Abstract
The population history of Aboriginal Australians remains largely uncharacterized. Here we generate high-coverage genomes for 83 Aboriginal Australians (speakers of Pama–Nyungan languages) and 25 Papuans from the New Guinea Highlands. We find that Papuan and Aboriginal Australian ancestors diversified 25–40 thousand years ago (kya), suggesting pre-Holocene population structure in the ancient continent of Sahul (Australia, New Guinea and Tasmania). However, all of the studied Aboriginal Australians descend from a single founding population that differentiated ~10–32 kya. We infer a population expansion in northeast Australia during the Holocene epoch (past 10,000 years) associated with limited gene flow from this region to the rest of Australia, consistent with the spread of the Pama–Nyungan languages. We estimate that Aboriginal Australians and Papuans diverged from Eurasians 51–72 kya, following a single out-of-Africa dispersal, and subsequently admixed with archaic populations. Finally, we report evidence of selection in Aboriginal Australians potentially associated with living in the desert.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
The Aboriginal Australian and Papuan whole genome sequence data generated in this study have been deposited at the European Genome-phenome Archive (EGA, http://www.ebi.ac.uk/ega/), which is hosted by the EBI, under the accession numbers EGAS00001001766 and EGAS00001001247, respectively. The Papuan SNP array data generated in this study can be found under http://geogenetics.ku.dk/latest-news/alle_nyheder/2016/data.
References
Davidson, I. The colonization of Australia and its adjacent islands and the evolution of modern cognition. Curr. Anthropol. 51, S177–S189 (2010)
Clarkson, C. et al. The archaeology, chronology and stratigraphy of Madjedbebe (Malakunanja II): A site in northern Australia with early occupation. J. Hum. Evol. 83, 46–64 (2015)
O’Connell, J. F. & Allen, J. The process, biotic impact, and global implications of the human colonization of Sahul about 47,000 years ago. J. Archaeol. Sci. 56, 73–84 (2015)
Barker, G. et al. The ‘human revolution’ in lowland tropical Southeast Asia: the antiquity and behaviour of anatomically modern humans at Niah Cave (Sarawak, Borneo). J. Hum. Evol. 52, 243–261 (2007)
Lahr, M. M. & Foley, R. Multiple dispersals and modern human origins. Evol. Anthropol. Issues News Rev . 3, 48–60 (1994)
Reyes-Centeno, H. et al. Genomic and cranial phenotype data support multiple modern human dispersals from Africa and a southern route into Asia. Proc. Natl Acad. Sci. USA 111, 7248–7253 (2014)
Wollstein, A. et al. Demographic history of Oceania inferred from genome-wide data. Curr. Biol. 20, 1983–1992 (2010)
Rasmussen, M. et al. An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334, 94–98 (2011)
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010)
Reeves, J. M. et al. Climate variability over the last 35,000 years recorded in marine and terrestrial archives in the Australian region: an OZ-INTIMATE compilation. Quat. Sci. Rev. 74, 21–34 (2013)
Hiscock, P. & Wallis, L. A. in Desert Peoples (eds Veth, P., Smith, M. & Hiscock, P. ) 34–57 (Blackwell Publishing Ltd, 2005)
Birdsell, J. B. Microevolutionary Patterns in Aboriginal Australia: A Gradient Analysis of Clines . (Oxford University Press, 1993)
Bowern, C. & Atkinson, Q. Computational phylogenetics and the internal structure of Pama–Nyungan. Language 88, 817–845 (2012)
Dixon, R. M. W. Australian Languages: Their Nature and Development . (Cambridge University Press, 2002)
Evans, N. & McConvell, P. in Archaeology and Language II: Archaeological Data and Linguistic Hypotheses (eds Blench, R. & Spriggs, M. ) Ch. 7 (Routledge, 1999)
Hiscock, P. Archaeology of ancient Australia . (Routledge, 2008)
Bellwood, P. First Migrants: Ancient Migration in Global Perspective . (Wiley-Blackwell, 2013)
Pugach, I., Delfin, F., Gunnarsdóttir, E., Kayser, M. & Stoneking, M. Genome-wide data substantiate Holocene gene flow from India to Australia. Proc. Natl Acad. Sci. USA 110, 1803–1808 (2013)
Ellinghaus, K. Absorbing the ‘Aboriginal problem’: controlling interracial marriage in Australia in the late 19th and early 20th centuries. Aborig. Hist . 27, 183–207 (2003)
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014)
Frichot, E., Mathieu, F., Trouillon, T., Bouchard, G. & François, O. Fast and efficient estimation of individual ancestry coefficients. Genetics 196, 973–983 (2014)
Patterson, N. J. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012)
Excoffier, L., Dupanloup, I., Huerta-Sánchez, E., Sousa, V. C. & Foll, M. Robust demographic inference from genomic and SNP data. PLoS Genet . 9, e1003905 (2013)
Thorne, A. G. in The Origin of the Australians (eds Kirk, R. L. & Thorne, A. G. ) 95–112 (Canberra: Australian Institute of Aboriginal Studies, 1976)
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nat. Genet. 46, 919–925 (2014)
Qin, P. & Stoneking, M. Denisovan Ancestry in East Eurasian and Native American Populations. Mol. Biol. Evol. 32, 2665–2674 (2015)
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014)
Gutenkunst, R. N., Hernandez, R. D., Williamson, S. H. & Bustamante, C. D. Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genet . 5, e1000695 (2009)
Bergström, A. et al. Deep roots for Aboriginal Australian Y chromosomes. Curr. Biol. 26, 809–813 (2016)
Hudjashov, G. et al. Revealing the prehistoric settlement of Australia by Y chromosome and mtDNA analysis. Proc. Natl Acad. Sci. USA 104, 8726–8730 (2007)
Lippold, S. et al. Human paternal and maternal demographic histories: insights from high-resolution Y chromosome and mtDNA sequences. Investig. Genet . 5, 13 (2014)
Radcliffe-Brown, A. R. The social organization of Australian tribes. Oceania 1, 34–63 (1930)
Veth, P. Islands in the interior: a model for the colonization of Australia’s arid zone. Archaeol. Ocean. 24, 81–92 (1989)
Drummond, A. J., Rambaut, A., Shapiro, B. & Pybus, O. G. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol. Biol. Evol. 22, 1185–1192 (2005)
Lourandos, H. & David, B. in Bridging Wallace’s Line: the Environmental and Cultural History and Dynamics of the SE Asian-Australasian Region (eds Kershaw, A. P., David, B., Tapper, N., Penny, D. & Brown, J. ) Advances in GeoEcology 34, 97–118 (2002)
Cavalli-Sforza, L. L. Genes, peoples and languages. Proc. Natl Acad. Sci. USA 91, 7719–7724 (1997)
Evans, N. & Jones, R. in Archaeology and linguistics: Aboriginal Australia in global perspective (Oxford University Press Australia, 1997)
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010)
Qi, X., Chan, W. L., Read, R. J., Zhou, A. & Carrell, R. W. Temperature-responsive release of thyroxine and its environmental adaptation in Australians. Proc. Biol. Sci. 281, 20132747 (2014)
Tin, A. et al. Genome-wide association study for serum urate concentrations and gout among African Americans identifies genomic risk loci and a novel URAT1 loss-of-function allele. Hum. Mol. Genet. 20, 4056–4068 (2011)
Scally, A. & Durbin, R. Revising the human mutation rate: implications for understanding human evolution. Nat. Rev. Genet. 13, 745–753 (2012)
Fenner, J. N. Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. Am. J. Phys. Anthropol. 128, 415–423 (2005)
Holt, S. Palaeoenvironments of the Gulf of Carpentaria from the Last Glacial Maximum to the Present, as Determined by Foraminiferal Assemblages. PhD thesis, Univ. Wollongong (2005)
Heupink, T. H. et al. Ancient mtDNA sequences from the First Australians revisited. Proc. Natl Acad. Sci. USA 113, 6892–6897 (2016)
Horton, D. (ed) The Encyclopaedia of Aboriginal Australia . (Aboriginal Studies Press, 1994)
Migliano, A. B. et al. Evolution of the pygmy phenotype: evidence of positive selection from genome-wide scans in African, Asian, and Melanesian pygmies. Hum. Biol. 85, 251–284 (2013)
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014)
Wall, J. D. et al. Higher levels of Neanderthal ancestry in East Asians than in Europeans. Genetics 194, 199–209 (2013)
Vernot, B. & Akey, J. M. Resurrecting surviving Neandertal lineages from modern human genomes. Science 343, 1017–1021 (2014)
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015)
Wang, C. et al. Comparing spatial maps of human population-genetic variation using Procrustes analysis. Stat. Appl. Genet. Mol. Biol. 9, 13 (2010)
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007)
Skoglund, P. & Jakobsson, M. Archaic human ancestry in East Asia. Proc. Natl. Acad. Sci. USA 108, 18301–18306 (2011)
Sankararaman, S., Mallick, S., Patterson, N. & Reich, D. The combined landscape of Denisovan and Neanderthal ancestry in present-day humans. Curr Biol. 26, 1241–1247 (2016)
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature http://dx.doi.org/10.1038/nature18964 (this issue)
Pagani, L. et al. Genomic analyses inform on migration events during the peopling of Eurasia. Nature http://dx.doi.org/10.1038/nature19792 (this issue)
Reich, D., et al. Denisova admixture and the first modern human dispersals into Southeast Asia and Oceania. Am. J. Hum. Genet. 89, 516–528 (2011)
Cheng, J. Y., Mailund, T., & Nielsen, R. Ohana, a tool set for population genetic analyses of admixture components. bioRxiv doi:10.1101/071233 (2016)
Acknowledgements
We thank all sample donors for contributing to this study. We thank Macrogen (http://www.macrogen.com/) for sequencing of the Aboriginal Australian genomes, M. Rasmussen, C. Der Sarkissian, M. Allentoft, D. Cooper, R. Gray, S. Greenhill, A. Seguin-Orlando, T. Carstensen, M. Przeworski, J. D. Jensen and L. Orlando for helpful discussions. We thank E. Thorsby for sample collection and contributing the DNA extract for the P2077 genome, I. Lissimore for support with data storage and distribution. We thank T. Parks, K. Auckland, K. Robson, A. V. Hill, J. B. Clegg, D. Higgs, D. J. Weatherall and M. Alpers for assistance in sample collection and discussion. L.E., V.C.S., I.A., I.D. and S.P. are grateful to the High Performance Computation platform of the University of Bern for providing access to the UBELIX cluster. This work was supported by the Danish National Research Foundation, the Lundbeck Foundation, the KU2016 grant and the Australian Research Council. A.-S.M. was supported by an ambizione grant with reference PZ00P3_154717 from the Swiss National Science Foundation (SNSF). M.C.W. was supported by the Australian Research Council (ARC) Discovery grants DP110102635 and DP140101405 and by a Linkage grant LP140100387. V.C.S., I.D. and S.P. were supported by SNSF grants to L.E. with references 31003A-143393 and CRSII3_141940. O.L. was supported by a Ramón y Cajal grant from the Spanish Ministerio de Economia y Competitividad (MINECO) with reference RYC-2013-14797 and by a BFU2015-68759-P (MINECO/FEDER) grant. I.A. was supported by a grant with reference SFRH/BD/73150/2010 from the Portuguese Foundation for Science and Technology (FCT). A.B., S.Sc., Y.X., C.T.-S. and R.D. were supported by a Wellcome Trust grant with reference WT098051. E.M., C.Ba., I.P., S.N. and M.St. acknowledge the Max Planck Society. S.Su. was supported by an ARC Discovery grant with reference DP140101405. J.L.W. was supported by a PhD scholarship from Griffith University. A.A. acknowledges the Villum foundation. I.M. was supported by a grant from the Danish Council for Independent Research with reference DFF–4090-00244. J.V.M.-M. acknowledges the Consejo Nacional de Ciencia y Tecnología (Mexico) for funding. N.B. and F.-X.R. were supported by the French Ministry of Foreign and European Affairs and French ANR with the grant ANR14-CE31-0013-01. S.B. was supported by a Novo Nordisk Foundation grant with reference NNF14CC0001. P.G. and A.B.M. were supported by a Leverhulme Programme grant number RP2011-R-045 to A.B.M. at UCL Department of Anthropology and M.G.T. at UCL Department of Genetics, Evolution and Environment. A.J.M. was supported by a Wellcome Trust grant with reference 106289/Z/14/Z. M.M. acknowledges the EU European Regional Development Fund through the Centre of Excellence in Genomics to Estonian Biocentre; Estonian Institutional Research grant IUT24-1. M.G.T. was supported by a Wellcome Trust Senior Investigator Award with grant number 100719/Z/12/Z. S.J.O. was supported by a Wellcome Trust Core Award Grant Number 090532/Z/09/Z. A.Man. was supported by an ERC Consolidator Grant 647787 ‘LocalAdaptation’. M.E.P. would like to acknowledge the cardio-metabolic research cluster at Jeffrey Cheah School of Medicine & Health Sciences, Monash University Malaysia and Ministry of Science, Technology & Innovation, Malaysia for research grant 100-RM1/BIOTEK 16/6/2B. M.H.S. was supported by a grant from the Danish Independence Research Council with reference FNU 12-125062. R.A.F. was supported by the Leverhulme Trust. M.M.L. is supported by an ERC Advanced Grant 295907 ‘In-Africa’. C.Bo. was supported by USA National Science Foundation (NSF) grants BCS-0844550 and BCS-1423711, awarded to C.Bo. and Yale University. T.M. was supported by a grant from the Danish Independence Research Council with reference FNU 1323-00749. M.S.S. was supported by a Wellcome Trust grant with reference WT098051. L.E. was supported by Swiss NSF grant number 31003A-143393, D.M.L. was supported by ARC Discovery Grants DP110102635 and DP140101405 and Linkage grants LP140100387, LP120200144 and LP150100583. E.W. is grateful to St John’s College in Cambridge for help and support.
Author information
Authors and Affiliations
Contributions
G.A., J.Y.C., J.E.C., T.H.H., E.M., S.P., S.R., S.Sc., S.Su. and J.L.W. contributed equally and are listed alphabetically in the author list; A.A., C.Ba., I.D., A.E., A.Mar., I.M. and I.P. contributed equally and are listed alphabetically in the author list; T.S.K., I.P.L., J.V.M.-M., S.N., F.R., M.Si. and Y.X. contributed equally and are listed alphabetically in the author list. E.W. and D.M.L. initially conceived and headed the project. L.E. led the genetic load and the SFS-based demographic analyses. M.S.S. headed the research at the Wellcome Trust Sanger Institute. A.-S.M. planned and coordinated the genetic analyses and the sequencing of the Aboriginal Australian genomes. C.M., J.L.W., T.H.H., P.F.C., W.C., G.F., D.I., B.L., A.L., P.J.M., L.M., D.R., T.W., C.W., J.D., M.C.W. and E.W. collaborated with local groups to collect Aboriginal Australian samples. N.B., P.G., G.K., M.L., A.J.M., A.B.M., W.P., F.-X.R., P.S., M.G.T. and S.J.O. collaborated with local groups to collect Papuan samples. S.E. collaborated with local groups to collect the Rapanui sample. A.Mar. extracted DNA for the Aboriginal Australian genomes. M.S.S., A.B. and C.T.-S. coordinated the design and sequencing of the Papuan genomes. O.L., V.C.S., I.A., A.-S.M., A.B., G.A., J.Y.C., J.E.C., T.H.H., E.M., S.P., S.R., S.Sc., S.Su., J.L.W., A.A., C.Ba., I.D., A.E., A.Man., I.M., I.P., T.S.K., I.P.L., J.V.M.-M., S.N., F.R., M.Si., F.A., S.B., L.E., J.D.W. and T.M. analysed genetic data. C.Bo. collected and analysed linguistic data. L.E., E.W., D.M.L., Y.X., M.E.P., C.T.-S., R.D., M.S.S., A.Man., M.H.S., T.M., M.St. and R.N. supervised genetic analyses. M.C.W., C.M., W.C., G.F., D.I., B.L., A.L., P.J.M., L.M., D.R., T.W., C.W., E.A.M.-S., M.M., M.E.P., S.J.O., J.D., A.B.M., R.A.F. and M.M.L. provided archaeological, anthropological and historical context. A.-S.M., V.C.S., O.L., I.A., A.B., M.M.L., R.N., L.E., D.M.L. and E.W. wrote the manuscript with critical input from G.A., T.H.H., E.M., S.Sc., S.Su., J.L.W., C.Ba., A.E., I.P., E.A.M.-S., M.S.S., S.J.O., C.T.-S., R.D., M.G.T., J.D., A.Man., M.H.S., R.A.F., C.Bo., J.D.W., T.M., M.St. and all other coauthors. A.-S.M., V.C.S., O.L., I.A. and A.B. revised and compiled the Supplementary Information.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Per-individual admixture proportions of K = 7 ancestral components including Aboriginal Australians, New Guineans, Europeans, Africans, Melanesians and Polynesians.
The genome of each individual is depicted as a bar and is coloured according to the estimated genome-wide proportions of ancestry components. An unrooted tree showing the relationships between the identified ancestral components is also estimated by our method. Each ancestry has been labelled with the name of the population (see also map) showing the highest fraction of that ancestral component. The cross-validation error is minimized for this value of K for fivefold cross-validation. The rooted tree supports the shared genetic origin of Aboriginal Australians, Papuans and Bougainvilleans. Note that only individuals with more than 50% of Aboriginal Australian ancestry in their genomes (defined in Supplementary Information section S06) were included in the analyses. Refer to ref. 58 and Supplementary Information section S05 for details about the method and the analysis. Map data ©2016 Google, INEGI. Tree constructed with http://jade-cheng.com/trees/.
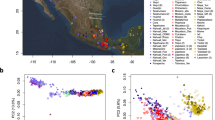
Extended Data Figure 2 Genetic relationships of Aboriginal Australians and Papuans.
a, Genetic affinities between a western central desert (WCD02) genome and Aboriginal Australians and Papuans. Outgroup f3 statistics between WCD02 and all other Aboriginal Australians and Highland Papuan individuals that were whole-genome sequenced for this study, using the genotypes called from the sequencing data. Because the widespread recent admixture in Aboriginal Australians has large confounding effects on the f3 statistics, the values were adjusted using the slope coefficient from a simple linear regression model fitted to the relationship between f3 and the fraction of non-indigenous (that is, neither Aboriginal Australian nor Papuan) ancestry in each individual genome. The adjusted f3 statistics display a genetic gradient that separates western and eastern Aboriginal Australian populations. However, we find no differences between Papuan population samples in their level of Aboriginal Australian affinity (Kruskal–Wallis test, P = 0.083). Horizontal lines correspond to ±1 standard error. b, Genetic affinities between a Papuan highlander genome and Aboriginal Australians and Papuans. The Papuan highlander sample MAR01 from the Marawaka area was arbitrarily chosen as a reference point for this analysis. f3 values were adjusted for recent admixture as in a. All Aboriginal Australian groups display a similar level of Highland Papuan affinity (with the exception of three outlier individuals from the north-eastern WPA and CAI populations: WPA06, WPA05 and CAI10, the latter two of which are known to have at least one parent with origins in Papua New Guinea or the Torres Strait Islands). While some differences between groups are actually statistically significant (Kruskal–Wallis test, P = 0.0002, after removing the three outliers), which could be consistent with, for example, low levels of Papuan gene flow into some Aboriginal Australian groups (see Supplementary Information sections S06 and S07), we caution that some of these differences are probably due to imperfect adjustment for Eurasian admixture (the adjusted f3 is highest in the WCD population, which has the least Eurasian admixture). Horizontal lines correspond to ±1 standard error. c, MSMC analyses. Linear interpolation through the midpoints of the time intervals of the relative cross coalescence rate estimates from MSMC25 using pairs of individuals including one HGDP-Papuan and one other individual as indicated. We used CAI01, PIL06, WCD01, WON03 and an ECCAC sample for this analysis (see Supplementary Information section S08 for details). The MSMC results were scaled using a mutation rate of 1.25×10−8 per generation per site as suggested in ref. 41 and a generation time of 29 years, corresponding to the average hunter–gatherer generation interval for males and females42.
Extended Data Figure 3 Introgressed archaic sites and putative Denisovan and Neanderthal haplotypes.
a, Distribution of number of putative introgressed sites per individual from archaic humans. The number of Neanderthal-specific introgressed sites per individual increases from Europe to Australia, and then decreases in Amerindians, which is consistent with recurrent Neanderthal (or Neanderthal-related archaic) gene flow during the expansion into Eurasia. Our results are thus indicative of several pulses of Neanderthal gene flow into modern humans, as inferred previously48,49,50. We note, however, that the apparent high levels of Neanderthal-specific introgressed sites in Australo-Papuans can be explained by the expected number of misclassified Neanderthal introgressed sites resulting from the shared ancestry with Denisovans (see Supplementary Information section S11 for details). b–e, Putative Denisovan (PDH) and Neanderthal haplotypes (PNH). The putative haplotypes correspond to clusters (four or more SNPs spanning at least 4 kb) of heterozygous or homozygous genotypes in complete linkage disequilibrium (‘diplotypes’) that are potentially the result of Neanderthal or Denisovan admixture. Those diplotypes are homozygous ancestral in 10 Africans, homozygous derived in the Denisovan for the PDH (respectively Neanderthal for the PNH), homozygous ancestral in the Neanderthal for the PDH (respectively Denisovan for the PNH), and with the derived allele segregating in all other contemporary non-African humans (see Supplementary Information section S10 for details). We report the average number of PDHs and PNHs (b), the correlation between the estimated amount of Australo-Papuan ancestry (see Fig. 2a, Extended Data Fig. 1, Supplementary Information section S05) and the number of identified PDHs for each Australian sample (c), the sum of the lengths (d) and the average length (e) of the PDHs and PNHs per individual for worldwide populations included in our reference panel (see Supplementary Information section S04).
Extended Data Figure 4 Out of Africa: admixture graphs based on D-statistics and MSMC analyses.
a, Admixture graphs representing some of the topologies considered for the two waves and one wave Out of Africa models assuming Denisovan admixture. All topologies are identical except for the coloured lineages representing Australo-Papuans (green), Neanderthal (Nea, orange) and Denisovan (Den, blue). The graphs differ in (1) the number of OoA events, and (2) the number of Neanderthal admixture pulses. Png, HGDP-Papuan. b, Sum of squared errors between the observed D-statistics and the expectations for each quartet in the graph involving the chimpanzee as an outgroup for each of the admixture graphs shown in a and the corresponding four without Denisovan admixture. Each point is the result of the optimization procedure with a different starting point. See Supplementary Information section S09 for details. c, Relative cross coalescence rate (CCR) estimates from MSMC25 for pairs of individuals including one African sample (Yoruba, Dinka and San) and one other, as indicated in the legend. d, Simulation study to assess the effect of archaic admixture on the CCR rates. Relative CCR estimated for data simulated under a simple two-population divergence model where one of the populations admixed at different rates with an archaic population. See Supplementary Information section S08 for details.
Extended Data Figure 5 Inferred deleterious mutations.
a, Box plot of the number of derived homozygous sites per individual for worldwide populations that are predicted to be deleterious. Deleteriousness of SNPs was inferred using genomic evolutionary rate profiling (GERP) rejected substitution scores. Derived alleles with a rejected substitution score larger than 2 were considered to be deleterious, see Supplementary Information section S11. b, c, Average rejected substitution score per individual calculated across heterozygous sites (b), and derived homozygous sites (c). Each coloured symbol corresponds to estimates from a single individual. Homozygosity is calculated as the number of derived homozygous sites divided by the number of sites at which an individual carries at least one copy of the derived allele. Solid lines show the linear regression of homozygosity against average rejected substitution score per individual for non-African modern humans. Dashed lines indicate the 95% confidence interval for the linear regression. See Supplementary Information S11 for details.
Extended Data Figure 6 Effective population size changes over time.
a, Population size estimates from MSMC for pairs of individuals from several populations within and outside of Australia. For each run, we used two individuals from each population, that is, four haplotypes in each run. MSMC results were scaled as in Fig. 3. b, Bayesian skyline plots (BSP) calculated from the mtDNA genome sequences, showing the effective population size estimates over time when considering either groups from northeastern Australia (CAI, WPA) or groups from southwestern Australia (ENY, NGA, WCD, WON). Solid lines are the estimates, dashed lines are the corresponding 95% credible intervals (see Supplementary Information section S12).
Extended Data Figure 7 Genetics mirrors geography and languages.
a, b, Procrustes analyses of the first two dimensions of a classical multidimensional scaling (MDS) analysis of the Aboriginal Australian genome sequences (autosomes). We considered two cases: an analysis including all variants (a), or only the variants remaining after genomic regions of putative recent European and East Asian origin are ‘masked’ (b, Supplementary Information section S06). Both MDS plots have been rotated towards the best overlap with geographic sampling locations as defined by Procrustes analysis51. In each plot, the connecting lines indicate the error of the MDS coordinates towards the assigned population-sampling geographic coordinates. We find that the genetic relationships within Australia mirrors geography, with a significant correlation for both cases, that is, rGEN,GEO = 0.59, P < 0.0005 for all variants and even higher, rGEN,GEO = 0.77, P <0.0005, for the masked data. We find using the bearing correlogram approach that the main axis of genetic differentiation in the masked Aboriginal Australian genomes is at an angle of 65° compared to the equator, that is, in the southwest to northeast direction (Supplementary Information section S13). c, d, Correspondence between genetics and linguistics. c, Unrooted neighbour-joining FST-based genetic tree (cladogram). Weir and Cockerham FST distance was computed between the Aboriginal Australian populations after masking the Eurasian tracts. Statistical robustness of each branch was estimated by means of a bootstrap analysis (1,000 replicates, Supplementary Information section S05). d, Bayesian phylogenetic tree for the 28 different Pama–Nyungan languages represented in this sample (from ref. 13, see Supplementary Information section S15). Posterior probabilities are also indicated. Note that one language group can be shared by different Aboriginal Australian groups. The linguistic tree was built with BEAST52. e–g, Gene flow across the continent. e, Mantel non-parametric r (estimating the goodness of fit between genetic differentiation and connectivity) versus ratios of resistance of inland to coastal nodes, showing a peak at 1.7. f, Best fit of pairwise population genetic differentiation, FST (computed between the nine Aboriginal Australian groups after masking Eurasian tracts (Supplementary Information section S06)), versus pairwise connectivity based on the environment (estimated as resistance) when moving inland is 1.7 times harder than moving along coastal nodes. g, Gene flow across the Australian landscape, quantified as the cumulative current for pairwise connections among Aboriginal Australian groups (black circles), with larger current (warmer colours) representing greater gene flow.
Extended Data Figure 8 European, East Asian and Papuan genomic tracts in Aboriginal Australians.
a, Distribution of the tracts assigned to Aboriginal Australian (WCD), Papuan, East Asian or European ancestry for 58 unrelated non-WCD Aboriginal Australian samples. Most of the shorter tracts were of Papuan origin, suggesting that a large fraction of the Papuan gene flow is much older than that from Europe and East Asia, consistent with a Papuan influence spreading slowly from northeastern to southwestern Australia by ancient migration. b, Corresponding scatter plot with fitted line of per-individual variance in Papuan tract length versus geographic distance from WCD, the latter calculated using the great-circle distance formula for pairs of individual GPS coordinates. Papuan tract distribution showed a strong and significant correlation with distance from WCD (r = 0.64; P < 1 × 10−5), with ‘younger tracts’ (that is, with a larger variance) closer to New Guinea and ‘older tracts’ (that is, with a smaller variance) closer to WCD. This is also consistent with continuous Papuan gene flow spreading from the northeast.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Figures, Supplementary Tables and additional references (see Contents for more details). (PDF 18492 kb)
Rights and permissions
About this article
Cite this article
Malaspinas, AS., Westaway, M., Muller, C. et al. A genomic history of Aboriginal Australia. Nature 538, 207–214 (2016). https://doi.org/10.1038/nature18299
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18299
This article is cited by
-
Whole-genome resequencing provides insights into the diversity and adaptation to desert environment in Xinjiang Mongolian cattle
BMC Genomics (2024)
-
Physiography, foraging mobility, and the first peopling of Sahul
Nature Communications (2024)
-
More than a decade of genetic research on the Denisovans
Nature Reviews Genetics (2024)
-
Incorporating uncertainty in Indigenous sea Country monitoring with Bayesian statistics: Towards more informed decision-making
Ambio (2024)
-
North African fox genomes show signatures of repeated introgression and adaptation to life in deserts
Nature Ecology & Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.