Abstract
Myocardial infarction results in compromised myocardial function and heart failure owing to insufficient cardiomyocyte self-renewal1. Unlike many vertebrates, mammalian hearts have only a transient neonatal renewal capacity2. Reactivating primitive reparative ability in the mature mammalian heart requires knowledge of the mechanisms that promote early heart repair. By testing an established Hippo-deficient heart regeneration mouse model for factors that promote renewal, here we show that the expression of Pitx2 is induced in injured, Hippo-deficient ventricles. Pitx2-deficient neonatal mouse hearts failed to repair after apex resection, whereas adult mouse cardiomyocytes with Pitx2 gain-of-function efficiently regenerated after myocardial infarction. Genomic analyses indicated that Pitx2 activated genes encoding electron transport chain components and reactive oxygen species scavengers. A subset of Pitx2 target genes was cooperatively regulated with the Hippo pathway effector Yap. Furthermore, Nrf2, a regulator of the antioxidant response3, directly regulated the expression and subcellular localization of Pitx2. Pitx2 mutant myocardium had increased levels of reactive oxygen species, while antioxidant supplementation suppressed the Pitx2 loss-of-function phenotype. These findings reveal a genetic pathway activated by tissue damage that is essential for cardiac repair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Xin, M., Olson, E. N. & Bassel-Duby, R. Mending broken hearts: cardiac development as a basis for adult heart regeneration and repair. Nature Rev. Mol. Cell Biol. 14, 529–541 (2013)
Porrello, E. R. et al. Transient regenerative potential of the neonatal mouse heart. Science 331, 1078–1080 (2011)
Itoh, K. et al. Keap1 represses nuclear activation of antioxidant responsive elements by Nrf2 through binding to the amino-terminal Neh2 domain. Genes Dev. 13, 76–86 (1999)
Heallen, T. et al. Hippo signaling impedes adult heart regeneration. Development 140, 4683–4690 (2013)
Lu, M. F., Pressman, C., Dyer, R., Johnson, R. L. & Martin, J. F. Function of Rieger syndrome gene in left-right asymmetry and craniofacial development. Nature 401, 276–278 (1999)
Semina, E. V. et al. Cloning and characterization of a novel bicoid-related homeobox transcription factor gene, RIEG, involved in Rieger syndrome. Nature Genet. 14, 392–399 (1996)
Wang, J. et al. Pitx2 prevents susceptibility to atrial arrhythmias by inhibiting left-sided pacemaker specification. Proc. Natl Acad. Sci. USA 107, 9753–9758 (2010)
Kirchhof, P. et al. PITX2c is expressed in the adult left atrium, and reducing Pitx2c expression promotes atrial fibrillation inducibility and complex changes in gene expression. Circ. Cardiovasc. Genet. 4, 123–133 (2011)
Giudice, J. et al. Alternative splicing regulates vesicular trafficking genes in cardiomyocytes during postnatal heart development. Nature Commun . 5, 3603 (2014)
Nord, A. S. et al. Rapid and pervasive changes in genome-wide enhancer usage during mammalian development. Cell 155, 1521–1531 (2013)
Bruning, J. C. et al. A muscle-specific insulin receptor knockout exhibits features of the metabolic syndrome of NIDDM without altering glucose tolerance. Mol. Cell 2, 559–569 (1998)
Chan, K., Lu, R., Chang, J. C. & Kan, Y. W. NRF2, a member of the NFE2 family of transcription factors, is not essential for murine erythropoiesis, growth, and development. Proc. Natl Acad. Sci. USA 93, 13943–13948 (1996)
Xin, M. et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc. Natl Acad. Sci. USA 110, 13839–13844 (2013)
L’Honoré, A. et al. Redox regulation by Pitx2 and Pitx3 is critical for fetal myogenesis. Dev. Cell 29, 392–405 (2014)
Dhalla, N. S., Temsah, R. M. & Netticadan, T. Role of oxidative stress in cardiovascular diseases. J. Hypertens. 18, 655–673 (2000)
Larsson, N. G. & Clayton, D. A. Molecular genetic aspects of human mitochondrial disorders. Annu. Rev. Genet. 29, 151–178 (1995)
Yue, F. et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355–364 (2014)
Zanconato, F. et al. Genome-wide association between YAP/TAZ/TEAD and AP-1 at enhancers drives oncogenic growth. Nature Cell Biol. 17, 1218–1227 (2015)
Galli, G. G. et al. YAP drives growth by controlling transcriptional pause release from dynamic enhancers. Mol. Cell 60, 328–337 (2015)
Stein, C. et al. YAP1 exerts its transcriptional control via TEAD-mediated activation of enhancers. PLoS Genet. 11, e1005465 (2015)
Morikawa, Y. et al. Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice. Sci. Signal. 8, ra41 (2015)
Puente, B. N. et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 157, 565–579 (2014)
Chouchani, E. T. et al. Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature 515, 431–435 (2014)
Shao, D. et al. A functional interaction between Hippo-YAP signalling and FoxO1 mediates the oxidative stress response. Nature Commun . 5, 3315 (2014)
Heallen, T. et al. Hippo pathway inhibits Wnt signaling to restrain cardiomyocyte proliferation and heart size. Science 332, 458–461 (2011)
Lavado, A., Lagutin, O. V., Chow, L. M., Baker, S. J. & Oliver, G. Prox1 is required for granule cell maturation and intermediate progenitor maintenance during brain neurogenesis. PLoS Biol. 8, e1000460 (2010)
Xin, M. et al. Regulation of insulin-like growth factor signaling by Yap governs cardiomyocyte proliferation and embryonic heart size. Sci. Signal 4, ra70 (2011)
Heallen, T. et al. Hippo signaling impedes adult heart regeneration. Development 140, 4683–4690 (2013)
Nascimento, D. S. et al. MIQuant–semi-automation of infarct size assessment in models of cardiac ischemic injury. PLoS ONE 6, e25045 (2011)
Porrello, E. R. et al. Regulation of neonatal and adult mammalian heart regeneration by the miR-15 family. Proc. Natl Acad. Sci. USA 110, 187–192 (2013)
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nature Protocols 8, 2281–2308 (2013)
Romanoski, C. E., Glass, C. K., Stunnenberg, H. G., Wilson, L. & Almouzni, G. Epigenomics: Roadmap for regulation. Nature 518, 314–316 (2015)
Amen, M. et al. Chromatin-associated HMG-17 is a major regulator of homeodomain transcription factor activity modulated by Wnt/β-catenin signaling. Nucleic Acids Res. 36, 462–476 (2008)
Acknowledgements
The project was supported in part by IDDRC grant number 1U54 HD083092 from the Eunice Kennedy Shriver National Institute of Child Health & Human Development. This project was supported by the Mouse Phenotyping Core at Baylor College of Medicine with funding from the National Institutes of Health (NIH) (U54 HG006348). The project was also supported by grants from the NIH (DE 023177 and HL 118761 to J.F.M.; DE 13941 to B.A.A.; HL-077439, HL-111665, HL-093039, DK-099653 and U01-HL-100401 to E.N.O.), and the Vivian L. Smith Foundation (J.F.M.). J.F.M. was supported by Transatlantic Network of Excellence Award LeDucq Foundation Transatlantic Networks of Excellence in Cardiovascular Research 14CVD01: “Defining the genomic topology of atrial fibrillation”. E.N.O. was supported by Fondation Leducq Networks of Excellence, Cancer Prevention and Research Institute of Texas and the Robert A. Welch Foundation (grant 1-0025). G.T. was supported by American Heart Association (AHA) (13POST17040027). P.C.K was supported by German Research Foundation (DFG) (KA4018/1-1).
Author information
Authors and Affiliations
Contributions
J.F.M. and G.T. conceived the project and designed the experiments. G.T., P.C.K., Y.M., M.R., T.R.H. and L.L. performed experiments and analysed data. Z.S. and B.A.A. provided reagents and performed experiments. E.N.O. provided the transgenic animal model. M.Z. and G.T. performed bioinformatics and statistical analyses. J.F.M. supervised the project and analysed data. G.T. and J.F.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
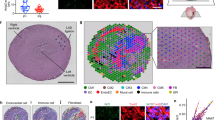
Extended Data Figure 1 Pitx2 is required in neonatal myocardial regeneration after LAD-O.
a, Serial trichrome images of control (Pitx2f/f), MCKcre;Pitx2f/f, and Mhccre-Ert;Pitx2f/f 21 days after LAD-O performed in P2 mice. Three representative hearts of each genotype were shown. b, Percentage of fibrotic left ventricular myocardium quantified at 3 weeks after LAD-O; n = 8 for control (Pitx2f/f), n = 7 for MCKcre;Pitx2f/f, and n = 4 for Mhccre-Ert;Pitx2f/f. c, d, Ejection fraction (c) and fractional shortening (d) of LAD-O and sham hearts (see Methods for n). L, left ventricle; R, right ventricle. Mean ± s.e.m. *P < 0.05 one-way ANOVA plus Bonferroni post-test (c, d) and Mann–Whitney test (b).
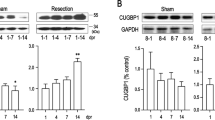
Extended Data Figure 2 Pitx2 promotes myocardial regeneration after apex resection at P8.
a, Schematic of Pitx2-expressing construct (Pitx2Gof). b–d, Pitx2Gof was crossed with the Mhccre-Ert strain to generate Mhccre-Ert/+;Pitx2Gof (Pitx2-overexpressing) mice. After tamoxifen treatment from P7–P10, qPCR (b, n = 4) and western blot (c, d, n = 3) show the overexpression of Pitx2 in the myocardium at P16. e, f, Trichrome-stained cross sections from 13-week-old sham hearts of control (e) and Pitx2-overexpressing (f) mice, with tamoxifen administrated at 7–8 weeks old. g, Heart weight over body weight ratio of adult sham and LAD-O hearts; n = 4 (control sham), 4 (control LAD-O), 5 (Pitx2-overexpressing sham), 9 (Pitx2-overexpressing LAD-O). h–j, Apex resection of Pitx2-overexpressing (i) and control (Mhccre-Ert/+) (h) hearts at P8 followed by trichrome staining at 28 DPR; the scar area was quantified in j; n = 10 (control mice), 7 (Pitx2-overexpressing mice). k, l, Echocardiography showed ejection fraction (k) and fractional shortening (l) at 28 DPR (see Methods for n). m–o, EdU labelling of Pitx2-overexpressing (n) and control (m) apical area, 8 days after P8 resection, sections were stained for cTnT (green), EdU (yellow), and DAPI (blue). Arrow indicates EdU-labelled cardiomyocytes, with quantification in o; n = 4 mice per group. Mean ± s.e.m. *P < 0.05, one-way ANOVA plus Bonferroni post-test (g, k, l) and Mann–Whitney test (b, d, j, o).
Extended Data Figure 3 Pitx2 is required for Hippo-deficient heart regeneration.
a, Schematic study plan for Fig. 3a–e. b–e, Trichrome-stained apical areas of control (b), Salv CKO (c) and double knockout (d) hearts 21 days after P8 apex resection. Scar area was quantified in e. f, Heart weight to body weight ratio of sham hearts at 28 days after tamoxifen administration. For n number, see Methods. Mean ± s.e.m. *P < 0.05, Mann–Whitney.
Extended Data Figure 4 Co-occurrence of Pitx2 and Tead DNA-binding motifs in fetal heart enhancers.
a, Consensus Pitx2 and Tead motifs. b, Pitx2 and Tead motif co-occurrence in fetal heart DHS peaks. c, Aggregate plot of H3K4me1 in fetal heart ChIP–seq reads within 6 kb range of DHS peaks. d, Heat map of fetal heart H3K4me1 ChIP–seq or input read density in 6-kb regions of DHS peaks. DHS peaks were centred on the Pitx2 motif, Tead motif, Pitx2-Tead motifs, or randomly selected. The read density was in log2 scale. Blue, negative values; yellow, positive values.
Extended Data Figure 5 Generation of GST-tagged proteins and interaction between Pitx2 and Yap in vivo.
a, The mouse Pitx2a, Pitx2c and truncated proteins were purified and run on a 10% SDS–PAGE gel, and Coomassie blue staining shows the GST fusion protein band with correct size (marked by asterisk). b, Coomassie blue staining of the purified GST–Yap, Yap cut by prescission protease and pure Yap protein. c, Co-immunoprecipitation of Flag in Pitx2flag ventricles at 5 DPR, and blotting of Yap, Pitx2 and Flag.
Extended Data Figure 6 Nuclei-shuttling of Nrf2 is independent of Pitx2.
a, Western blotting of Pitx2, α-tubulin, and TATA-binding protein (TBP) of P19 cell fraction after H2O2, with or without Nrf2 siRNA treatment. b, Immunofluorescent staining of Nrf2 (green) in P19 control and Pitx2 knockout cells after vehicle or H2O2 treatment. DAPI, blue. Scale bars, 50 μm. c, The ratio of cells with nuclear Nrf2 over total cell number; n = 6 biological repeats. d, Blotting of α-tubulin and TBP to show cell fraction of P19 cells used in e. e, Co-immunoprecipitation Nrf2 from nuclear and cytoplasmic fraction of P19 cells after vehicle or H2O2 treatment, blotting shows Nrf2 and Pitx2. f–h, 4 DPMI control (C57BL6) (f) and Nrf2nu/nu (g) cross-sections stained for Pitx2 (red), cTnT (green), and DAPI (blue), with the ratio of cardiomyocytes with nuclei-localized Pitx2 quantified in h; n = 4 mice per group. Arrows, Pitx2+ cardiomyocytes. Mean ± s.e.m. *P < 0.05, one-way ANOVA plus Bonferroni post-test (c) and Mann–Whitney test (h).
Extended Data Figure 7 Nrf2 is required for neonatal myocardial regeneration.
a, Trichrome images of Nrf2nu/nu and control heart (C57BL6) at 21 days after P2 LAD-O, along with sham controls. b, c, Ejection fraction (b) and fractional shortening (c) of LAD-O and sham hearts (see Methods for n). Mean ± s.e.m. *P < 0.05, Mann–Whitney test.
Extended Data Figure 8 Yap1 and Nrf2 are essential for Pitx2-induced myocardial regeneration.
a–d, Trichrome staining showing apical scarring of different groups at 28 DPR, apex resection was performed at P8. e, Quantification of scar area; n = 4 mice per group. Mean ± s.e.m. *P < 0.05, Mann–Whitney test, control compared to the other three groups individually.
Extended Data Figure 9 Pitx2 regulates antioxidant scavenger genes.
a, Overall change of genes in Pitx2 CKO mice compared to control. b, Upregulated genes in 5 DPR control over wild-type sham heart (n = 480) overlaid with downregulated genes in 5 DPR Pitx2 CKO over 5 DPR control heart (n = 1,002). c, GO analysis of genes upregulated (left) and downregulated (right) in Pitx2 CKO ventricles over controls at 5 DPR. d, GO analysis of genes upregulated (right) and downregulated (left) in 5 DPR control ventricles over age matching sham hearts. e, ChIP–qPCR confirming the binding of Pitx2 to the regulatory regions of target genes; n = 4 biological replicates. f, qPCR detecting Pitx2 and antioxidant genes in wild-type and Pitx2nu/nu ES cells after vehicle or H2O2 treatment; n = 4 biological replicates. g, qPCR of antioxidant genes in P19 cells after doxorubicin or H2O2 treatment; n = 5 biological replicates. h, qPCR of Pitx2 in P19 cells after doxorubicin or H2O2 treatment; n = 5 biological replicates. Mean ± s.e.m. *P < 0.05; ***P < 0.001, Mann–Whitney test.
Extended Data Figure 10 Mechanism model of Pitx2, Nrf2 and Yap1 responding to oxidative stress.
When oxidative stress is low, Nrf2 is sequestered in cytoplasm by its degradation complex (Cul3, Keap1), and Pitx2 stays either in the cytoplasm or at low expression levels. When the redox balance is disturbed by ROS, Nrf2 breaks away from the degradation complex, and enters nuclei to upregulate Pitx2 gene expression; Nrf2 also binds cytoplasmic Pitx2 and shuttles it to the nuclei, where Pitx2 and Yap co-regulate their common targets including critical antioxidant genes. In wild-type adult mouse heart, active Yap is maintained at a low level, even after ischaemic injury, and is thus not able to repair myocardium efficiently. When Pitx2 is overexpressed in cardiomyocytes, sufficient amounts of Pitx2 will cooperate with low levels of resident active Yap to induce the expression of beneficial antioxidant scavengers in a synergetic pattern, rendering protection to injured myocardium. Red arrow, supported by in vitro evidence; Blue arrows, supported by in vivo evidence.
Supplementary information
Supplementary Figure
This file contains the uncropped scans with size marker indications. (PDF 1104 kb)
Rights and permissions
About this article
Cite this article
Tao, G., Kahr, P., Morikawa, Y. et al. Pitx2 promotes heart repair by activating the antioxidant response after cardiac injury. Nature 534, 119–123 (2016). https://doi.org/10.1038/nature17959
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17959
This article is cited by
-
A modified apical resection model with high accuracy and reproducibility in neonatal mouse and rat hearts
npj Regenerative Medicine (2023)
-
Remdesivir increases mtDNA copy number causing mild alterations to oxidative phosphorylation
Scientific Reports (2023)
-
TMEM11 regulates cardiomyocyte proliferation and cardiac repair via METTL1-mediated m7G methylation of ATF5 mRNA
Cell Death & Differentiation (2023)
-
Inhibition of fatty acid oxidation enables heart regeneration in adult mice
Nature (2023)
-
Redifferentiated cardiomyocytes retain residual dedifferentiation signatures and are protected against ischemic injury
Nature Cardiovascular Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.