Abstract
Regulation of messenger RNA translation is central to eukaryotic gene expression control1. Regulatory inputs are specified by the mRNA untranslated regions (UTRs) and often target translation initiation. Initiation involves binding of the 40S ribosomal small subunit (SSU) and associated eukaryotic initiation factors (eIFs) near the mRNA 5′ cap; the SSU then scans in the 3′ direction until it detects the start codon and is joined by the 60S ribosomal large subunit (LSU)2,3,4,5 to form the 80S ribosome. Scanning and other dynamic aspects of the initiation model have remained as conjectures because methods to trap early intermediates were lacking. Here we uncover the dynamics of the complete translation cycle in live yeast cells using translation complex profile sequencing (TCP-seq), a method developed from the ribosome profiling6 approach. We document scanning by observing SSU footprints along 5′ UTRs. Scanning SSU have 5′-extended footprints (up to ~75 nucleotides), indicative of additional interactions with mRNA emerging from the exit channel, promoting forward movement. We visualized changes in initiation complex conformation as SSU footprints coalesced into three major sizes at start codons (19, 29 and 37 nucleotides). These share the same 5′ start site but differ at the 3′ end, reflecting successive changes at the entry channel from an open to a closed state following start codon recognition. We also observe SSU ‘lingering’ at stop codons after LSU departure. Our results underpin mechanistic models of translation initiation and termination, built on decades of biochemical and structural investigation, with direct genome-wide in vivo evidence. Our approach captures ribosomal complexes at all phases of translation and will aid in studying translation dynamics in diverse cellular contexts. Dysregulation of translation is common in disease and, for example, SSU scanning is a target of anti-cancer drug development7. TCP-seq will prove useful in discerning differences in mRNA-specific initiation in pathologies and their response to treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
The sequences of reads with poly(A) tracts were deposited to the Sequence Read Archive, under accession code SRP074093. An online interface for browsing and analysing the TCP-seq data is available at (http://bioapps.erc.monash.edu/TCP/) along with (http://dx.doi.org/10.6084/m9.figshare.3206725). The underlying mapped dataset is also available (http://dx.doi.org/10.6084/m9.figshare.3206698).
References
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011)
Hinnebusch, A. G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 83, 779–812 (2014)
Jackson, R. J., Hellen, C. U. & Pestova, T. V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nature Rev. Mol. Cell Biol. 11, 113–127 (2010)
Hershey, J. W., Sonenberg, N. & Mathews, M. B. Principles of translational control: an overview. Cold Spring Harb. Perspect. Biol . 4, a011528 (2012)
Merrick, W. C. Eukaryotic protein synthesis: still a mystery. J. Biol. Chem. 285, 21197–21201 (2010)
Ingolia, N. T. Ribosome profiling: new views of translation, from single codons to genome scale. Nature Rev. Genet. 15, 205–213 (2014)
Bhat, M. et al. Targeting the translation machinery in cancer. Nature Rev. Drug Discov . 14, 261–278 (2015)
Kozak, M. How do eucaryotic ribosomes select initiation regions in messenger RNA? Cell 15, 1109–1123 (1978)
Paek, K. Y. et al. Translation initiation mediated by RNA looping. Proc. Natl. Acad. Sci. USA 112, 1041–1046 (2015)
Valásek, L., Szamecz, B., Hinnebusch, A. G. & Nielsen, K. H. In vivo stabilization of preinitiation complexes by formaldehyde cross-linking. Methods Enzymol. 429, 163–183 (2007)
Li, Z. et al. Rational extension of the ribosome biogenesis pathway using network-guided genetics. PLoS Biol. 7, e1000213 (2009)
Lareau, L. F., Hite, D. H., Hogan, G. J. & Brown, P. O. Distinct stages of the translation elongation cycle revealed by sequencing ribosome-protected mRNA fragments. eLife 3, e01257 (2014)
Kozak, M. & Shatkin, A. J. Migration of 40S ribosomal subunits on messenger RNA in the presence of edeine. J. Biol. Chem. 253, 6568–6577 (1978)
Siridechadilok, B., Fraser, C. S., Hall, R. J., Doudna, J. A. & Nogales, E. Structural roles for human translation factor eIF3 in initiation of protein synthesis. Science 310, 1513–1515 (2005)
Spirin, A. S. How does a scanning ribosomal particle move along the 5′-untranslated region of eukaryotic mRNA? Brownian Ratchet model. Biochemistry 48, 10688–10692 (2009)
Pöyry, T. A., Kaminski, A. & Jackson, R. J. What determines whether mammalian ribosomes resume scanning after translation of a short upstream open reading frame? Genes Dev. 18, 62–75 (2004)
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010)
Gilbert, W. V., Zhou, K., Butler, T. K. & Doudna, J. A. Cap-independent translation is required for starvation-induced differentiation in yeast. Science 317, 1224–1227 (2007)
Llácer, J. L. et al. Conformational differences between open and closed states of the eukaryotic translation initiation complex. Mol. Cell 59, 399–412 (2015)
Saini, A. K. et al. Eukaryotic translation initiation factor eIF5 promotes the accuracy of start codon recognition by regulating Pi release and conformational transitions of the preinitiation complex. Nucleic Acids Res. 42, 9623–9640 (2014)
Lomakin, I. B. & Steitz, T. A. The initiation of mammalian protein synthesis and mRNA scanning mechanism. Nature 500, 307–311 (2013)
Luna, R. E. et al. The C-terminal domain of eukaryotic initiation factor 5 promotes start codon recognition by its dynamic interplay with eIF1 and eIF2β. Cell Rep. 1, 689–702 (2012)
Yamamoto, H. et al. Structure of the mammalian 80S initiation complex with initiation factor 5B on HCV-IRES RNA. Nature Struct. Mol. Biol . 21, 721–727 (2014)
Fernández, I. S. et al. Molecular architecture of a eukaryotic translational initiation complex. Science 342, 1240585 (2013)
Becker, T. et al. Structural basis of highly conserved ribosome recycling in eukaryotes and archaea. Nature 482, 501–506 (2012)
Chen, Z. Q. et al. The essential vertebrate ABCE1 protein interacts with eukaryotic initiation factors. J. Biol. Chem. 281, 7452–7457 (2006)
Khoshnevis, S. et al. The iron-sulphur protein RNase L inhibitor functions in translation termination. EMBO Rep. 11, 214–219 (2010)
Roque, S. et al. Interaction between the poly(A)-binding protein Pab1 and the eukaryotic release factor eRF3 regulates translation termination but not mRNA decay in Saccharomyces cerevisiae. RNA 21, 124–134 (2015)
Jackson, R. J., Hellen, C. U. & Pestova, T. V. Termination and post-termination events in eukaryotic translation. Adv. Protein Chem. Struct. Biol. 86, 45–93 (2012)
Kurata, S. et al. Possible steps of complete disassembly of post-termination complex by yeast eEF3 deduced from inhibition by translocation inhibitors. Nucleic Acids Res. 41, 264–276 (2013)
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003)
Kressler, D. et al. Synchronizing nuclear import of ribosomal proteins with ribosome assembly. Science 338, 666–671 (2012)
Archer, S. K., Shirokikh, N. E., Hallwirth, C. V., Beilharz, T. H. & Preiss, T. Probing the closed-loop model of mRNA translation in living cells. RNA Biol. 12, 248–254 (2015)
Archer, S. K., Shirokikh, N. E. & Preiss, T. Probe-directed degradation (PDD) for flexible removal of unwanted cDNA sequences from RNA-seq libraries. Curr. Protoc. Hum. Genet. 85, 11 15 11–11 15 36 (2015)
Archer, S. K., Shirokikh, N. E. & Preiss, T. Selective and flexible depletion of problematic sequences from RNA-seq libraries at the cDNA stage. BMC Genomics 15, 401 (2014)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Zhang, Z., Hesselberth, J. R. & Fields, S. Genome-wide identification of spliced introns using a tiling microarray. Genome Res. 17, 503–509 (2007)
O’Connor, P., Andreev, D. & Baranov, P. Surveying the relative impact of mRNA features on local ribosome profiling read density in 28 datasets. bioRxiv http://dx.doi.org/10.1101/018762 (2015)
Artieri, C. G. & Fraser, H. B. Accounting for biases in riboprofiling data indicates a major role for proline in stalling translation. Genome Res. 24, 2011–2021 (2014)
McManus, C. J., May, G. E., Spealman, P. & Shteyman, A. Ribosome profiling reveals post-transcriptional buffering of divergent gene expression in yeast. Genome Res. 24, 422–430 (2014)
Chan, P. P. & Lowe, T. M. GtRNAdb: a database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 37, D93–D97 (2009)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Pop, C. et al. Causal signals between codon bias, mRNA structure, and the efficiency of translation and elongation. Mol. Syst. Biol. 10, 770 (2014)
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008)
Acknowledgements
This work was supported by an ARC Discovery Grant (DP1300101928) and an NHMRC Senior Research Fellowship (514904) awarded to T.P. N.E.S. was supported by a Go8 European Fellowship. We are grateful to A. G. Hinnebusch, C. G. Proud and R. D. Hannan for discussions and suggestions for this work. We acknowledge technical support from the Australian Cancer Research Foundation Biomolecular Resource Facility (JCSMR, ANU), D. Powell and S. Androulakis at the Monash Bioinformatics Platform.
Author information
Authors and Affiliations
Contributions
S.K.A., T.H.B. and T.P. designed the research, S.K.A. and N.E.S. performed the experiments, S.K.A, N.E.S., T.H.B. and T.P. analysed the data, discussed the results and wrote the paper.
Corresponding author
Additional information
Reviewer Information Nature thanks E. Alkalaeva, P. Baranov and Y. Mechulam for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Current model of eukaryotic translation with emphasis on substeps with scarce, missing or ambiguous evidence.
a, Schematic representing the current model of the translation cycle starting from SSU attachment to mRNA (top) and ending with recycling of the post-termination complexes on mRNA (bottom). Each step represents complexes with existing in vivo or in vitro evidence. b–e, Alternative models suggested for different substeps, with key differences highlighted in red. b, Type of scanning motion to locate the cognate start codon: linear unidirectional motion of SSU along the 5′ UTR (top) or random hopping over 5′ UTR regions from mRNA 5′-end (bottom). c, Mode of SSU attachment during scanning: cap-severed mRNA scanning in which initiating SSU lose their interaction with the 5′ cap structure as they encroach into the inner 5′ UTR regions (top), as opposed to cap-tethered scanning where the SSU 5′-end cap connection is preserved at least until start codon recognition. d, Position of the scanning factors relative to the SSU: SSU-led scanning in which initiation factors responsible for the scanning motion are located and interact with mRNA behind the SSU to push it in 5′→3′ direction (top), or an SSU-trailing mechanism where these factors interact with mRNA in front of the moving SSU to pull it along. e, Alternative recycling of termination complexes: mRNA is released from terminating ribosomes before they are disassembled into subunits (left), in contrast to a staged recycling where the LSU is first ejected from the post-termination ribosomes, leaving post-termination SSU attached to mRNA (right).
Extended Data Figure 2 Overview of the in vivo distribution of translation complexes on mRNAs measured by TCP-seq.
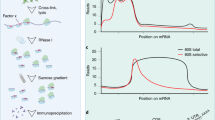
a, Snapshots of translation as represented by footprint 5′ ends on mRNA from wild-type strain ribosomes and SSU fractions (middle and bottom panels), total input mRNA is shown for reference (top panel). Footprint reads are mapped to all annotated mRNAs up to 1,000 nt of mRNA 5′ and 3′ flanking regions. Footprint 5′ end positions are plotted (horizontal axis) against the ORF length rank (vertical axis). Horizontal location (both within and outside the ORF) is scaled relative to ORF length, with start and stop codon positions indicated at the first nucleotide of these triplets (red lines). Footprints with 5′ ends accumulating at a constant distance upstream from start and stop appear as curved vertical trails due to the scaling. Transparency of points is increased by log10(2 + read number per transcript) to allow the distribution in less abundant transcripts to be observed. b, c, Same as a, but the horizontal location is scaled to 5′ (b) or 3′ UTR (c) length, and footprint 5′ ends are plotted against 5′ or 3′UTR length, respectively. d, Scheme showing classification of the footprints based on position along mRNAs for e, f. e, Profiles of the frequency of the footprint lengths in the 5′ UTR, start codon, ORF, stop codon and 3′ UTR regions for the ribosome and SSU fractions of the wild-type yeast. Only footprints mapping to mRNAs within annotated UTRs are analysed. The maximum frequency value for each plot is scaled to 1. f, Proportions of the footprints in the different mRNA regions for the ribosome and SSU libraries of wild-type and SYO1 yeast strains. For raw read counts, see Supplementary Table 1.
Extended Data Figure 3 Characteristics of start-codon-associated, stop-codon-associated and ORF-associated ribosome footprints and comparison of TCP-seq with ribosome profiling.
a, Aggregated densities of wild-type yeast footprint 5′ ends (FP-5′s; red) and footprint 3′ ends (FP-3′s; blue) from the ribosome fraction, mapped to all annotated mRNAs and shown within the vicinity of the start (top) or end (bottom) of the ORFs (aligned as in Fig. 4a, c). Positions of the major footprint end density modalities are indicated (dashed lines). The error regions shown as shading of the same colour indicate the 95th percentiles of 1,000 iterations of random gene-wise resampling (see Methods for details). b, Schematics illustrating the calculated position and 3′ and 5′ footprint extensions for the major ribosome footprint types (alignment as for the SSU in Fig. 4b, d) across the start (top) or stop (bottom) codons. c, Correlation of the ribosome density on mRNAs in exponentially propagating wild-type yeast as measured by TCP-seq (vertical axis) or translation (ribosome) profiling43 (horizontal axis). Note that total input denotes either total polysomal RNA (TCP-seq) or total cellular mRNA (ribosome profiling). d, End density for ribosome footprint 3′ ends plotted relative to the last sense codon from all ORFs, only for footprints with 5′ ends within the main peak for terminating ribosome complexes (from −14 nt to −10 nt). Error regions are depicted as in a. e, Same as in Fig. 2a (top panel), but showing ORF region from +20 to +120 nt from the start codon. f, Register bias (triplet periodicity) for 5′ and 3′ ends of ORF-associated SSU and ribosome footprints. Footprints falling between +10 and +200 nt from the start codons of all mRNAs are binned into one of three registers according to the remainder when dividing the footprint 5′ or 3′ ends position (relative to the ORF start) by three. The relative difference between the frequency of the most common and least common register is 16.5% for ribosome and 9.9% for SSU. The SSU register is shifted 1 nt downstream to that of the ribosome, possibly due to a 1 nt recession in the footprint 5′ ends after LSU dissociation.
Extended Data Figure 4 Assessment of potential library or sequence bias.
a, Comparison of sequence bias in the TCP-seq ribosome fraction to other ribosomal profiling experiments in yeast39,40, using codon-level Kullbach–Liebler divergence as implemented in the RUST package38. Peaks in skewness at the fragment ends likely result from library preparation artefacts while the peak at the central A-site (+17 nt from the footprint 5′ end) probably results from biological differences in codon decoding speed38. b, Probabilities for occurrence of nucleotides around the 5′ or 3′ ends of TCP-seq reads (footprint 5′ and 3′ ends) generated using Weblogo 3 (ref. 44), c, d, A depletion of footprint 3′ ends at the uridyl nucleotides. c, Depletion of footprint 3′ ends over U of the AUG triplets is characteristic of all AUG triplets within ORFs, and not only for start AUG codons (visible for example, in Extended Data Fig. 6a, c). Internal AUG triplets were selected from 50 nt downstream of start codons up to ORF ends, and footprint 3′ end densities aligned by these codons. d, Same as c, but footprint 3′ ends were aligned around the first nucleotides of out-of-frame stop codons located within ORF regions.
Extended Data Figure 5 Assorted analyses of TCP-seq data for the SYO1 yeast strain with mild deficiency in cytoplasmic LSU levels11.
a, b, As in Fig. 2a, b. c, d, Top panels as in Extended Data Fig. 3a, bottom panels as in Fig. 4a, c. e, As in Extended Data Fig. 3d, but for the SYO1 cells. Note that horizontal axes in c, d are aligned by the position of the first nucleotide in the P-site of initiating (left panels) or terminating (right panels) to allow for consistency in footprint 5′ and 3′ end extensions across translation complexes. For terminating complexes, this corresponds to the first nucleotide of the last sense codon (not the stop codon). Error regions in c–e indicate the 95th percentiles of 1,000 iterations of gene-wise bootstrapping (see Methods).
Extended Data Figure 6 Snapshots of SSU and ribosome footprint distribution reveal translation initiation by a cap-severed, SSU-led scanning mechanism and staged ribosome recycling after termination, visualized as footprint length versus footprint 3′ end positions aggregated across all mRNAs.
a, b, As in Fig. 2a, b. c, d, As in Extended Data Fig. 5a, b, but for footprint 3′ ends. e, Heatmap of footprint 3′ end positions relative to start codons, aggregated amongst mRNAs having an annotated 5′ UTR of at least 100 nt in length45. The extended scanning SSU are arrested when the leading edge (footprint 3′ end) overruns the start codon by a set margin, irrespective of the trailing edge (footprint 5′ end) position. This is evidenced by the scarcity of longer footprints with 3′ end > +30 nt position and 5′ end < −13 nt position (window 2; encompassing 175 footprints) relative to that expected from an equivalently bounded region (footprint 3′ end > −30 nt and footprint 5′ end < −73 nt) that is fully within the 5′ UTR (window 1; encompassing 1,552 footprints). Isoline iso-3′ and iso-5′ margins forming window 1 and 2 represent positions of the footprint 3′ end and footprint 5′ end isolines of SSU located upstream of (window 1) or on (window 2) the start codons. f, Schematic of plausible topologies of scanning SSU complexes as they approach and align over the start codons, to assist in interpreting data in e.
Extended Data Figure 7 TCP-seq data for individual mRNAs from wild-type and SYO1 cells demonstrating uninterrupted 5′ UTR scanning and high frequency of initiation.
a, Translation initiation on the 5′UTR of ACT1 mRNA coding for actin. Individual SSU footprints observed on the ACT1 mRNA of wild-type (upper panel) and SYO1 (middle panel) cells. Footprint 5′ ends of each footprint are shown in black. Annotated mRNA 5′ end45 and ORF start are indicated by blue and red arrows above the panels and lines of the same colour; potential start codons and stop codons along the 5′ UTR are indicated below in each of three reading frames (lower panel), colour-coded by bars (with the legend on the right). Initiation codons located in perfect nucleotide context (for yeast) are shown as bars of double width. b, Same as a, but for a collection of additional mRNAs. See also the web interface (http://dx.doi.org/10.6084/m9.figshare.3206725) for browsing and analysing the TCP-seq data for wild-type and SYO1 yeast strains, available at (http://bioapps.erc.monash.edu/TCP/).
Extended Data Figure 8 TCP-seq data for individual mRNAs from wild-type and SYO1-TAP cells demonstrating SSU pile-ups within 5′ UTRs and decreased frequency of initiation.
Designations and abbreviations as in Extended Data Fig. 7. a, Translation initiation on the 5′ UTR of GCN4 mRNA coding for general control nonderepressible bZIP transcriptional activator of amino acid biosynthetic genes. b, Same as a, but for a collection of other mRNAs. See also the web interface for browsing and analysing the TCP-seq data for wild-type and SYO1 yeast strains (http://bioapps.erc.monash.edu/TCP/).
Extended Data Figure 9 Efficiency of scanning and start codon selection across yeast mRNAs.
a, Dependency of the ratio of 5′ UTR to start codon-associated SSU footprints on 5′ UTR length. b, Dependency of the relative SSU count at start codons on the length of the 5′ UTR. c, Dependency of 5′ UTR SSU footprint counts (vertical axis) on the abundance and 5′ UTR length of mRNA (horizontal axis; as determined by 5′ UTR RNA fragment counts in the total input fraction) and translation initiation rates (colour; as determined by count of start-codon occupying SSU footprints). Data shown in a–c is for the wild-type yeast strain. d, e, Cross-correlation of the three major start codon footprint types by 3′ end position (refer to Fig. 4a, b) observed across mRNAs of the wild-type (WT) (d) and SYO1-TAP (SYO1) (e) strains. f, Correlation between wild-type and SYO1 yeast cells of the gene-wise footprint count of each of the three major start codon footprint types. Pearson’s correlation coefficients are indicated (r). All counts were incremented by 0.5 to allow logarithmic transformation.
Extended Data Figure 10 Summary of TCP-seq findings and their interpretation in the context of existing mechanistic models and information on ribosome structure.
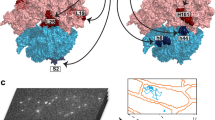
a, Schematic depicting the proposed progression of translation initiation, including cap-severed, SSU-led scanning, and structural rearrangements at start codons. Post-termination events featuring lingering of SSU at stop codons are also shown. b, View of the translation cycle with emphasis on the nexus between TCP-seq footprint types and recent structural information on the SSU in complex with translation factors. The schematic models represent structures obtained for in vitro assembled complexes that are taken to represent particular sub-steps of the translation cycle. Note that in vivo translation intermediates might not completely match the composition and/or structural configuration of in vitro assembled complexes. Schematic models representing the anticipated arrangement of scanning and start codon pausing SSU14,19,20,21,22, start codon recognition14,19,20,21,22, pre-subunit joining23,24 and post-termination recycling25 SSU complexes are shown. eIF1 (blue), eIF1A (red), eIF2 (dark green), initiator tRNAiMet (light green), eIF3 (purple), eIF4A/B/G/H and ABCE1 (amber), eIF5B (yellow), and elongator tRNAeAa (light green).
Supplementary information
Supplementary Table 1
Summary of sequencing read counts for each of the RNA fragment libraries, mapped step-wise to different RNA biotypes (rRNA, other non-coding RNAs, mRNAs). The libraries included RNA fragments from either ribosomal small subunit (SSU), full ribosome (RS) and unseparated total input (TI) fractions of the wild-type or SYO1-TAP yeast strains. (XLSX 14 kb)
Supplementary Table 2
Ranking of Saccharomyces cerevisiae mRNAs by the degree of disrupted ribosomal scanning. To derive a measure for scanning disruption, for each mRNA a ratio was calculated of SSU peak coverage in the 5'UTR over SSU peak coverage in the start codon region. (XLSX 366 kb)
Supplementary Table 3
List of sequences of DNA probes used to deplete libraries for cDNA molecules containing ribosomal RNA (rRNA) sequences by the probe-directed degradation method. (XLSX 13 kb)
Supplementary Discussion
The introduction provides an in-depth and referenced account of eukaryotic translation initiation focusing on the major questions addressed in this work. Results and discussion further integrate particulars of the TCP-Seq method, and the findings obtained with it, into the current knowledge of eukaryotic protein synthesis. (PDF 436 kb)
Rights and permissions
About this article
Cite this article
Archer, S., Shirokikh, N., Beilharz, T. et al. Dynamics of ribosome scanning and recycling revealed by translation complex profiling. Nature 535, 570–574 (2016). https://doi.org/10.1038/nature18647
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18647
This article is cited by
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
The molecular basis of translation initiation and its regulation in eukaryotes
Nature Reviews Molecular Cell Biology (2024)
-
Streamlined and sensitive mono- and di-ribosome profiling in yeast and human cells
Nature Methods (2023)
-
Principles, challenges, and advances in ribosome profiling: from bulk to low-input and single-cell analysis
Advanced Biotechnology (2023)
-
Distance-dependent inhibition of translation initiation by downstream out-of-frame AUGs is consistent with a Brownian ratchet process of ribosome scanning
Genome Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.