Abstract
The drug/metabolite transporter (DMT) superfamily is a large group of membrane transporters ubiquitously found in eukaryotes, bacteria and archaea, and includes exporters for a remarkably wide range of substrates, such as toxic compounds and metabolites1. YddG is a bacterial DMT protein that expels aromatic amino acids and exogenous toxic compounds, thereby contributing to cellular homeostasis2,3. Here we present structural and functional analyses of YddG. Using liposome-based analyses, we show that Escherichia coli and Starkeya novella YddG export various amino acids. The crystal structure of S. novella YddG at 2.4 Å resolution reveals a new membrane transporter topology, with ten transmembrane segments in an outward-facing state. The overall structure is basket-shaped, with a large substrate-binding cavity at the centre of the molecule, and is composed of inverted structural repeats related by two-fold pseudo-symmetry. On the basis of this intramolecular symmetry, we propose a structural model for the inward-facing state and a mechanism of the conformational change for substrate transport, which we confirmed by biochemical analyses. These findings provide a structural basis for the mechanism of transport of DMT superfamily proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jack, D. L., Yang, N. M. & Saier, M. H. The drug/metabolite transporter superfamily. Eur. J. Biochem. 268, 3620–3639 (2001)
Santiviago, C. A. et al. The Salmonella enterica sv. Typhimurium smvA, yddG and ompD (porin) genes are required for the efficient efflux of methyl viologen. Mol. Microbiol. 46, 687–698 (2002)
Doroshenko, V. et al. YddG from Escherichia coli promotes export of aromatic amino acids. FEMS Microbiol. Lett. 275, 312–318 (2007)
Nikaido, H. Prevention of drug access to bacterial targets: permeability barriers and active efflux. Science 264, 382–388 (1994)
Paulsen, I. T., Brown, M. H. & Skurray, R. A. Proton-dependent multidrug efflux systems. Microbiol. Rev. 60, 575–608 (1996)
Poole, K. Efflux-mediated multiresistance in Gram-negative bacteria. Clin. Microbiol. Infect. 10, 12–26 (2004)
Piddock, L. J. V. Multidrug-resistance efflux pumps - not just for resistance. Nature Rev. Microbiol. 4, 629–636 (2006)
Krämer, R. Systems and mechanisms of amino acid uptake and excretion in prokaryotes. Arch. Microbiol. 162, 1–13 (1994)
Livshits, V. A., Zakataeva, N. P., Aleshin, V. V. & Vitushkina, M. V. Identification and characterization of the new gene rhtA involved in threonine and homoserine efflux in Escherichia coli. Res. Microbiol. 154, 123–135 (2003)
Bellman, A. et al. Expression control and specificity of the basic amino acid exporter LysE of Corynebacterium glutamicum. Microbiology 147, 1765–1774 (2001)
Ishida, N. & Kawakita, M. Molecular physiology and pathology of the nucleotide sugar transporter family (SLC35). Pflugers Arch. 447, 768–775 (2004)
Song, Z. Roles of the nucleotide sugar transporters (SLC35 family) in health and disease. Mol. Aspects Med. 34, 590–600 (2013)
Lolkema, J. S., Dobrowolski, A. & Slotboom, D. J. Evolution of antiparallel two-domain membrane proteins: tracing multiple gene duplication events in the DUF606 family. J. Mol. Biol. 378, 596–606 (2008)
Västermark, Å., Almén, M. S., Simmen, M. W., Fredriksson, R. & Schiöth, H. B. Functional specialization in nucleotide sugar transporters occurred through differentiation of the gene cluster EamA (DUF6) before the radiation of Viridiplantae. BMC Evol. Biol. 11, 123 (2011)
Bay, D. C. & Turner, R. J. Diversity and evolution of the small multidrug resistance protein family. BMC Evol. Biol. 9, 140 (2009)
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006)
Hattori, M., Hibbs, R. E. & Gouaux, E. A fluorescence-detection size-exclusion chromatography-based thermostability assay for membrane protein precrystallization screening. Structure 20, 1293–1299 (2012)
Airich, L. G. et al. Membrane topology analysis of the Escherichia coli aromatic amino acid efflux protein YddG. J. Mol. Microbiol. Biotechnol. 19, 189–197 (2010)
Rapp, M., Seppälä, S., Granseth, E. & von Heijne, G. Emulating membrane protein evolution by rational design. Science 315, 1282–1284 (2007)
Schuldiner, S. EmrE, a model for studying evolution and mechanism of ion-coupled transporters. Biochim. Biophys. Acta 1794, 748–762 (2009)
Chen, Y.-J. et al. X-ray structure of EmrE supports dual topology model. Proc. Natl Acad. Sci. USA 104, 18999–19004 (2007)
Forrest, L. R. Structural biology. (Pseudo-)symmetrical transport. Science 339, 399–401 (2013)
Liao, J. et al. Structural insight into the ion-exchange mechanism of the sodium/calcium exchanger. Science 335, 686–690 (2012)
Mancusso, R., Gregorio, G. G., Liu, Q. & Wang, D.-N. Structure and mechanism of a bacterial sodium-dependent dicarboxylate transporter. Nature 491, 622–626 (2012)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
De La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997)
Abrahams, J. P. & Leslie, A. G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
Cowtan, K. D. & Main, P. Improvement of macromolecular electron-density maps by the simultaneous application of real and reciprocal space constraints. Acta Crystallogr. D 49, 148–157 (1993)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: Building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Hattori, M. et al. Mg2+-dependent gating of bacterial MgtE channel underlies Mg2+ homeostasis. EMBO J. 28, 3602–3612 (2009)
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006)
Masuda, A. & Dohmae, N. Examination of an absolute quantity of less than a hundred nanograms of proteins by amino acid analysis. Anal. Bioanal. Chem. 405, 8073–8081 (2013)
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS ONE 6, e28766 (2011)
Hopf, T. A. et al. Three-dimensional structures of membrane proteins from genomic sequencing. Cell 149, 1607–1621 (2012)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–8, 27–8 (1996)
Brooks, B. R. et al. CHARMM: the biomolecular simulation program. J. Comput. Chem. 30, 1545–1614 (2009)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005)
Feller, S. E., Zhang, Y., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation: The Langevin piston method. J. Chem. Phys. 103, 4613 (1995)
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089 (1993)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
Acknowledgements
We thank H. Nishimasu and M. Hattori for comments on the manuscript; T. Tsukazaki and D. Drew for discussion; Y. Lee and A. Kurabayashi for technical assistance; the RIKEN BioResource Center for providing Starkeya novella genomic DNA; and the beam-line scientists at BL41XU and BL32XU of SPring-8 for assistance with data collection. The diffraction experiments were performed at SPring-8 BL41XU and BL32XU (proposals 2014A1091, 2014A1061, 2014A1093, 2014A1116 and 2014B1194). This work was supported by grants from the Platform for Drug Discovery, Informatics and Structural Life Science by the Ministry of Education, Culture, Sports, Science and Technology (MEXT), JSPS KAKENHI (grants 24227004, 25291011 and 26711003), the FIRST program, and a Grant-in-Aid for JSPS Fellows. Partial calculations were performed on the HOKUSAI GreatWave supercomputer system at RIKEN and the NIG supercomputer at ROIS National Institute of Genetics.
Author information
Authors and Affiliations
Contributions
H.T., S.D., T.H., and E.T. constructed the expression systems. H.T. expressed, purified, and crystallized SnYddG, collected the diffraction data, determined the structure, and performed liposome-based assays. K.T. and S.N. assisted with the liposome-based assays. M.T., T.N. and R.I. assisted with the structure determinations. K.I. constructed the E. coli knockout strain. K.F. and Y.U. performed the metabolomics analysis. M.T. and T.I. performed the molecular dynamics simulations. N.D. performed disulphide-bond quantification. H.T., R.I. and O.N. wrote the manuscript. R.I. and O.N. directed and supervised all of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
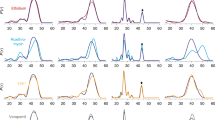
Extended Data Figure 1 Metabolomics analysis of the E. coli W3110 wild-type (WT) and YddG knockout (ΔyddG) strains, containing either the empty vector or the EcYddG protein expression vector.
We analysed the 110 well-known metabolites in the culture media of wild-type and ΔyddG cells, transformed with an empty vector or the pET-derived vector containing the EcYddG gene. The amounts of the metabolites in the media that exhibited more than threefold increases in both the wild-type and ΔyddG strains are plotted. The results showed the consistent increases of valine, threonine and isoleucine, in the medium of the EcYddG-expressing cells, as compared to the negative controls. These results are consistent with those from the in vitro liposome-based analysis of EcYddG (Fig. 1), supporting the notion that YddG transports a wide range of metabolites, as well as amino acids.
Extended Data Figure 2 The 2.4-Å electron density map and molecular surface of SnYddG.
a, b, A stereo view of the unbiased 2mFo − DFc maps around TM4 and TM9 (a), and an overall view of molecule B in the crystal asymmetric unit, contoured at 1.1σ (b) are shown. c, Molecular surface of SnYddG, viewed from different directions. The transmembrane segments are coloured as in Fig. 2.
Extended Data Figure 3 Conserved residues of YddG and SMR homologues.
a, Amino acid sequence alignment of SnYddG and other YddG homologues, from Bradyrhizobium japonicum (UniProtKB ID: A0A0A3YL06), Virgibacillus halodenitrificans (GenBank ID: CDQ34001.1), Serratia marcescens (UniProtKB ID: L0MHZ3), Pectobacterium carotovorum (UniProtKB ID: A0A0E2ZTH4), E. coli (UniProtKB ID: P46136), Vibrio vulnificus (UniProtKB ID: Q7MDM6), Acinetobacter baumannii (UniProtKB ID: A0A009YGQ0), Bifidobacterium merycicum (UniProtKB ID: A0A087BD75), Phaeobacter inhibens (UniProtKB ID: I7EXD9), and Pseudomonas sp. (UniProtKB ID: B2Z3V9). The conserved amino acids involved in the transport mechanism and discussed in the main text are indicated. b, Amino acid sequence alignment of the N and C halves of SnYddG. c, Amino acid sequence alignment of EmrE homologues, from E. coli (UniProtKB ID: P23895), S. enterica (UniProtKB ID: X5AXH0), Xanthobacter autotrophicus (UniProtKB IDs: A7IJY9 and A7IN30), Saccharomonospora viridis (UniProtKB IDs: C7MRM1 and C7MSE0), Thermobifida fusca (UniProtKB IDs: Q47R92 and Q47QH0), Thermus thermophilus (UniProtKB ID: Q72K82), Thermobispora bispora (UniProtKB IDs: D6YBB5 and D6Y4B6), Thermomonospora curvata (UniProtKB IDs: D1A404 and D1AD18), and Zymomonas mobilis (UniProtKB IDs: Q5NPN9 and Q5NRC2).
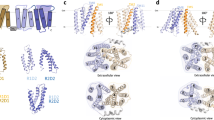
Extended Data Figure 4 Molecular evolution of the DMT superfamily.
a, Crystal structure of E. coli EmrE (PDB accession 3B5D)21. The bound tetraphenylphosphonium (TPP) is shown as a stick model. The transmembrane segments are coloured and labelled as in d. b, The present crystal structure of SnYddG. Only the transmembrane segments are shown in cylinder models. The transmembrane segments are coloured and labelled as in d. c, Molecular superimposition of EmrE and SnYddG, calculated by the SSM algorithm43. The cylinder model of EmrE is shown with the same colouring as in d. The semi-transparent cylinder model of SnYddG is shown with the same colouring as in d. d, Schematic illustration of the possible evolutionary path of the DMT family transporters. The transmembrane segments are indicated by circles, and the connecting loops are indicated by solid and dotted curves. The black arrows indicate the folding of the transporter, while the grey arrows indicate the possible evolutional pathway. Transmembrane segments originating from one protomer of SMR are coloured blue, while the newly inserted transmembrane segments (TM a) are coloured red.
Extended Data Figure 5 Preparation of the reconstituted proteoliposomes containing the SnYddG mutants.
a, The chromatograms of the final purification step by the gel-filtration chromatography column. The results of the wild type, as well as the mutants with the decreased activities in the liposome-based assay (that is, G71A, H79A and G222A), are shown. b, Reconstitution rates of the wild-type and SnYddG mutants, determined by the SDS–PAGE analysis. The reconstitution rates were measured as a percentage of the total amount of protein used for the reconstitution.
Extended Data Figure 6 Cys-crosslink analysis of the residues on the periplasmic sides of TM1 (Ala21, Thr24) and TM3 (Ala85, Leu86, Pro91, Ala92).
a, The Cys-free mutant of SnYddG (C159A/C185A/C195A/C271A) was created, and then pairs of Cys residues to the residues in TM1 and TM3 or TM1 and TM4, which may form the intracellular gate, were introduced. The purified double-Cys mutant proteins were oxidized or reduced, and then modified by the Cys-reactive fluorescent reagent, tetramethylrhodamine maleimide (TMRM). The double-Cys mutants were completely masked from the Cys-modification by TMRM under the oxidized conditions, indicating that these residues are close enough to form disulfide bonds. b, The results of the quantification of the amino acid composition of the cross-linking products of the YddG double-Cys mutants. The percentages of disulfide bond formation for each mutant were calculated from the amounts of carboxymethylated Cys under the oxidizing and reducing conditions. c, The positions of the mutated residues are indicated in the crystal structure viewed from the periplasmic side.
Extended Data Figure 7 Evolutionary covariation analysis of YddG and DMT protein homologues.
a, Contact maps of the top-ranked 170 evolutionary constraints (ECs), calculated from the 59,114 homologue sequences using the program EVcoupling. The ECs are indicated by stars, coloured proportionally according to their EC scores, from orange to red. The solid and dashed rectangles indicate the ECs corresponding to the observed extracellular gate and the putative intracellular gate interactions, respectively. b, The ECs mapped onto the outward-open crystal structure of SnYddG. The ECs indicated in a and tabulated in the top and bottom panels are indicated as arrows connecting the corresponding residues in the SnYddG structure. The solid and dashed arrows indicate the ECs corresponding to the observed extracellular gate and the putative intracellular gate interactions, respectively. c, The SnYddG structure predicted from the top-ranked 170 ECs, using the program EVfold_membrane.
Extended Data Figure 8 The kinks of TM3 and TM8 around the conserved Gly and Pro residues.
a, The kinks of TM3 around the conserved Gly residues, in the outward-open crystal structure and the inward-open model. b, The kinks of TM8 around the conserved Gly and Pro residues, in the outward-open crystal structure and the inward-open model. In a and b, the Gly and Pro residues are shown by stick models, and the axes of the transmembrane helices are indicated by cylinders. In the right panel, the activities of SnYddG mutants of the TM3 and TM8 kinks, measured by the liposome-based analysis using threonine as a substrate, are plotted, respectively. The substrate uptake activities measured after 30 min were plotted as percentages of the wild-type transport activity. Error bars, s.d.; n = 3.
Extended Data Figure 9 Molecular dynamics simulation of SnYddG.
a, The results of the 500-ns run of the non-biased simulation. Left, a plot of the root mean square fluctuations of each Cα atom from the initial crystal structure. Right, a plot of the root mean square deviation over all Cα atoms from the initial crystal structure, during the course of the simulation. b, The results of the outward-to-occluded simulation. Top left, a plot of the distances between the Cα atoms of TM4a (Pro91–Ala98) and TM9b (Ala246–Leu253), and between TM4b (Trp101–Phe108) and TM9a (Val237–Ser244). The equilibrium distance of the harmonic restraint applied between TM4a and TM9b is plotted with a red line. Top right, time series of the distances between the Cα atoms of Ser167 (TM6) and Trp228 (TM8), and between Ala21 (TM1) and Leu86 (TM3). Bottom left, time series of the distances between the Cα atoms of TM1 (17–24) and TM3 (81–88) and the Cα atoms of TM6 (163–170) and TM8 (225–232). Bottom right, the final snapshot of the outward-to-occluded simulation. The Cα atoms of Ala21 (TM1) and Leu86 (TM3) are shown as spheres. The parts of TM4a and TM9b subjected to harmonic restraints are coloured grey. The arrows indicate the distances plotted in the graphs in b and c. c, The results of the occluded-to-inward simulation. Top left, plot of the distance between the Cα atoms of TM4a (Pro91–Ala98) and TM9b (Ala246–Leu253), and between TM4b (Trp101–Phe108) and TM9a (Val237–Ser244). The equilibrium distance of the harmonic restraint applied between TM4b and TM9a is plotted with a red line. Top right, time series of the distances between the Cα atoms of Ser167 (TM6) and Trp228 (TM8), and between Ala21 (TM1) and Leu86 (TM3). Bottom left, time series of the distances between the Cα atoms of TM1 (17–24) and TM3 (81–88) and between the Cα atoms of TM6 (163–170) and TM8 (225–232). Bottom right, the final snapshot of the occluded-to-inward simulation. The Cα atoms of Ser167 (TM6) and Trp228 (TM8) are shown as spheres. The parts of TM4b and TM9a subjected to harmonic restraints are coloured grey. The arrows indicate the distances plotted in the graphs in b and c.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion and Supplementary References. (PDF 319 kb)
The outward-inward conformational change of SnYddG
The video sequence showing the conformational changes from the outward- to inward-facing state of SnYddG, viewed from four different directions. The structure is shown with rainbow coloring, with the N- and C-termini colored blue and red, respectively. The residues involved in the intracellular and extracellular gates are shown in stick models. (MP4 10466 kb)
Rights and permissions
About this article
Cite this article
Tsuchiya, H., Doki, S., Takemoto, M. et al. Structural basis for amino acid export by DMT superfamily transporter YddG. Nature 534, 417–420 (2016). https://doi.org/10.1038/nature17991
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17991
This article is cited by
-
Advances and prospects in metabolic engineering of Escherichia coli for L-tryptophan production
World Journal of Microbiology and Biotechnology (2022)
-
Structural and evolutionary analyses of the Plasmodium falciparum chloroquine resistance transporter
Scientific Reports (2020)
-
The structural basis of promiscuity in small multidrug resistance transporters
Nature Communications (2020)
-
A major and stable QTL controlling wheat thousand grain weight: identification, characterization, and CAPS marker development
Molecular Breeding (2020)
-
Antibiotic resistance related to biofilm formation in Streptococcus suis
Applied Microbiology and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.