Abstract
Many hybrid devices integrate functional molecular or nanoparticle components with microstructures, as exemplified by the nanophotonic devices that couple emitters to optical resonators1 for potential use in single-molecule detection2,3, precision magnetometry4, low threshold lasing5,6 and quantum information processing7,8,9,10,11,12. These systems also illustrate a common difficulty for hybrid devices: although many proof-of-principle devices exist, practical applications face the challenge of how to incorporate large numbers of chemically diverse functional components into microfabricated resonators at precise locations. Here we show that the directed self-assembly13,14 of DNA origami15 onto lithographically patterned binding sites allows reliable and controllable coupling of molecular emitters to photonic crystal cavities (PCCs). The precision of this method is sufficient to enable us to visualize the local density of states within PCCs by simple wide-field microscopy and to resolve the antinodes of the cavity mode at a resolution of about one-tenth of a wavelength. By simply changing the number of binding sites, we program the delivery of up to seven DNA origami onto distinct antinodes within a single cavity and thereby digitally vary the intensity of the cavity emission. To demonstrate the scalability of our technique, we fabricate 65,536 independently programmed PCCs on a single chip. These features, in combination with the widely used modularity of DNA origami16,17,18,19,20, suggest that our method is well suited for the rapid prototyping of a broad array of hybrid nanophotonic devices.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Benson, O. Assembly of hybrid photonic architectures from nanophotonic constituents. Nature 480, 193–199 (2011)
Armani, A. M. et al. Label-free, single-molecule detection with optical microcavities. Science 317, 783–787 (2007)
Yang, D. et al. High sensitivity and high Q-factor nanoslotted parallel quadrabeam photonic crystal cavity for real-time and label-free sensing. Appl. Phys. Lett. 105, 063118 (2014)
Jensen, K. et al. Cavity-enhanced room-temperature magnetometry using absorption by nitrogen-vacancy centers in diamond. Phys. Rev. Lett. 112, 160802 (2014)
Yakunin, S. et al. Low-threshold amplified spontaneous emission and lasing from colloidal nanocrystals of caesium lead halide perovskites. Nat. Commun. 6, 8056 (2015)
Strauf, S. et al. Self-tuned quantum dot gain in photonic crystal lasers. Phys. Rev. Lett. 96, 127404 (2006)
Hennessy, K. et al. Quantum nature of a strongly coupled single quantum dot–cavity system. Nature 445, 896–899 (2007)
Englund, D. et al. Controlling cavity reflectivity with a single quantum dot. Nature 450, 857–861 (2007)
Sapienza, L., Davanço, M., Badolato, A. & Srinivasan, K. Nanoscale optical positioning of single quantum dots for bright and pure single-photon emission. Nat. Commun. 6, 7833 (2015)
Barth, M., Nüsse, N., Löchel, B. & Benson, O. Controlled coupling of a single-diamond nanocrystal to a photonic crystal cavity. Opt. Lett. 34, 1108–1110 (2009)
Englund, D. et al. Deterministic coupling of a single nitrogen vacancy center to a photonic crystal cavity. Nano Lett. 10, 3922–3926 (2010)
Lyasota, A. et al. Integration of multiple site-controlled pyramidal quantum dot systems with photonic-crystal membrane cavities. J. Cryst. Growth 414, 192–195 (2015)
Kershner, R. J. et al. Placement and orientation of individual DNA shapes on lithographically patterned surfaces. Nat. Nanotechnol. 4, 557–561 (2009)
Gopinath, A. & Rothemund, P. W. K. Optimized assembly and covalent coupling of single-molecule DNA origami nanoarrays. ACS Nano 8, 12030–12040 (2014)
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Acuna, G. P. et al. Fluorescence enhancement at docking sites of DNA-directed self-assembled nanoantennas. Science 338, 506–510 (2012)
Schreiber, R. et al. Hierarchical assembly of metal nanoparticles, quantum dots and organic dyes using DNA origami scaffolds. Nat. Nanotechnol. 9, 74–78 (2014)
Bui, H. et al. Programmable periodicity of quantum dot arrays with DNA origami nanotubes. Nano Lett. 10, 3367–3372 (2010)
Ko, S. H., Du, K. & Liddle, J. A. Quantum-dot fluorescence lifetime engineering with DNA origami constructs. Angew. Chem. Int. Ed. 52, 1193–1197 (2013)
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nat. Nanotechnol. 5, 61–66 (2010)
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003)
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009)
Zhang, T. et al. DNA-based self-assembly of fluorescent nanodiamonds. J. Am. Chem. Soc. 137, 9776–9779 (2015)
Barth, M. et al. Modification of visible spontaneous emission with silicon nitride photonic crystal nanocavities. Opt. Express 15, 17231–17240 (2007)
Louvion, N. et al. Local observation and spectroscopy of optical modes in an active photonic-crystal microcavity. Phys. Rev. Lett. 94, 113907 (2005)
Mivelle, M. et al. Light funneling from a photonic crystal laser cavity to a nano-antenna: overcoming the diffraction limit in optical energy transfer down to the nanoscale. Opt. Express 22, 15075–15087 (2014)
Rotenberg, N. & Kuipers, L. Mapping nanoscale light fields. Nat. Photon. 8, 919–926 (2014)
Sapienza, R. et al. Deep-subwavelength imaging of the modal dispersion of light. Nat. Mater. 11, 781–787 (2012)
Knowles, H. S., Kara, D. M. & Atatüre, M. Observing bulk diamond spin coherence in high-purity nanodiamonds. Nat. Mater. 13, 21–25 (2014)
Neu, E. et al. Low-temperature investigations of single silicon vacancy colour centres in diamond. New J. Phys. 15, 043005 (2013)
Stahl, E., Martin, T. G., Praetorius, F. & Dietz, H. Facile and scalable preparation of pure and dense DNA origami solutions. Angew. Chem. Int. Ed. 53, 12735–12740 (2014)
Shaw, A., Benson, E. & Högberg, B. Purification of functionalized DNA origami nanostructures. ACS Nano 9, 4968–4975 (2015)
Klein, M. J. K. Wafer-Scale Fabrication of Thin SiN Membranes and Au Films and Membranes with Arrays of Sub-micron Holes Using Nanosphere Lithography . PhD thesis, École Polytechnique Fédérale de Lausanne (2010)
Noy, A. Handbook of Molecular Force Spectroscopy (Springer, 2007)
Akahane, Y., Asano, T., Song, B.-S. & Noda, S. High-Q photonic nanocavity in a two-dimensional photonic crystal. Nature 425, 944–947 (2003)
Acknowledgements
We acknowledge funding from the Army Research Office (award W911NF-11-1-0117), the Office of Naval Research (award N000141410702), the Air Force Office of Scientific Research (Young Investigator award FA9550-15-1-0252), and the US National Science Foundation (Expeditions in Computing numbers 0832824 and 1317694, Molecular Programming Project; http://molecular-programming.org). We thank J. Fakonas for discussions and B. Fultz for use of his spectrometer. Device fabrication was done at Caltech’s Kavli Nanoscience Institute.
Author information
Authors and Affiliations
Contributions
A.G. and P.W.K.R. conceived the project. A.G. performed origami synthesis, nanofabrication, AFM, SEM and fluorescence microscopy. A.G. and E.M. built the set-up for microphotoluminescence spectroscopy. All authors contributed to data interpretation and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 DNA origami design showing position of Cy5 fluorophores.
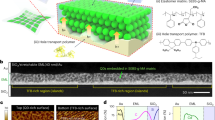
a, We used a triangular DNA origami shape, ~130 nm on each edge, as an adaptor to position fluorophores within a photonic crystal cavity (PCC). The origami is composed of a 7,249-nucleotide (nt) single-stranded scaffold (the commercially available single-stranded genome of the M13mp18 bacteriophage) and approximately 200 commercially synthesized ‘staple strands’ which are typically 32 nucleotides in length. Mixed together and annealed from 90 to 20 °C over the course of about 6 h, the strands self-assemble to form equilateral triangles in high yield. Three trapezoidal domains make up this ‘sameside sharp triangle’ design. Red filled circles indicate 15 positions at which staples were extended on their 5′ end with 18-nt poly-T linkers for the synthesis of origami bearing n = 15 Cy5. Blue rings indicate the subset of positions used for origami bearing n = 3 Cy5. To these linkers, 21-nt poly-A ‘subcomponent’ strands modified with a 5′ Cy5 were hybridized. Because the Cy5 modification is on their 5′ ends, subcomponent strands put Cy5 within a few nucleotides of the origami surface, less than 1 nm from where linkers extend from the origami surface (rather than at the 6-nm-away distal end of the linker/Cy5 strand hybrid). b, Schema showing both components of the fine structure of a DNA origami (gaps between helices, and crossover positions, where helices are tangent) with linker/Cy5-label positions noted as in a. The white outer ring has a 62.4-nm diameter and the white inner ring has a 31.2-nm diameter. As in a, red dots indicate the subset of positions used for origami bearing n = 15 Cy5 and blue rings around red dots indicate positions used for origami bearing n = 3 Cy5. Rainbow colours on the origami trace the path of the scaffold as it progresses through the origami structure, from red to purple. c, Buffer AFM image showing fine structure of origami without linkers/Cy5 strands for comparison with b. d, Buffer AFM image showing origami with n = 3 linkers and Cy5 strands. In this AFM image the Cy5-labelled duplexes appear to fall on the inside of the inner triangular hole, perhaps due to adhesion to the mica substrate used in these high-resolution experiments. High-resolution imaging of origami dried onto SiN is difficult, and so the exact conformation of Cy5-labelled duplexes on resonators was not determined. However, because the carboxylated sites bind DNA strongly, it seems likely that Cy5-labelled duplexes would behave similarly and fall within the 54-nm triangular hole of the origami, between 30 and 60 nm from the centre of the origami.
Extended Data Figure 2 Process flow for fabricating PCCs.
a, Fabrication of isolated PCCs for widely spaced arrays (Fig. 2). b, Fabrication of PCCs in close-packed arrays (Figs 3, 4). Note that back-side alignment markers are omitted from b for clarity. After either fabrication process substrates are incubated in origami solution, rinsed of excess origami, subjected to an ethanol dilution series, and air-dried.
Extended Data Figure 3 SEM imaging of isolated PCCs used for taking fluorescence spectra as a function of x-position.
An 8 × 21 array was used. a, Positions 01 to 21 each indicate a row of eight copies of isolated PCCs having origami positioned at the same x-offset, as exemplified in Extended Data Fig. 4. Red squares indicate broken PCC membranes which were not used. b, Intact membrane. c, Typical broken membrane. d, Zoom-in of PCC. e, Cross-section of PCC membrane. Scale bars: a, 300 μm; b, 2 μm; d, 1 μm; e, 500 nm.
Extended Data Figure 4 AFM of isolated PCCs and control cavity.
a, Schema and representative AFM for each of 21 offsets at which spectra (Fig. 2b) were measured. b, Dry AFM of a control PCC filled with origami (n = 15 Cy5/origami). Scale bar, 250 nm. c, PL reference spectra (red) for Cy5-origami on open SiN, and for Cy5-origami filling a cavity, as in b.
Extended Data Figure 5 Optical set-up for reflectance and fluorescence spectroscopy.
The 650-nm long-pass filter (650 LP) was used only for fluorescence spectroscopy. Details in Methods.
Extended Data Figure 6 Comparison of unaveraged and averaged epifluorescence images of PCC array used for 2D mode map.
a, A single epifluorescence image of a single array. b, An average of five images taken of five different PCC arrays, as in Fig. 3c. Scale bars, 23 μm.
Extended Data Figure 7 Raw fluorescence data demonstrating digital control of cavity emission.
The central panel diagrams eight different arrangements of origami (white triangles) within PCCs, with the number of origami ranging from zero to seven. The left panel shows an epifluorescence image of eight PCC arrays, each containing 512 copies of a PCC with the origami arrangement diagrammed in the adjacent row of the central panel; each origami has n = 3 Cy5. The right panel shows an analogous epifluorescence image of eight PCC arrays for which each origami has n = 15 Cy5. Histograms summarizing these epifluorescence data are given in Extended Data Fig. 8. See Fig. 4b for plots showing the linearity of intensity as the number of origami are increased. Scale bars, 50 μm.
Extended Data Figure 8 Histograms of numerical data demonstrating digital control of cavity emission.
Eight different arrangements of origami were loaded into PCCs, with the number of origami ranging from zero to seven (see centre panel, Extended Fig. 7). 512 identical PCCs of each of the nonzero arrangements were imaged by epifluorescence microscopy (Extended Data Fig. 7, left and right panels) and their intensity was quantified. These data, for n = 3 Cy5 origami (a) and n = 15 Cy5 origami (b) are represented by coloured histograms labelled ‘one’ through ‘seven’. The number of PCCs with a given fluorescence intensity is plotted on the y-axis. c, Summary of means and standard deviations (for intensity measured as counts) from Gaussian fits in a, b. The peaks of these histograms are plotted in Fig. 4b.
Extended Data Figure 9 Schema for recreation of Vincent van Gogh’s painting The Starry Night (1889).
See Fig. 4c for recreation. a, Original image from Wikimedia Commons (https://commons.wikimedia.org/wiki/File:VanGogh-starry_night_ballance1.jpg). Because our current emitters and cavities provide only one color, with only eight levels of intensity, the original image was converted to greyscale (b) and then further converted to have three-bit colour depth (c). Because of the difficulty of handling extremely large suspended SiN membranes (which easily break) we limited our recreation to a low-resolution 256 × 256 = 65,536 pixel version, which was then further broken down into sixteen 64 × 64 pixel arrays (d). With one PCC per pixel, these arrays were small enough that we could fabricate and handle the resulting SiN membranes without too much breakage, image them separately, and then reassemble the sixteen images into a 4 × 4 array for the final recreation (Fig. 4c).
Supplementary information
Supplementary Data
This file contains Supplementary Data including (1) a DNA origami caDNAno design file and corresponding Microsoft Excel sequence list, (2) PCC design files as Matlab and Autocad scripts, and (3) FDTD simulations as Lumerical scripts. These files have been combined as a single .zip file. (ZIP 287 kb)
Rights and permissions
About this article
Cite this article
Gopinath, A., Miyazono, E., Faraon, A. et al. Engineering and mapping nanocavity emission via precision placement of DNA origami. Nature 535, 401–405 (2016). https://doi.org/10.1038/nature18287
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18287
This article is cited by
-
Site-directed placement of three-dimensional DNA origami
Nature Nanotechnology (2023)
-
A DNA origami rotary ratchet motor
Nature (2022)
-
Deterministic assembly of single emitters in sub-5 nanometer optical cavity formed by gold nanorod dimers on three-dimensional DNA origami
Nano Research (2022)
-
Spontaneous emission in micro- or nanophotonic structures
PhotoniX (2021)
-
A synthetic tubular molecular transport system
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.