Abstract
Ribosome-associated factors must properly decode the limited information available in nascent polypeptides to direct them to their correct cellular fate1. It is unclear how the low complexity information exposed by the nascent chain suffices for accurate recognition by the many factors competing for the limited surface near the ribosomal exit site2,3. Questions remain even for the well-studied cotranslational targeting cycle to the endoplasmic reticulum, involving recognition of linear hydrophobic signal sequences or transmembrane domains by the signal recognition particle (SRP)4,5. Notably, the SRP has low abundance relative to the large number of ribosome–nascent-chain complexes (RNCs), yet it accurately selects those destined for the endoplasmic reticulum6. Despite their overlapping specificities, the SRP and the cotranslationally acting Hsp70 display precise mutually exclusive selectivity in vivo for their cognate RNCs7,8. To understand cotranslational nascent chain recognition in vivo, here we investigate the cotranslational membrane-targeting cycle using ribosome profiling9 in yeast cells coupled with biochemical fractionation of ribosome populations. We show that the SRP preferentially binds secretory RNCs before their targeting signals are translated. Non-coding mRNA elements can promote this signal-independent pre-recruitment of SRP. Our study defines the complex kinetic interaction between elongation in the cytosol and determinants in the polypeptide and mRNA that modulate SRP–substrate selection and membrane targeting.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pechmann, S., Willmund, F. & Frydman, J. The ribosome as a hub for protein quality control. Mol. Cell 49, 411–421 (2013)
Bornemann, T., Holtkamp, W. & Wintermeyer, W. Interplay between trigger factor and other protein biogenesis factors on the ribosome. Nat. Commun. 5, 4180 (2014)
Nyathi, Y. & Pool, M. R. Analysis of the interplay of protein biogenesis factors at the ribosome exit site reveals new role for NAC. J. Cell Biol. 210, 287–301 (2015)
Akopian, D., Shen, K., Zhang, X. & Shan, S. O. Signal recognition particle: an essential protein-targeting machine. Annu. Rev. Biochem. 82, 693–721 (2013)
Zhang, X. & Shan, S. O. Fidelity of cotranslational protein targeting by the signal recognition particle. Annu. Rev. Biophys. 43, 381–408 (2014)
Ogg, S. C. & Walter, P. SRP samples nascent chains for the presence of signal sequences by interacting with ribosomes at a discrete step during translation elongation. Cell 81, 1075–1084 (1995)
Willmund, F. et al. The cotranslational function of ribosome-associated Hsp70 in eukaryotic protein homeostasis. Cell 152, 196–209 (2013)
del Alamo, M. et al. Defining the specificity of cotranslationally acting chaperones by systematic analysis of mRNAs associated with ribosome-nascent chain complexes. PLoS Biol. 9, e1001100 (2011)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Rapoport, T. A., Matlack, K. E., Plath, K., Misselwitz, B. & Staeck, O. Posttranslational protein translocation across the membrane of the endoplasmic reticulum. Biol. Chem. 380, 1143–1150 (1999)
Ast, T., Cohen, G. & Schuldiner, M. A network of cytosolic factors targets SRP-independent proteins to the endoplasmic reticulum. Cell 152, 1134–1145 (2013)
Ng, D. T., Brown, J. D. & Walter, P. Signal sequences specify the targeting route to the endoplasmic reticulum membrane. J. Cell Biol. 134, 269–278 (1996)
Walter, P. & Johnson, A. E. Signal sequence recognition and protein targeting to the endoplasmic reticulum membrane. Annu. Rev. Cell Biol. 10, 87–119 (1994)
Kutay, U., Ahnert-Hilger, G., Hartmann, E., Wiedenmann, B. & Rapoport, T. A. Transport route for synaptobrevin via a novel pathway of insertion into the endoplasmic reticulum membrane. EMBO J. 14, 217–223 (1995)
Williams, C. C., Jan, C. H. & Weissman, J. S. Targeting and plasticity of mitochondrial proteins revealed by proximity-specific ribosome profiling. Science 346, 748–751 (2014)
Jan, C. H., Williams, C. C. & Weissman, J. S. Principles of ER cotranslational translocation revealed by proximity-specific ribosome profiling. Science 346, 1257521 (2014)
Lakkaraju, A. K., Mary, C., Scherrer, A., Johnson, A. E. & Strub, K. SRP keeps polypeptides translocation-competent by slowing translation to match limiting ER-targeting sites. Cell 133, 440–451 (2008)
Mason, N., Ciufo, L. F. & Brown, J. D. Elongation arrest is a physiologically important function of signal recognition particle. EMBO J. 19, 4164–4174 (2000)
Pechmann, S., Chartron, J. W. & Frydman, J. Local slowdown of translation by nonoptimal codons promotes nascent-chain recognition by SRP in vivo. Nat. Struct. Mol. Biol. 21, 1100–1105 (2014)
Lu, J. & Deutsch, C. Electrostatics in the ribosomal tunnel modulate chain elongation rates. J. Mol. Biol. 384, 73–86 (2008)
Woolstenhulme, C. J., Guydosh, N. R., Green, R. & Buskirk, A. R. High-precision analysis of translational pausing by ribosome profiling in bacteria lacking EFP. Cell Reports 11, 13–21 (2015)
Voorhees, R. M. & Hegde, R. S. Structures of the scanning and engaged states of the mammalian SRP-ribosome complex. eLife 4, e07975 (2015)
Berndt, U., Oellerer, S., Zhang, Y., Johnson, A. E. & Rospert, S. A signal-anchor sequence stimulates signal recognition particle binding to ribosomes from inside the exit tunnel. Proc. Natl Acad. Sci. USA 106, 1398–1403 (2009)
Hainzl, T. & Sauer-Eriksson, A. E. Signal-sequence induced conformational changes in the signal recognition particle. Nat. Commun. 6, 7163 (2015)
Holtkamp, W. et al. Dynamic switch of the signal recognition particle from scanning to targeting. Nat. Struct. Mol. Biol. 19, 1332–1337 (2012)
Loya, A. et al. The 3′-UTR mediates the cellular localization of an mRNA encoding a short plasma membrane protein. RNA 14, 1352–1365 (2008)
Gao, X. et al. Quantitative profiling of initiating ribosomes in vivo. Nat. Methods 12, 147–153 (2015)
Kraut-Cohen, J. et al. Translation- and SRP-independent mRNA targeting to the endoplasmic reticulum in the yeast Saccharomyces cerevisiae. Mol. Biol. Cell 24, 3069–3084 (2013)
Naranda, T., MacMillan, S. E. & Hershey, J. W. Purified yeast translational initiation factor eIF-3 is an RNA-binding protein complex that contains the PRT1 protein. J. Biol. Chem. 269, 32286–32292 (1994)
Elvekrog, M. M. & Walter, P. Dynamics of co-translational protein targeting. Curr. Opin. Chem. Biol. 29, 79–86 (2015)
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003)
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003)
Zhong, T. & Arndt, K. T. The yeast SIS1 protein, a DnaJ homolog, is required for the initiation of translation. Cell 73, 1175–1186 (1993)
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999)
Freidlin, P. J. & Patterson, R. J. Heparin releases monosomes and polysomes from rough endoplasmic reticulum. Biochem. Biophys. Res. Commun. 93, 521–527 (1980)
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protocols 7, 1534–1550 (2012)
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10 (2011)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013)
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013)
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015)
Bendtsen, J. D., Nielsen, H., von Heijne, G. & Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340, 783–795 (2004)
Käll, L., Krogh, A. & Sonnhammer, E. L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 338, 1027–1036 (2004)
Reynolds, S. M., Käll, L., Riffle, M. E., Bilmes, J. A. & Noble, W. S. Transmembrane topology and signal peptide prediction using dynamic bayesian networks. PLOS Comput. Biol. 4, e1000213 (2008)
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001)
Breker, M., Gymrek, M., Moldavski, O. & Schuldiner, M. LoQAtE–Localization and Quantitation ATlas of the yeast proteomE. A new tool for multiparametric dissection of single-protein behavior in response to biological perturbations in yeast. Nucleic Acids Res. 42, D726–D730 (2014)
Becker, A. H., Oh, E., Weissman, J. S., Kramer, G. & Bukau, B. Selective ribosome profiling as a tool for studying the interaction of chaperones and targeting factors with nascent polypeptide chains and ribosomes. Nat. Protocols 8, 2212–2239 (2013)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Beckert, B. et al. Translational arrest by a prokaryotic signal recognition particle is mediated by RNA interactions. Nat. Struct. Mol. Biol. 22, 767–773 (2015)
Murtagh, F. & Legendre, P. Ward’s hierarchical agglomerative clustering method: which algorithms implement ward’s criterion? J. Classif. 31, 274–295 (2014)
Pédelacq, J. D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006)
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009)
Hessa, T. et al. Recognition of transmembrane helices by the endoplasmic reticulum translocon. Nature 433, 377–381 (2005)
Pechmann, S. & Frydman, J. Evolutionary conservation of codon optimality reveals hidden signatures of cotranslational folding. Nat. Struct. Mol. Biol. 20, 237–243 (2013)
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011)
Doerfel, L. K. & Rodnina, M. V. Elongation factor P: Function and effects on bacterial fitness. Biopolymers 99, 837–845 (2013)
Acknowledgements
We thank P. Walter, J. S. Weissman and C. Jan for discussions; R. Andino and R. Hegde for critical reading of the manuscript. We thank S. Pechmann, K. M. Dalton, E. M. Sontag, P. T. Dolan and other members of the Frydman laboratory for advice on analysis. Sequencing was performed at the UCSF Center for Advanced Technology with assistance from E. Chow, J. Lund and A. Acevedo. J.W.C. is supported by an NIH NRSA award. This work was additionally supported by grants to J.F. from the NIH and HFSP.
Author information
Authors and Affiliations
Contributions
J.W.C. and J.F. designed the study. K.C.L.H. performed experiments with prt1-1. J.W.C. performed all other experiments and analysis. J.W.C. and J.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks R. Keenan and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Cotranslational membrane enrichment.
a, Crude lysates were fractionated, and then polysomes were recovered by sucrose gradient ultracentrifugation and used for ribosome profiling. b, Enrichment of ribosome-protected mRNA reads in the membrane polysome fractions over the soluble polysome fractions from two biological replicates. Every dot represents one ORF. c, Metagene plots of soluble polysome ribosome-protected reads of transcripts encoding proteins lacking ER-targeting signals (top), or of membrane-bound polysome-protected reads of transcripts encoding secretory proteins that were at least twofold membrane-enriched (bottom). For each ORF, ribosome-protected reads at each position were scaled by dividing by the mean reads per codon of the ORF, excluding the first two and last two sense codons. The median scaled reads at each position are plotted as a line, and the interquartile range is shaded in grey. d, Ribosome-protected reads at each codon of an example secreted protein, β-1,3-glucanosyltransferase (GAS1), a model SRP-independent protein12. Topology is indicated above, with the signal sequence in lavender. The position where the signal begins to emerge from the ribosome exit tunnel is indicated. e, The number of codons remaining after the encoding of the first residue of an SS, and the corresponding membrane enrichment per SS-containing ORF. Signal sequences were divided between those that bind Ssh1p directly upon exposure and those that require a looped conformation (>90 codons after the first SS codon)16. f, Transcripts remain at the membrane by subsequent translocon binding, thus the small soluble fraction comprises mRNA undergoing initial targeting.
Extended Data Figure 2 Cotranslational enrichment of SRP.
a, Enrichment of ribosome-protected mRNA reads in the soluble SRP-bound polysome fractions over the total soluble polysome fractions from two biological replicates. b, The number of codons remaining after encoding of first SS or TMD residue, and the corresponding SRP and membrane enrichment scores per ORF. Scores are determined from cultures harvested without added CHX. Enrichment scores are indicated with filled dots, and the scores from the same transcript are linked with a grey line. The vertical dashed line indicates 50 codons, the boundary for tail-anchored proteins. Here, only SSs that bind Ssh1p directly after exposure from the RNC are shown. c, Secretory transcripts were classified into two groups based on the ribosome-protected-read distributions from SRP-bound polysomes. Some showed a pronounced increase in reads at positions coincident with the initial exposure of an SS or TMD by the ribosome, whereas others did not. Shown here are metagene analysis plots of soluble polysome-protected reads from the categorized TMD proteins. For each ORF, the reads at each codon position were divided by the mean reads per codon within the range +20 to +40 after the first signal codon. The first 30 codons of each ORF are excluded to avoid the characteristic low-density region near the start codon. The lavender line indicates when the first TMD begins to emerge from the exit tunnel, and the dashed line indicates the position of the read peak. Notably, the total soluble polysome reads depleted in a similar manner for both classes, a read increase was not observed in the total soluble reads, and reads from the SRP-bound transcripts with a peak did not deplete faster than the total soluble reads. These features are consistent with a model in which SRP is recruited at the peak site, and elongation then proceeds at the same rate. d, The number of codons remaining after encoding of the first SS or TMD and corresponding SRP enrichment. Transcripts are classified by the presence or absence of a read increase following signal exposure, as in c. Note that for SRP-enriched transcripts with signals closest to the terminus (<100 codons), evidence of direct binding between SRP and the nascent chain was always observed. SRP can therefore bind late TMDs immediately after they become exposed by the ribosome. e, Maximum hydrophobicity across targeting signals using an 8-residue averaging window. Only signals with peaks that could be unambiguously attributed to a targeting signal were included. Hydrophobicity was determined by attributing the biological hydrophobicity score to each encoded amino acid54. ***P ≤ 0.001, Wilcoxon rank-sum test. f, Distribution of the distance between the first codon of a targeting signal and the position of the downstream read increase. Only transcripts wherein the increase can be unambiguously attributed to a specific targeting signal were included.
Extended Data Figure 3 Elongation pausing and local SRP recruitment.
a, b, Local increases in ribosome-protected reads from membrane-bound polysomes, indicated by orange lines, were coincident with rare codons, as in cell division cycle protein 1 (CDC1, a) or polybasic nascent chains, as in the plasma membrane G-protein-coupled receptor (GPR1, b). Soluble SRP-bound polysome-protected reads were further increased at the same positions. c, In these cases, hydrophobic sequences in the nascent chain were exposed to the cytosol at the locations of increased reads, which were coincident with elongation attenuators. d, Translational efficiencies for the 6 codons following, and the number of stalling residues within the 10 residues preceding, the sites of increased SRP-bound ribosome reads. Translational efficiency was determined by attributing the normalized translational efficiency (nTE) score to each codon55. Residues that were found to stall the ribosome, based on previous investigation20,56,57, were lysine, arginine, glutamate, aspartate, proline and glycine. Because of variation in specific motifs, and uncertainty in whether these motifs are additive, we simply compared the total number of these residues in the indicated 10 residue spans. Sets of 10,000 random sequences, at least 10 amino acids from the stop codon, were sampled from 5,907 non-dubious ORFs, and translational efficiency and stalling residues were determined over 6 or 10 codon spans. *P ≤ 0.05, **P ≤ 0.01, Wilcoxon rank-sum tests. e, The targeting signals that recruited SRP directly to the nascent chain unusually far from the encoding of the signal had SRP-binding sites coincident with intrinsic elongation attenuation. Secretory protein transcripts that showed an increase in SRP-bound protected reads (see Extended Data Fig. 2c, f) were further classified by the position of the peak relative to the first signal codon. Transcripts with peaks found at least 80 codons after the signal had significantly lower translational efficiency in the 6 codons following the peak. These transcripts also had a greater, but not statistically significant, amount of stalling amino acids in the 10 residues preceding the peak. *P ≤ 0.05, Wilcoxon rank-sum tests. f, Similar increases in SRP-bound reads were observed for certain non-secretory proteins as exemplified by phosphoacetylglucosamine mutase (PCM1) and tRNASer Um44 2′-O-methyltransferase (TRM44). Hydrophobic sequences in non-secretory proteins, coupled with attenuation of elongation, may lead to SRP recruitment.
Extended Data Figure 4 Ribosome profiling of monosomes.
a, Ribosomes transition from monosomes to polysomes during elongation. The pioneer round of initiation will be a monosome, and during elongation there is a chance of additional initiation converting the transcript to a polysome. Similarly, a polysome will become a monosome if all ribosomes but one terminate. As mRNA is sampled closer to the stop codon, the likelihood of observing a footprint from the final ribosome will increase. b, Metagene analysis of soluble monosome- or polysome-protected reads from proteins lacking an ER-targeting signal. Data were obtained using CHX treatment. ORFs are at least 400 codons long and have an average of at least 0.5 reads per codon in each data set. For each ORF, ribosome reads at each position were divided by the mean reads per codon over the range +160 to +240 codons. The median normalized read value at each codon position is plotted, and the interquartile range is shaded in grey. c, Relative reads at the start codon from ORFs normalized in b. d, Distributions of the ratio of ribosome-protected reads found in soluble monosomes over soluble polysomes. e, A pioneer round of translation deposits mRNA on the membrane. Polysomes will be retained at the membrane and are therefore depleted from the soluble fraction.
Extended Data Figure 5 Ribosome profiling of SRP-bound monosomes.
a, Ribosome-protected reads, in tags per million (TPM) for each ORF, from SRP-bound monosome fractions from two biological replicates. b, Ribosome-protected reads from the soluble SRP-bound monosome and SRP-bound polysome fractions of the same biological replicate, with CHX treatment. c, Distribution of ribosome reads within example ORFs that display SRP-bound monosome and polysome profiles consistent with direct recognition of the nascent chain. d, If RNCs can recruit SRP while a TMD is within the exit tunnel, then there will be an increase in ribosome-protected reads from SRP-bound monosomes when the TMD begins to translate (lavender). This increase will maximize when the TMD is exposed to the cytosol (orange). e, Distribution of ribosome reads within example ORFs that display SRP-bound monosome profiles consistent with recruitment to transcripts before targeting signal synthesis. Examples are arranged for an increasing distance from the start codon to the first TMD. Only the first 600 codons for each ORF are shown.
Extended Data Figure 6 The role of the UTR from PMP1 and PMP2.
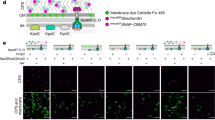
a, The cotranslational SRP enrichment of the PMP1 and PMP2 ORFs was similar to other bona fide secretory proteins, such as SEC61. By contrast, cytosolic proteins such as tubulin (TUB2) were not enriched. The enrichment scores are determined from the SRP-bound and total soluble polysomes from two biological replicates collected without added CHX. b, Distribution of ribosome-protected reads from soluble polysomes within the PMP1 and PMP2 ORFs. c, Membrane enrichment, determined by qPCR, of the mRNA of GFP fused to the indicated 3′ UTRs. The coding sequence of endogenous SEC61 transcript was also amplified as a control for a membrane-localized transcript. **P ≤ 0.01, n = 3 biological replicates, Welch’s t-test. d, Localization of mature GFP. Scale bar, 5 μm. Yeast were grown to mid-log phase and imaged using an Axio Observer Z1 with a Plan-Apochromat 100 × /1.4 oil immersion objective (Zeiss). Z-stacks were deconvoluted by the iterative maximum likelihood algorithm in ZEN (Zeiss) and single planes are shown. Images were representative from a set of two replicated assays. e, Yeast growth after replacement of the endogenous 3′ UTR of PMP1 with the 3′ UTR of tubulin. Also shown is a complete deletion of PMP1 ORF34. Gibson assembly53 was used to fuse the 300-nucleotide TUB2 3′ UTR to the KlURA3 cassette into SmaI digested pUC19. The TUB2-UTR-URA3 element was PCR amplified, including 40-nucleotide overhangs matching genomic sequences, and replaced the 650 nucleotides immediately following the PMP1 coding sequence in strain BY4741 by homologous recombination. Image is representative from a set of 3 replicated assays. f, Nascent-chain-independent SRP recognition may require ribosomes. Puromycin treatment of lysates disrupts elongating, but not initiating, ribosomes. g, Transcripts showing only canonical recognition are more sensitive to puromycin. This is consistent with puromycin resistance of SRP that has pre-recruited to initiating ribosomes. h, Membrane enrichment of the GFP-PMP1 construct or SEC61 mRNA after lysates were incubated with puromycin. **P ≤ 0.01, n = 3 biological replicates, Welch’s t-test.
Extended Data Figure 7 The role translation in membrane enrichment.
a, Lysates were treated with puromycin before membrane fractionation. mRNA recovered from the soluble and membrane fractions were used for RNA-seq b, Membrane enrichment of secretory protein transcripts (SS, TMD, SS-TMD, or TA, n = 729) following puromycin treatment of lysates.
Supplementary information
Supplementary Discussion
This file contains a Supplementary Discussion and additional references. (PDF 355 kb)
Supplementary Table 1
This file contains gene annotations, enrichment scores, and sequencing counts derived from sequencing experiments. The first tab provides descriptions for the data presented in the second tab. The third and fourth tabs provide raw sequencing counts from the Ribo-seq and RNA-seq samples. (XLSX 2039 kb)
Rights and permissions
About this article
Cite this article
Chartron, J., Hunt, K. & Frydman, J. Cotranslational signal-independent SRP preloading during membrane targeting. Nature 536, 224–228 (2016). https://doi.org/10.1038/nature19309
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19309
This article is cited by
-
The ER-SURF pathway uses ER-mitochondria contact sites for protein targeting to mitochondria
EMBO Reports (2024)
-
Circular RNA circGlis3 protects against islet β-cell dysfunction and apoptosis in obesity
Nature Communications (2023)
-
Selectivity of mRNA degradation by autophagy in yeast
Nature Communications (2021)
-
Comparison of RNA isolation methods on RNA-Seq: implications for differential expression and meta-analyses
BMC Genomics (2020)
-
Signal recognition particle prevents N-terminal processing of bacterial membrane proteins
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.