Abstract
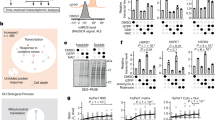
The mitochondrial matrix is unique in that it must integrate the folding and assembly of proteins derived from the nuclear and mitochondrial genomes. In Caenorhabditis elegans, the mitochondrial unfolded protein response (UPRmt) senses matrix protein misfolding and induces a program of nuclear gene expression, including mitochondrial chaperonins, to promote mitochondrial proteostasis1,2,3. While misfolded mitochondrial-matrix-localized ornithine transcarbamylase induces chaperonin expression4,5,6, our understanding of mammalian UPRmt is rudimentary7, reflecting a lack of acute triggers for UPRmt activation. This limitation has prevented analysis of the cellular responses to matrix protein misfolding and the effects of UPRmt on mitochondrial translation to control protein folding loads. Here we combine pharmacological inhibitors of matrix-localized HSP90/TRAP1 (ref. 8) or LON protease9, which promote chaperonin expression, with global transcriptional and proteomic analysis to reveal an extensive and acute response of human cells to UPRmt. This response encompasses widespread induction of nuclear genes, including matrix-localized proteins involved in folding, pre-RNA processing and translation. Functional studies revealed rapid but reversible translation inhibition in mitochondria occurring concurrently with defects in pre-RNA processing caused by transcriptional repression and LON-dependent turnover of the mitochondrial pre-RNA processing nuclease MRPP3 (ref. 10). This study reveals that acute mitochondrial protein folding stress activates both increased chaperone availability within the matrix and reduced matrix-localized protein synthesis through translational inhibition, and provides a framework for further dissection of mammalian UPRmt.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Haynes, C. M., Petrova, K., Benedetti, C., Yang, Y. & Ron, D. ClpP mediates activation of a mitochondrial unfolded protein response in C. elegans. Dev. Cell 13, 467–480 (2007)
Haynes, C. M., Yang, Y., Blais, S. P., Neubert, T. A. & Ron, D. The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans. Mol. Cell 37, 529–540 (2010)
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012)
Zhao, Q. et al. A mitochondrial specific stress response in mammalian cells. EMBO J. 21, 4411–4419 (2002)
Horibe, T. & Hoogenraad, N. J. The chop gene contains an element for the positive regulation of the mitochondrial unfolded protein response. PLoS ONE 2, e835 (2007)
Aldridge, J. E., Horibe, T. & Hoogenraad, N. J. Discovery of genes activated by the mitochondrial unfolded protein response (mtUPR) and cognate promoter elements. PLoS ONE 2, e874 (2007)
Haynes, C. M., Fiorese, C. J. & Lin, Y.-F. Evaluating and responding to mitochondrial dysfunction: the mitochondrial unfolded-protein response and beyond. Trends Cell Biol. 23, 311–318 (2013)
Kang, B. H. et al. Combinatorial drug design targeting multiple cancer signaling networks controlled by mitochondrial Hsp90. J. Clin. Invest. 119, 454–464 (2009)
Bernstein, S. H. et al. The mitochondrial ATP-dependent Lon protease: a novel target in lymphoma death mediated by the synthetic triterpenoid CDDO and its derivatives. Blood 119, 3321–3329 (2012)
Holzmann, J. et al. RNase P without RNA: identification and functional reconstitution of the human mitochondrial tRNA processing enzyme. Cell 135, 462–474 (2008)
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011)
Kim, Y. E., Hipp, M. S., Bracher, A., Hayer-Hartl, M. & Hartl, F. U. Molecular chaperone functions in protein folding and proteostasis. Annu. Rev. Biochem. 82, 323–355 (2013)
Lamech, L. T. & Haynes, C. M. The unpredictability of prolonged activation of stress response pathways. J. Cell Biol. 209, 781–787 (2015)
Siegelin, M. D. et al. Exploiting the mitochondrial unfolded protein response for cancer therapy in mice and human cells. J. Clin. Invest. 121, 1349–1360 (2011)
Guyton, K. Z., Xu, Q. & Holbrook, N. J. Induction of the mammalian stress response gene GADD153 by oxidative stress: role of AP-1 element. Biochem. J. 314, 547–554 (1996)
Zajac, M., Gomez, G., Benitez, J. & Martínez-Delgado, B. Molecular signature of response and potential pathways related to resistance to the HSP90 inhibitor, 17AAG, in breast cancer. BMC Med. Genomics 3, 44 (2010)
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014)
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011)
Lopez Sanchez, M. I. G. et al. RNA processing in human mitochondria. Cell Cycle 10, 2904–2916 (2011)
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011)
Reinhard, L., Sridhara, S. & Hällberg, B. M. Structure of the nuclease subunit of human mitochondrial RNase P. Nucleic Acids Res. 43, 5664–5672 (2015)
Jiang, H. Y. et al. Activating transcription factor 3 is integral to the eukaryotic initiation factor 2 kinase stress response. Mol. Cell. Biol. 24, 1365–1377 (2004)
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE 5, e13984 (2010)
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011)
Bozidis, P., Williamson, C. D. & Colberg-Poley, A. M. in Current Protocols in Cell Biology (eds Bonifacino, J. S. et al.) Ch. 3, Unit 3.27 (Wiley, 2007)
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010)
Pagliarini, D. J. et al. A mitochondrial protein compendium elucidates complex I disease biology. Cell 134, 112–123 (2008)
McKenzie, M., Lazarou, M., Thorburn, D. R. & Ryan, M. T. Mitochondrial respiratory chain supercomplexes are destabilized in Barth syndrome patients. J. Mol. Biol. 361, 462–469 (2006)
Acknowledgements
We thank D. C. Altieri for GTPP, J. Paulo and S. P. Gygi for assistance with mass spectrometry, and L. Pontano and S. H. Sui for assistance with RNA-seq. This work was supported by National Institutes of Health grant R37NS083524 and Biogen, Inc. (J.W.H.), and an EMBO Fellowship (C.M.).
Author information
Authors and Affiliations
Contributions
C.M. and J.W.H. were responsible for project conception, data analysis and manuscript preparation. All experiments were performed by C.M.
Corresponding author
Ethics declarations
Competing interests
J.W.H. is a consultant for Biogen Inc. J.W.H.'s interests were reviewed and are managed by Harvard Medical School in accordance with their conflict of interest policies.
Additional information
Reviewer Information Nature thanks G. Shadel and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Mitochondrial HSP90 inhibition induces UPRmt.
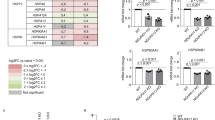
a, Quantitative PCR monitoring chaperonin (HSPD1 and HSPE1) mRNA levels upon treatment of cells with GTPP. Shown are means of levels relative to untreated ± s.d. (n = 3 biological replicates). b, Measurement of cell viability upon GTPP treatment with CCK8. Shown are means of levels relative to untreated ± s.d. (n = 5 biological replicates). c, Measurement of mitochondrial membrane potential upon GTPP or CCCP (mitochondrial membrane potential uncoupler) treatment, measured with JC-1 and analysed on a BD FACSCalibur. Shown are means of levels relative to untreated ± s.d. (n = 3 biological replicates). d, Measurement of cellular ATP levels upon GTPP or antimycin A (electron transport chain inhibitor) treatment. Shown are means of levels relative to untreated ± s.d. (n = 4 biological replicates) and two-tailed P values ***P ≤ 0.001; NS, not significant. e, Blue native gel analysis of mitochondrial respiratory chain complexes upon 6 h treatments of DMSO or GTPP. f, Changes in chaperonin and TRAP1 mRNA levels upon knockdown with shRNA targeting GFP or TRAP1 mRNA. Shown are means of log2 fold changes relative to control and s.d. (n = 3 biological replicates).
Extended Data Figure 2 UPRmt signals distinctly from the integrated stress response.
a, Table with summarized results of data shown in Fig. 1a–c. Induced genes are labelled green and compounds are clustered into their molecular function. GTPP induces UPRmt and tunicamycin ERUPR. All other compounds affect mitochondrial respiration and/or the mitochondrial membrane potential. b, Schematic showing how different stresses signal through the integrated stress response pathway based on the results shown in b and Fig. 1a–c. c, Quantitative PCR to assess the mRNA knockdown of the four EIF2A kinases by siRNA smart pools in biological duplicate; repl., replicate d, Quantitative PCR monitoring CHOP mRNA levels in untreated or GTPP-treated cells with or without knockdown of the EIF2A kinases as in c. e, Quantitative PCR to monitor PERK mRNA levels upon PERK knockdown with individual siRNAs in biological duplicate. f, Quantitative PCR monitoring CHOP mRNA levels in GTPP-treated cells with or without knockdown of PERK by individual siRNAs in biological duplicate.
Extended Data Figure 3 Global analysis of transcriptional responses to UPRmt induction.
a, Heatmap of measured transcript abundances of cells treated with DMSO, 10 μM GTPP or 2.5 μM CDDO for 6 h (n = 3 biological replicates). Values not passing the cuffdiff threshold of FPKM abundance and read number were excluded (white). b, Correlation of replicates for DMSO-, GTPP- and CDDO-treated samples with R values depicting correlation value; log10-transformed FPKM values (≥0) are plotted. c, Quantitative PCR monitoring induction of mtUPR by measuring chaperonin mRNA levels upon treatment with DMSO or CDDO. Shown are means of levels relative to DMSO-treated ± s.d. (n = 3 biological replicates). d, Correlation between the abundance of transcripts significantly altered in GTPP- versus CDDO-treated cells (Fig. 1c, combined panel). e, Table representing changed transcripts upon GTPP or CDDO treatment (Fig. 1c) compared with the number of transcripts changed upon 17AAG previously reported16.
Extended Data Figure 4 Promoter analysis of UPRmt-induced transcripts encoding mitochondrial proteins.
Analysis of UPRmt-induced (GTPP and CDDO) transcripts encoding mitochondrial proteins for the occurrence of CHOP, MURE1, MURE2, or ATF4 promoter elements. We used FIMA version 4.11.1 with the consensus sequences shown. Cells marked in green represent the presence of the consensus sequence in the gene shown.
Extended Data Figure 5 Changes in the mitochondrial proteome upon UPRmt induction.
a, Assay design. b, Summary of proteomic data. c, Analysis of changes in the average abundance of mitochondrial ribosome (left) or for individual ribosomal subunits (right). Values are mean values ± s.d. of scaled signal to noise values (that is, relative abundance) derived from the quantitative proteomics (Fig. 2) for identified mitochondrial ribosomal subunits (right, n = 2 biological replicates) and the average of all these values ± s.d. (left); repl., replicate. d, Analysis of the abundance of the different mitochondrial electron transport chain complexes and ATP synthase. Values are derived from quantitative proteomics (Fig. 2) and shown as mean values ± s.d. across all quantified subunits (top left) or separately per subunit for the different complexes (n = 2 biological replicates). All data depict scaled signal to noise values (that is, relative abundance).
Extended Data Figure 6 Mitochondrial pre-RNA processing defects upon UPRmt.
a, Primer design for monitoring pre-RNA processing. Primer pairs 1 and 3, and 2 and 4, will only produce PCR products for uncleaved mitochondrial pre-RNAs and allow quantitation of non-processed pre-RNAs. Primer pair 2 and 3 will monitor total levels for normalization. b, Quantitative PCR of MRPP3 mRNA levels upon knockdown with siRNA targeting a scrambled sequence or MRPP3. Shown are averages ± s.d. (n = 3 biological replicates). c, qPCR of mitochondrial pre-RNA at tRNAMet and tRNALys RNaseP processing sites upon depletion of MRPP3 by RNAi. Error bars, ±s.d. (n = 3 biological replicates). d, Quantitative PCR monitoring levels of non-processed pre-RNA upon treatment of cells with GTPP or the uncoupler CCCP in biological duplicate; repl., replicate e, LON protein levels as determined by quantitative proteomics (Fig. 2) in biological duplicate. Shown are scaled signal to noise values observed (that is, relative abundance). f. Western blot analysis of LON levels upon control or 10 μM GTPP treatment (6 h).
Extended Data Figure 7 Rescue of UPRmt-induced mitochondrial pre-RNA processing by MRPP3 overexpression.
a, Western Blot analysis of MRPP3 levels upon DMSO or GTPP treatment in the context of wild-type or MRPP3-overexpressing (o/e) cells. b, Quantitative PCR analysis of non-processed mitochondrial pre-RNA levels at the tRNAMet and tRNALys cut sites in wild-type cells or cells overexpressing MRPP3. Shown are mean values ± s.d. (n = 3 biological replicates). c, Quantitative PCR analysis of non-processed mitochondrial pre-RNA levels at the tRNAMet and tRNALys cut sites in wild-type cells or cells overexpressing MRPP3 upon GTPP treatment. Shown are mean values ± s.d. (n = 3 biological replicates).
Extended Data Figure 8 Mitochondrial translation defects upon UPRmt.
a, Coomassie gel staining as a loading control of the same experiment as in Fig. 4b. b, Analysis of cytosolic translation upon treatment with DMSO or GTPP with the same experimental procedure as in Fig. 4a without the addition of emetine. Newly synthesized proteins were monitored by phospho-imager (left) with Coomassie staining of the same gel as loading control (right). c, Table of experiment number, mass spectrometer used, analysis method, peptides sequence, protein encoded and heavy-to-light ratios (H/L) used to determine protein synthesis rates in Fig. 4d. Fusion and QE are Orbitrap Fusion or Q Exactive (Thermo Scientific), respectively; Core depicts in-house mass spectrometry analysis pipeline; #, oxidative modification on methionine; *, could not be determined by Core/Maxquant.
Extended Data Figure 9 Proteomic determination of mitochondrial translation upon UPRmt.
SILAC spectra for the data shown in Fig. 4d and Extended Data Fig. 9c for experiment 1.
Extended Data Figure 10 Reversibility of UPRmt-induced mitochondrial pre-RNA processing and translation defects.
a, Coomassie gel staining as a loading control of the same experiment as in Fig. 4e. b, Mitochondrial pre-RNA processing was measured by qPCR in cells subjected to GTTP pulse-chase wash-out for 1–4 h. Data are averages of fold changes versus untreated ± s.d., two-tailed P values *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, n = 3 biological replicates. c, Analysis of mitochondrial translation in wild-type or MRPP3 overexpressing cells with or without GTPP treatment. Newly synthesized proteins were labelled with 35S and analysed by phospho-imager. d, Immunoblot of TFB1M expression with or without 6 h GTPP treatment (left). Quantification of control normalized TFB1M levels from immunoblots of two independent experiments.
Supplementary information
Supplementary Figure
This file contains the uncropped scans with size marker indications. (PDF 1155 kb)
Supplementary Data
This file contains Supplementary Tables 1-4. (XLSX 4614 kb)
Rights and permissions
About this article
Cite this article
Münch, C., Harper, J. Mitochondrial unfolded protein response controls matrix pre-RNA processing and translation. Nature 534, 710–713 (2016). https://doi.org/10.1038/nature18302
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18302
This article is cited by
-
Mammalian integrated stress responses in stressed organelles and their functions
Acta Pharmacologica Sinica (2024)
-
Emerging roles of mitochondrial functions and epigenetic changes in the modulation of stem cell fate
Cellular and Molecular Life Sciences (2024)
-
Celastrol attenuates hepatitis C virus translation and inflammatory response in mice by suppressing heat shock protein 90β
Acta Pharmacologica Sinica (2023)
-
A cytosolic surveillance mechanism activates the mitochondrial UPR
Nature (2023)
-
Proteolytic rewiring of mitochondria by LONP1 directs cell identity switching of adipocytes
Nature Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.