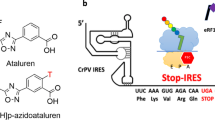
Abstract
Rocaglamide A (RocA) typifies a class of protein synthesis inhibitors that selectively kill aneuploid tumour cells and repress translation of specific messenger RNAs1,2,3,4. RocA targets eukaryotic initiation factor 4A (eIF4A), an ATP-dependent DEAD-box RNA helicase; its messenger RNA selectivity is proposed to reflect highly structured 5′ untranslated regions that depend strongly on eIF4A-mediated unwinding5. However, rocaglate treatment may not phenocopy the loss of eIF4A activity, as these drugs actually increase the affinity between eIF4A and RNA1,2,6. Here we show that secondary structure in 5′ untranslated regions is only a minor determinant for RocA selectivity and that RocA does not repress translation by reducing eIF4A availability. Rather, in vitro and in cells, RocA specifically clamps eIF4A onto polypurine sequences in an ATP-independent manner. This artificially clamped eIF4A blocks 43S scanning, leading to premature, upstream translation initiation and reducing protein expression from transcripts bearing the RocA–eIF4A target sequence. In elucidating the mechanism of selective translation repression by this lead anti-cancer compound, we provide an example of a drug stabilizing sequence-selective RNA–protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cencic, R. et al. Antitumor activity and mechanism of action of the cyclopenta[b]benzofuran, silvestrol. PLoS ONE 4, e5223 (2009)
Bordeleau, M. E. et al. Therapeutic suppression of translation initiation modulates chemosensitivity in a mouse lymphoma model. J. Clin. Invest. 118, 2651–2660 (2008)
Liu, T. et al. Synthetic silvestrol analogues as potent and selective protein synthesis inhibitors. J. Med. Chem. 55, 8859–8878 (2012)
Santagata, S. et al. Tight coordination of protein translation and HSF1 activation supports the anabolic malignant state. Science 341, 1238303 (2013)
Wolfe, A. L. et al. RNA G-quadruplexes cause eIF4A-dependent oncogene translation in cancer. Nature 513, 65–70 (2014)
Sadlish, H. et al. Evidence for a functionally relevant rocaglamide binding site on the eIF4A-RNA complex. ACS Chem. Biol. 8, 1519–1527 (2013)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Ingolia, N. T. et al. Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Reports 8, 1365–1379 (2014)
Sonenberg, N. & Hinnebusch, A. G. Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell 136, 731–745 (2009)
Pestova, T. V., Shatsky, I. N., Fletcher, S. P., Jackson, R. J. & Hellen, C. U. A prokaryotic-like mode of cytoplasmic eukaryotic ribosome binding to the initiation codon during internal translation initiation of hepatitis C and classical swine fever virus RNAs. Genes Dev. 12, 67–83 (1998)
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J. S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo . Nature 505, 701–705 (2014)
Bordeleau, M. E. et al. Functional characterization of IRESes by an inhibitor of the RNA helicase eIF4A. Nature Chem. Biol. 2, 213–220 (2006)
Lindqvist, L. et al. Selective pharmacological targeting of a DEAD box RNA helicase. PLoS ONE 3, e1583 (2008)
Hsieh, A. C. et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 485, 55–61 (2012)
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012)
Lambert, N. et al. RNA Bind-n-Seq: quantitative assessment of the sequence and structural binding specificity of RNA binding proteins. Mol. Cell 54, 887–900 (2014)
Linder, P. & Jankowsky, E. From unwinding to clamping – the DEAD box RNA helicase family. Nature Rev. Mol. Cell Biol. 12, 505–516 (2011)
Abramson, R. D. et al. The ATP-dependent interaction of eukaryotic initiation factors with mRNA. J. Biol. Chem. 262, 3826–3832 (1987)
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nature Struct. Mol. Biol . 17, 909–915 (2010)
Pestova, T. V. & Kolupaeva, V. G. The roles of individual eukaryotic translation initiation factors in ribosomal scanning and initiation codon selection. Genes Dev. 16, 2906–2922 (2002)
Parsyan, A. et al. mRNA helicases: the tacticians of translational control. Nature Rev. Mol. Cell Biol. 12, 235–245 (2011)
Pause, A. & Sonenberg, N. Mutational analysis of a DEAD box RNA helicase: the mammalian translation initiation factor eIF-4A. EMBO J. 11, 2643–2654 (1992)
Shah, P., Ding, Y., Niemczyk, M., Kudla, G. & Plotkin, J. B. Rate-limiting steps in yeast protein translation. Cell 153, 1589–1601 (2013)
Shirokikh, N. E. et al. Quantitative analysis of ribosome-mRNA complexes at different translation stages. Nucleic Acids Res. 38, e15 (2010)
Balvay, L., Soto Rifo, R., Ricci, E. P., Decimo, D. & Ohlmann, T. Structural and functional diversity of viral IRESes. Biochim. Biophys. Acta 1789, 542–557 (2009)
Kozak, M. Downstream secondary structure facilitates recognition of initiator codons by eukaryotic ribosomes. Proc. Natl Acad. Sci. USA 87, 8301–8305 (1990)
Medenbach, J., Seiler, M. & Hentze, M. W. Translational control via protein-regulated upstream open reading frames. Cell 145, 902–913 (2011)
Arribere, J. A. & Gilbert, W. V. Roles for transcript leaders in translation and mRNA decay revealed by transcript leader sequencing. Genome Res. 23, 977–987 (2013)
Lee, S. et al. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl Acad. Sci. USA 109, E2424–E2432 (2012)
Pommier, Y. & Marchand, C. Interfacial inhibitors: targeting macromolecular complexes. Nature Rev. Drug Discov . 11, 25–36 (2011)
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nature Protocols 7, 1534–1550 (2012)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011)
Dmitriev, S. E., Pisarev, A. V., Rubtsova, M. P., Dunaevsky, Y. E. & Shatsky, I. N. Conversion of 48S translation preinitiation complexes into 80S initiation complexes as revealed by toeprinting. FEBS Lett. 533, 99–104 (2003)
Iwasaki, S. et al. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 39, 292–299 (2010)
Busso, D., Delagoutte-Busso, B. & Moras, D. Construction of a set Gateway-based destination vectors for high-throughput cloning and expression screening in Escherichia coli . Anal. Biochem. 343, 313–321 (2005)
Galicia-Vázquez, G., Cencic, R., Robert, F., Agenor, A. Q. & Pelletier, J. A cellular response linking eIF4AI activity to eIF4AII transcription. RNA 18, 1373–1384 (2012)
Acknowledgements
We are grateful to J. Tanaka for providing hippuristanol, to Y. Tomari for sharing DNA constructs, to H. Asahara and University of California, Berkeley DNA sequencing facility for help with the toeprinting assay, and to A. Pinder and F. Tan for support with deep sequencing analysis. We also thank the members of Ingolia, Lareau, and Tomari laboratories for discussion and technical support. N.T.I. is a Damon-Runyon-Rachleff Innovator supported in part by the Damon Runyon Cancer Research Foundation (DRR-37-15), the Searle Scholars Program (11-SSP-229), and the National Institute of General Medical Sciences of the National Institutes of Health (P50GM102706). This work used the Vincent J. Coates Genomics Sequencing Laboratory at University of California, Berkeley, supported by National Institutes of Health S10 Instrumentation Grants S10RR029668, S10RR027303, and OD018174. S.I. is a recipient of Human Frontier Science Program long-term fellowship. S.N.F. is a Howard Hughes Medical Institute Fellow of the Helen Hay Whitney Foundation.
Author information
Authors and Affiliations
Contributions
S.I. performed all experiments and analysed the data. Recombinant protein purification and the fluorescence polarization assay were performed with the help of S.N.F. S.I. and N.T.I. designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 RocA represses translation, targeting to eIF4A.
a, Polysome profiling experiments with RocA and PP242 treatments. RocA disrupts polysomes dose-dependently. b, Western blot of phospho-eIF2α and phospho-4EBP shows that effect of RocA is independent of known translation control targeting to eIFs. Phosphorylation of eIF2α and dephosphorylation of 4EBP were induced by thapsigargin and PP242, respectively. c, d, Luciferase reporter assay possessing PTGES3 5′ UTR (Fig. 1c) with exogenous expression of WT or RocA-resistant eIF4A mutants (c) and western blot of endogenous and exogenous eIF4A (d). eIF4A is the main molecular target of RocA. Data represent mean and s.d. (n = 3). e, f, Correlation of sum of the footprint reads to 13 mitochondrial mRNAs among different conditions (e) and correlation of sum of the footprint reads from cytoplasmic ribosomes to each transcript between biological replicates (f). Symbol r is Pearson’s correlation. P value is calculated by Student’s t-test. g, h, Tile plot of codon periodicity along length of mitochondria footprints (g, left) and mitochondria footprint length distribution (g, right) and codon periodicities of 31-nt mitochondrial footprints among different conditions (h). Footprints with 31-nt length showed most homogenous codon periodicity, and this periodicity was retained with RocA treatment, showing that mitochondrial ribosome translates even in high doses of RocA.
Extended Data Figure 2 RocA represses translation without mRNA degradation.
a, Metabolic labelling of nascent peptides with OP-puro. The OP-puro incorporated nascent peptides were visualized by Click reaction with Alexa Fluor 488 Azide (middle) and quantified (right). Data represent mean and s.d. (n = 3). b, Correlation of translation -fold change among different concentrations of RocA treatments. c, MA plot of mean footprint reads between 0.03 μM RocA treatment and non-treatment normalized to library sizes to footprints -fold change by 0.03 μM RocA treatment (left) and the correlation of translation -fold change between 0.03 and 3 μM of RocA treatments (right), highlighting high-sensitivity mRNAs at 0.03 μM RocA treatment. d, Scatter plots of footprint -fold change normalized to mitochondrial footprints and mRNA -fold change by RocA treatments. RocA represses translation without significatnt mRNA change. e, qPCR from the samples of Fig. 1c. Data represent mean and s.d. (n = 3).
Extended Data Figure 3 Secondary structure in 5′ UTR is not strong determinant of RocA sensitivity.
a, Cumulative fractions along length of 5′ UTR, minimum ΔG among all 30-mer windows along a 5′ UTR, ΔG in cap-proximal region (30 nt) of 5′ UTR, and Gini difference are plotted to total, RocA high-sensitivity, and RocA low-sensitivity mRNAs. Significance is calculated by Mann–Whitney U-test. b, Cumulative fractions along translation -fold change by RocA are plotted to total mRNAs and mRNAs with predicted G-quadruplexes in 5′ UTRs. Significance is calculated by Mann–Whitney U-test. The impact of presence of G-quadruplex in 5′ UTR is modest in RocA sensitivity. c, The 5′ UTRs with G-quadruplexes and randomized control sequence were fused to Renilla luciferase and these reporter mRNAs were transfected before treatment with RocA as indicated. Data represent mean and s.d. (n = 3). G-quadruplex does not show the prominent RocA sensitivity.
Extended Data Figure 4 Characterization of translational inhibition by Hippuristanol and PP242.
a, Polysome profiling experiments with Hipp treatments. Hipp disrupts polysomes dose-dependently. b, Histograms of number of transcripts along footprints -fold change with 0.01 and 1 μM Hipp treatment compared with non-treatment, normalized to mitochondrial footprints. Median -fold change is shown. Bin width is 0.1. c, MA plot of mean footprint reads between 1 μM Hipp treatment and non-treatment normalized to library sizes to translation -fold change by 1 μM Hipp treatment, highlighting high-sensitivity and low-sensitivity mRNAs. d, Cumulative fractions along length of 5′ UTR, minimum ΔG among all 30-mer windows along a 5′ UTR, ΔG in cap-proximal region (30 nt) of 5′ UTR, and Gini difference are plotted to total, Hipp high-sensitivity, and Hipp low-sensitivity mRNAs. Significance is calculated by Mann–Whitney U-test. e, Translation -fold changes by RocA and Hipp are modestly correlated. f, MA plot of mean footprint reads between 2.5 μM PP242 treatment and non-treatment normalized to library sizes to translation -fold change by PP242 treatment, highlighting PP242 target mRNAs. g, Cumulative distributions of translation -fold change caused by RocA and Hipp treatment are plotted for total and PP242-target mRNAs. Significance is calculated by Mann–Whitney U-test.
Extended Data Figure 5 Purification of SBP-tagged eIF4A and co-purified RNA from HEK 293 cells.
a, Western blot of exogenous SBP-eIF4A and endogenous eIF4A in tetracycline-inducible stable cell line. Expression of physiological levels of the tagged allele attenuated endogenous eIF4A expression but preserved overall eIF4A levels, probably reflecting the same feedback loop previously reported between eIF4AI and eIF4AII37. b, CBB staining of purified SBP-eIF4A and SYBR Gold staining of purified RNA bound to SBP-eIF4A with or without micrococcal nuclease (MNase). c, Correlation of sum of the mRNA fragment reads of each transcript between biological replicates of RIP-seq. P value is calculated by Student’s t-test. d, Histogram of the number of transcripts along RNA/eIF4A interaction -fold change by RIP-seq when cells are treated with 0.03 or 0.3 μM RocA normalized to spiked-in RNA. Data present the same mRNAs analysed in Fig. 1a. Median -fold change is shown. Bin width is 0.1. e, Correlation of RIP -fold change between different concentration of RocA treatments. f, Correlation of translation -fold change to RIP -fold change with the same concentration of RocA treatment.
Extended Data Figure 6 Motif enrichment by Bind-n-Seq.
a, Nucleotide composition in each length of reads in input RNAs for Bind-n-Seq. Input RNAs are random in entire read length. b, Length distribution of reads from Bind-n-Seq. RNAs bound to eIF4A showed longer length distribution, indicating that eIF4A has preference for longer RNAs. c, Correlations of tetramer motif enrichment in Bind-n-Seq by 0.03 μM RocA treatment to that by 0.3 μM RocA treatment. d, Correlations between pentamer and hexamer motif enrichment in Bind-n-Seq by 0.03 μM RocA treatment and motif prediction of 0.03 μM RocA effect in RIP-seq. e, Highest-scoring pentamer and hexamer motifs in Bind-n-Seq and RIP-seq. f, Cumulative fractions along number of tetramer motifs (Fig. 2b) in 5′ UTR are plotted to total, RocA high-sensitivity, and RocA low-sensitivity mRNAs. Significance is calculated by Mann–Whitney U-test. g, Correlations of Bind-n-Seq motif enrichment (pentamer) by eIF4A to that by 0.03 μM RocA treatment. The motifs appearing in RNAs used in Extended Data Fig. 8 are highlighted. h, Correlation of Bind-n-Seq motif enrichment (pentamer) by eIF4A to motif prediction of Hipp effect in translation change, which is defined as Spearman’s correlation of motif number in 5′ UTR to translation -fold change by Hipp. mRNAs with high-affinity motif to eIF4A in 5′ UTR are resistant to Hipp treatment. i, The correlation between enriched motifs of replicates in Bind-n-Seq with ADP + Pi.
Extended Data Figure 7 Characterization of iCLIP data.
a, CBB staining of purified SBP-eIF4A protein in iCLIP procedure. b, Visualization of RNA-crosslinked with SBP-eIF4A and unknown proteins by 32P labelling of RNA. We avoided the contamination of RNAs cross-linked to the additional, co-purifying, unknown proteins. c, Distribution of read length in iCLIP libraries. Avoidance of contaminating RNAs restricted us to short RNAs, which probably correspond to the region of RNA physically protected by eIF4A binding, or footprint. d, Nucleotide bias along the reads in iCLIP libraries. The crosslinking bias for U may underestimate the preference for polypurine motifs. e, Correlations of iCLIP motif enrichment (tetramer) by different RocA concentrations. f, Correlations of iCLIP motif enrichment (tetramer) by 3 μM RocA and motif prediction of 0.03 μM RocA effect in RIP-seq. The motifs shown in Fig. 2a are highlighted.
Extended Data Figure 8 eIF4A/RNA affinity measured by fluorescence polarization.
a, CBB staining of recombinant proteins used in this study. b, Summary of Kd between RNA and eIF4A among the conditions assayed. c, e–g, i, Direct measurement of the eIF4A/RNA affinity by fluorescence polarization for eIF4A WT, eIF4A (VX4GKT), or eIF4A (D296A–T298K) and 5′ FAM-labelled RNAs in the presence or absence of RocA. Data represent mean and s.d. (n = 3). d, ATP crosslinking assay with eIF4A WT and eIF4A (VX4GKT). h, Pulldown assay with His-MBP-eIF4A expressed in E. coli and eIF4E/G in RRL.
Extended Data Figure 9 Characterization of toeprinting assay.
a, Diagram of the reporters used in this study. b, c, In vitro translation in RRL with mRNAs containing seven polypurine motif (AGAGAG) insertions (b) and qPCR from the samples (c). d, Dideoxy-terminated sequencing of RNA by reverse transcription verified the toeprinting product length terminated by 48S ribosomes. e, Ribosome toeprinting assay performed in RRL in the presence of m7-GTP in the presence or absence of 3 μM RocA treatment. f, Toeprinting assay using 10 μM recombinant eIF4A in the presence or absence of 10 μM RocA treatment. g, Toeprinting assay (top) and RNase I footprinting assay (bottom) using 10 μM recombinant eIF4A with mRNA containing one AGAGAG motif at the middle in the presence or absence of 10 μM RocA treatment. h, i, Toeprinting assay using 10 μM recombinant eIF4A (VX4GKT) or (D296A-T298K) with mRNA containing seven AGAGAG motifs in the presence or absence of 10 μM RocA treatment. j, Pre-formation of the complex with RocA and eIF4A (VX4GKT) or (D296A-T298K) on the mRNA bearing seven polypurine motifs represses the translation from the mRNA in RRL. k, Basal translation level from mRNA containing seven AGAGAG motifs with the supplementation of recombinant eIF4A. l, In vitro translation in RRL with mRNAs with a single polypurine motif (AGAGAG) insertion at the different positions in 5′ UTR. m, Basal translation level from mRNAs bearing polio virus IRES and polio virus IRES with three AGAGAG motifs. In b, c, and h–j, data represent mean and s.d. (n = 3).
Extended Data Figure 10 The 5′ UTR footprints accumulated in RocA treatments come from uORFs.
a, The distributions of specific footprint length, which is a hallmark of 80S ribosomes8, from CDS and 5′ UTR are indistinguishable. b, The change in ribosome footprint counts for 5′ UTRs and CDSs when cells are treated with 3 μM RocA or 1 μM Hipp compared with non-treatment, normalized to mitochondrial footprints. Median -fold change is shown. Bin width is 0.1. Analysis is restricted to mRNAs bearing footprints in the 5′ UTR in the non-treatment condition. c, Meta-gene analysis of low-sensitivity transcripts to RocA. Reads are normalized to the sum of mitochondrial footprints reads. d, The illustration of the definition of uORF translation intensity. e, Transcripts sensitive to RocA contain more active uORFs, as measured by cumulative distributions of the uORF translation intensity c. Significance is calculated by Mann–Whitney U-test. f, The summary of deep sequencing-based approaches used in this study and corresponding figures.
Supplementary information
Supplementary Table 1
High-sensitivity (a) and low-sensitivity (b) mRNAs to 3 µM RocA. Each transcript is listed with its UCSC identifier, translation -fold change to mean [log2], q value, translation -fold change normalized to mitochondria footprints [log2], length of 5′ UTR, minimum ΔG calculated along 5′ UTR with 30-mer window, gene name, and gene description. (c) High-sensitivity mRNAs to 0.03 µM RocA are listed the same as (a). (XLSX 179 kb)
Supplementary Table 2
High-sensitivity (a) and low-sensitivity (b) mRNAs to 1 µM Hipp are listed the same as Supplemental Table 1. (XLSX 248 kb)
Supplementary Table 3
(a) PP242 target mRNAs. Each transcript is listed with its UCSC identifier, translation -fold change to mean [log2], q value, translation -fold change normalized to mitochondria footprints [log2], gene name, and gene description. (XLSX 66 kb)
Supplementary Figure 1
This file contains the raw data for Figure 4e. (PDF 468 kb)
Rights and permissions
About this article
Cite this article
Iwasaki, S., Floor, S. & Ingolia, N. Rocaglates convert DEAD-box protein eIF4A into a sequence-selective translational repressor. Nature 534, 558–561 (2016). https://doi.org/10.1038/nature17978
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17978
This article is cited by
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
Nature Communications (2023)
-
Cellular functions of eukaryotic RNA helicases and their links to human diseases
Nature Reviews Molecular Cell Biology (2023)
-
Distance-dependent inhibition of translation initiation by downstream out-of-frame AUGs is consistent with a Brownian ratchet process of ribosome scanning
Genome Biology (2022)
-
Altered proteome in translation initiation fidelity defective eIF5G31R mutant causes oxidative stress and DNA damage
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.