypeptide. Mol. Gen. Genet. 221, 443–452 (1990)
Gera, J. F. & Baker, E. J. Deadenylation-dependent and -independent decay pathways for α1-tubulin mRNA in Chlamydomonas reinhardtii. Mol. Cell. Biol. 18, 1498–1505 (1998)
Takahashi, H., Iwai, M., Takahashi, Y. & Minagawa, J. Identification of the mobile light-harvesting complex II polypeptides for state transitions in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 103, 477–482 (2006)
Iwai, M. et al. Isolation of the elusive supercomplex that drives cyclic electron flow in photosynthesis. Nature 464, 1210–1213 (2010)
Longoni, P., Douchi, D., Cariti, F., Fucile, G. & Goldschmidt-Clermont, M. Phosphorylation of the light-harvesting complex II isoform Lhcb2 is central to state transitions. Plant Physiol. 169, 2874–2883 (2015)
Johnson, X. et al. A new setup for in vivo fluorescence imaging of photosynthetic activity. photosynth. Res. 102, 85–93 (2009)
Petroutsos, D. et al. PGRL1 participates in iron-induced remodeling of the photosynthetic apparatus and in energy metabolism in Chlamydomonas reinhardtii. J. Biol. Chem. 284, 32770–32781 (2009)
Genty, B., Briantais, J.-M. & Baker, N. R. The relationship between the quantum yield of photosynthetic electron transport and quenching of chlorophyll fluorescence. Biochim. Biophys. Acta 990, 87–92 (1989)
Depège, N., Bellafiore, S. & Rochaix, J. D. Role of chloroplast protein kinase Stt7 in LHCII phosphorylation and state transition in Chlamydomonas. Science 299, 1572–1575 (2003)
Bergner, S. V. et al. STATE TRANSITION7-dependent phosphorylation is modulated by changing environmental conditions, and its absence triggers remodeling of photosynthetic protein complexes. Plant Physiol. 168, 615–634 (2015)
Acknowledgements
We thank K. Kamada for help with genetic crossing, Y. Yari Kamrani for help with the Okazaki Large Spectrograph experiments and K. Kosuge for help with immunoblotting experiments. We thank G. Kreimer and Y. Aihara for discussions, M. Hippler for the antibody against LHCSR3 and G. Allorent for help during the initial phase of this project. We also thank P. Longoni and M. Goldschmidt-Clermont for advice and technical assistance with Phos-tag PAGE and T. Yamasaki for assistance with quantitative PCR. This work was supported by grants from Agence Nationale de la Recherche (ANR-12-BIME-0005, GRAL Labex, ANR-10-LABX-49-01, DiaDomOil to D.P. and G.F.); the Marie Curie Initial Training Network Accliphot (FP7-PEOPLE-2012-ITN; 316427 to G.F., D.P., S.F.); the CNRS Défi (ENRS 2013 to G.F. and D.P.); the CEA Bioénergies program (to G.F. and D.P.); the HFSP (HFSP0052) to G.F. and D.P.; the NIBB Cooperative Research Program for the Okazaki Large Spectrograph (13-514 and 14-508 to G.F. and 15-609 and 16-705 to J.M.); JSPS KAKENHI (JP15H05599 to R.T., JP26251033 and JP16H06553 to J.M. and R.T.); the NEDO (P07015 to J.M.); the MEXT (through the Network of Centres of Carbon Dioxide Resource Studies in Plants to J.M.); the German Research Foundation, DFG, grants FOR1261 (to P.H., M.M. and T.K.) and Heisenberg fellowship (to T.K.). L.M. acknowledges support from the Alexander von Humboldt Stiftung/Foundation.
Author information
Authors and Affiliations
Contributions
D.P., R.T., J.M. and G.F. designed the study. D.P., R.T., S.F. and L.M performed biochemical analyses; D.P. and R.T. performed fluorescence measurements; D.P. performed pharmacological tests; D.P., R.T., J.M. and G.F. performed action spectra; R.T. performed mutant generation; R.T. and S.M. performed quantitative PCR analysis; S.F. performed pigment quantification; L.C. performed the photosensitivity tests; J.M. and G.F. performed non-photochemical quenching experiments. D.P., R.T., S.M., A.G., L.M., T.K., M.M., P.H., J.M. and G.F. provided strains and analysed the data. D.P., R.T., J.M. and G.F. wrote the manuscript, and all authors revised and approved it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks J.-D. Rochaix and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Photosynthetic properties of acry and phot mutants.
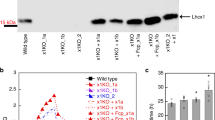
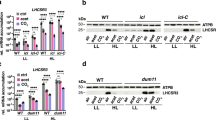
a, Immunoblot analysis of aCRY accumulation in wild-type (137c) and acry cells. The two strains were grown as previously described16. Cells were harvested at the beginning of the light phase (LD2 phase16). Whole-cell samples with 2 μg of chlorophyll were loaded on each lane. Representative dataset of an experiment replicated 3 times on different biological samples. b, Immunoblot analysis of PHOT in wild-type (cw15-302) and phot cells after 4 h exposure to high-intensity white light. Representative dataset of an experiment replicated 3 times on different biological samples. c, Immunoblot analysis of LHCSR3 and ATPB in wild-type (137c), phot and acry cells in low light intensity and after 4 h exposure to high-intensity white light (HL). ATPB was used as a loading control. Representative dataset of an experiment replicated five times on different biological samples. d, e, NPQ induction kinetics of wild-type (137c) and acry cells (d) and wild-type (cw15-302) and phot cells (e) after 4 h exposure to high-intensity white light. qE was recorded for 16 min upon illumination with 440 μmol photons m−2 s−1 (white bar) followed by 2 min of darkness (black bar), to measure qE relaxation. Representative dataset of an experiment replicated 4 (d) and 7 (e) times on different biological samples. f, g, DES indicates the xanthophyll cycle de-epoxidation state ([zeaxanthin] + 1/2[antheraxanthin])/([zeaxanthin] + [antheraxanthin] + [violaxanthin]) in wild-type (137c) and acry cells (f) and wild-type (cw15-302) and phot cells (g) after exposure to high light intensity for 4 h (n = 3 biological samples, mean ± s.d.). h, i, ETR of wild-type (137c) and acry (h) and wild-type (cw15-302) and phot (i) cells exposed at 20, 240 and 750 μmol photons m−2 s−1 for 10 h. ETR (μmol photons m−2 s−1) was measured at actinic illumination of 41 μmol photons m−2 s−1 (n = 3 biological samples, mean ± s.d.).
Extended Data Figure 2 Accumulation of major photosynthetic complexes is unaltered in the phot cells upon exposure to different wavelengths of visible light.
a, Immunoblot analyses of ATPB, PsaA/B, D1, D2 and CP26 accumulation in wild-type (137c), phot and acry cells after 4 h exposure to 250 μmol photons m−2 s−1 of monochromatic light at the different wavelengths of the visible spectrum. b, Immunoblot analyses of major photosynthetic complexes of PSII, cyt b6f and PSII in wild-type, phot and acry cells after 4 h exposure at 250 μmol photons m−2 s−1 of white light. Representative dataset of an experiment replicated three times on different biological samples.
Extended Data Figure 3 Diminished LHCSR3 induction in phot is not caused by diminished photosynthesis.
a, Comparison of ETR in DCMU-titrated wild-type cells and DCMU-untreated phot cells exposed to high light intensity for 3 h. ETR (μmol photons m−2 s−1) was measured upon exposure to light of 170 μmol photons m−2 s−1 (n = 3 biological samples, mean ± s.d.). b, Immunoblot analyses of LHCSR3 accumulations in the wild-type and phot samples described in a. ATPB was used as a loading control. Representative dataset of an experiment replicated five times on different biological samples.
Extended Data Figure 4 PHOT protein levels pigment content in phot mutants.
a, Immunoblot analyses of PHOT accumulation in wild-type (cw15-302) and PPHOT strains. PPHOT expresses a fused PHOT–BLE protein, which has a higher molecular mass than wild-type PHOT. ATPB served as a loading control. Representative dataset of experiment replicated four times on different biological samples. b, Total cellular chlorophyll (a + b) content in wild-type, phot, PPHOT, PKIN and PLOV cells exposed to light of 20, 200 or 750 μmol photons m−2 s−1 for 20 h as in Fig. 2d (n = 3 biological samples, mean ± s.d.). Diminished chlorophyll content is a signature of pigment bleaching following photo-damage.
Extended Data Figure 5 PHOT-independent LHCSR3 expression restores photoprotection in the phot mutant.
Relative qE and LHCSR3 expression under high-light-intensity conditions in wild-type (cw15-302), phot, and two phot-transformed lines expressing an additional copy of the LHCSR3.1 gene under the control of the PsaD promoter (PLHCSR3(B5) and PLHCSR3(G6)). ATPB was used as a loading control. qE values were normalized to wild-type cells (n = 3 biological samples, mean ± s.d.).
Extended Data Figure 6 Phosphorylation levels of LHCSR3 in phot, PPHOT, PLOV, PKIN, stt7 and wild-type cells.
Assessment of the phosphorylation levels of LHCSR3 in wild-type (cw15-302), phot, PPHOT, PLOV and PKIN cells by a mobility-shift detection of phosphorylated proteins (Phos-tag). The upper and lower bands correspond to the phosphorylated and non-phosphorylated forms of LHCSR3 (P-LHCSR3 and LHCSR3), respectively, as confirmed by treatment of the samples with either calf intestinal phosphatase (CIP) or λPP. The stt7 mutant41 was used to test the involvement of the chloroplastic serine/threonine kinase STT7 in the LHCSR3 phosphorylation. In this mutant, LHCSR3 was mostly present in the non-phosphorylated form, while an STT7-independent phosphorylation was also seen in agreement with recent findings42. Representative dataset of an experiment replicated three times on different biological samples.
Extended Data Figure 7 Phenotypic traits of the WTKIN genotype.
a, Immunoblot analyses of LHCSR3 accumulation after exposure to 240 μmol photons m−2 s−1 of white light for 20 h in wild-type (137c) and WTKIN cells. ATPB was used as a loading control. b, Erlenmeyer flasks containing wild-type and WTKIN (expressing the kinase domain of PHOT in the wild-type background) after 20 h exposure to white light of 20, 200 and 750 μmol photons m−2 s−1 and schematic drawings of the PHOT gene constructs in the two lines. Representative pictures from an experiment replicated three times on different biological samples. c, Action spectrum of LHCSR3 accumulation in wild-type and WTKIN. ATPB was used as a loading control. d, Densitometric quantification of LHCSR3 accumulation in wild-type (137c) and WTKIN (data normalized to ATPB). e, LHCSR3 protein accumulation in WTKIN at low and high light intensity in the absence and presence of the PSII inhibitor DCMU. Representative dataset of experiment replicated three times on different biological samples.
Extended Data Figure 8 Second messengers involved in the PHOT-dependent regulation of LHCSR3 expression.
a, Immunoblotting analyses of LHCSR3 accumulation after exposure to high-intensity white light for 20 h in wild-type (cw15-302) and phot cells under control conditions (0.34 mM Ca2+) or in the presence of increased Ca2+ concentration (3.4 mM Ca2+). Representative dataset of an experiment replicated three times on different biological samples. b, Statistical analyses of LHCSR3 accumulation in high-light-intensity-treated wild-type or phot cells in the absence (control) and presence of IBMX, DB-cGMP and DB-cAMP (see conditions as in Fig. 4a). Data are normalized to LHCSR3 levels of wild-type control cells (n = 5 biological samples, mean ± s.d.). Asterisks indicate statistically significant difference from wild-type control cells (t-test, P < 0.0001)
Extended Data Figure 9 LHCSR3.1 and LHCSR3.2 transcript stability in wild-type and phot cells.
Relative amounts of LHCSR3.1 and LHCSR3.2 mRNA in wild-type (cw15-302; black) and phot (red) cells were quantified by quantitative PCR. Cells exposed for 20 h to 470 nm LED light at 100 μmol photons m−2 s−1 were transfered to darkness at t = 0 and treated with actinomycin D to stop further mRNA synthesis. mRNA samples were collected at 0, 0.25, 0.5, 1, 2 and 4 h after transition to darkness. LHCSR3.1 and LHCSR3.2 transcript amounts were normalized to the amounts of 18S rRNA as endogenous control and their values were set at 100% at t = 0 (n = 3 biological samples, mean ± s.d.).
Extended Data Figure 10 qE and LHCSR3 induction requires high light in C. reinhardtii.
a, Action spectrum of qE induction in wild-type cells as a function of the light intensity. b, Immunoblot analysis of LHCSR3 accumulation in darkness (D) and under white (WL), blue (470 nm) and red (660 nm) light of low (20 μmol photons m−2 s−1) and high intensity (250 μmol photons m−2 s−1). ATPB was used as a loading control. Representative dataset of an experiment replicated two times on different biological samples.
Rights and permissions
About this article
Cite this article
Petroutsos, D., Tokutsu, R., Maruyama, S. et al. A blue-light photoreceptor mediates the feedback regulation of photosynthesis. Nature 537, 563–566 (2016). https://doi.org/10.1038/nature19358
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19358
This article is cited by
-
Macroscale structural changes of thylakoid architecture during high light acclimation in Chlamydomonas reinhardtii
Photosynthesis Research (2024)
-
Small Papillae Regulated by SPD25 are Critical for Balancing Photosynthetic CO2 Assimilation and Water Loss in Rice
Rice (2023)
-
Proteomic characterization of a lutein-hyperaccumulating Chlamydomonas reinhardtii mutant reveals photoprotection-related factors as targets for increasing cellular carotenoid content
Biotechnology for Biofuels and Bioproducts (2023)
-
Light-independent regulation of algal photoprotection by CO2 availability
Nature Communications (2023)
-
Widening the landscape of transcriptional regulation of green algal photoprotection
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.