Abstract
Ribosome biogenesis is a highly complex process in eukaryotes, involving temporally and spatially regulated ribosomal protein (r-protein) binding and ribosomal RNA remodelling events in the nucleolus, nucleoplasm and cytoplasm1,2. Hundreds of assembly factors, organized into sequential functional groups3,4, facilitate and guide the maturation process into productive assembly branches in and across different cellular compartments. However, the precise mechanisms by which these assembly factors function are largely unknown. Here we use cryo-electron microscopy to characterize the structures of yeast nucleoplasmic pre-60S particles affinity-purified using the epitope-tagged assembly factor Nog2. Our data pinpoint the locations and determine the structures of over 20 assembly factors, which are enriched in two areas: an arc region extending from the central protuberance to the polypeptide tunnel exit, and the domain including the internal transcribed spacer 2 (ITS2) that separates 5.8S and 25S ribosomal RNAs. In particular, two regulatory GTPases, Nog2 and Nog1, act as hub proteins to interact with multiple, distant assembly factors and functional ribosomal RNA elements, manifesting their critical roles in structural remodelling checkpoints and nuclear export. Moreover, our snapshots of compositionally and structurally different pre-60S intermediates provide essential mechanistic details for three major remodelling events before nuclear export: rotation of the 5S ribonucleoprotein, construction of the active centre and ITS2 removal. The rich structural information in our structures provides a framework to dissect molecular roles of diverse assembly factors in eukaryotic ribosome assembly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The cryo-EM density maps of state 1 and state 2 have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-6615 and EMD-6616, respectively. The atomic model of state 1 has been deposited in the Protein Data Bank (PDB) under accession number 3JCT. The XL–MS data have been deposited to the ProteomeXchange Consortium (http://www.proteomexchange.org/) with the dataset identifier PXD003736.
References
Woolford, J. L., Jr & Baserga, S. J. Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195, 643–681 (2013)
Panse, V. G. & Johnson, A. W. Maturation of eukaryotic ribosomes: acquisition of functionality. Trends Biochem. Sci. 35, 260–266 (2010)
Lebreton, A. et al. 60S ribosomal subunit assembly dynamics defined by semi-quantitative mass spectrometry of purified complexes. Nucleic Acids Res. 36, 4988–4999 (2008)
McCann, K. L., Charette, J. M., Vincent, N. G. & Baserga, S. J. A protein interaction map of the LSU processome. Genes Dev. 29, 862–875 (2015)
Saveanu, C. et al. Nog2p, a putative GTPase associated with pre-60S subunits and required for late 60S maturation steps. EMBO J. 20, 6475–6484 (2001)
Leidig, C. et al. 60S ribosome biogenesis requires rotation of the 5S ribonucleoprotein particle. Nature Commun. 5, 3491 (2014)
Saveanu, C. et al. Sequential protein association with nascent 60S ribosomal particles. Mol. Cell. Biol. 23, 4449–4460 (2003)
Matsuo, Y. et al. Coupled GTPase and remodelling ATPase activities form a checkpoint for ribosome export. Nature 505, 112–116 (2014)
Sengupta, J. et al. Characterization of the nuclear export adaptor protein Nmd3 in association with the 60S ribosomal subunit. J. Cell Biol. 189, 1079–1086 (2010)
Greber, B. J., Boehringer, D., Montellese, C. & Ban, N. Cryo-EM structures of Arx1 and maturation factors Rei1 and Jjj1 bound to the 60S ribosomal subunit. Nature Struct. Mol. Biol. 19, 1228–1233 (2012)
Bradatsch, B. et al. Structure of the pre-60S ribosomal subunit with nuclear export factor Arx1 bound at the exit tunnel. Nature Struct. Mol. Biol. 19, 1234–1241 (2012)
Kharde, S., Calviño, F. R., Gumiero, A., Wild, K. & Sinning, I. The structure of Rpf2–Rrs1 explains its role in ribosome biogenesis. Nucleic Acids Res. 43, 7083–7095 (2015)
Madru, C. et al. Chaperoning 5S RNA assembly. Genes Dev. 29, 1432–1446 (2015)
Barrio-Garcia, C. et al. Architecture of the Rix1–Rea1 checkpoint machinery during pre-60S-ribosome remodeling. Nature Struct. Mol. Biol. 23, 37–44 (2016)
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011)
Greber, B. J. et al. Insertion of the biogenesis factor Rei1 probes the ribosomal tunnel during 60S maturation. Cell 164, 91–102 (2016)
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011)
Talkish, J., Zhang, J., Jakovljevic, J., Horsey, E. W. & Woolford, J. L. Jr. Hierarchical recruitment into nascent ribosomes of assembly factors required for 27SB pre-rRNA processing in Saccharomyces cerevisiae. Nucleic Acids Res. 40, 8646–8661 (2012)
Dembowski, J. A., Kuo, B. & Woolford, J. L., Jr. Has1 regulates consecutive maturation and processing steps for assembly of 60S ribosomal subunits. Nucleic Acids Res. 41, 7889–7904 (2013)
Feng, B. et al. Structural and functional insights into the mode of action of a universally conserved Obg GTPase. PLoS Biol. 12, e1001866 (2014)
Pertschy, B. et al. Cytoplasmic recycling of 60S preribosomal factors depends on the AAA protein Drg1. Mol. Cell. Biol. 27, 6581–6592 (2007)
Kappel, L. et al. Rlp24 activates the AAA-ATPase Drg1 to initiate cytoplasmic pre-60S maturation. J. Cell Biol. 199, 771–782 (2012)
Granneman, S., Petfalski, E. & Tollervey, D. A cluster of ribosome synthesis factors regulate pre-rRNA folding and 5.8S rRNA maturation by the Rat1 exonuclease. EMBO J. 30, 4006–4019 (2011)
Granato, D. C., Machado-Santelli, G. M. & Oliveira, C. C. Nop53p interacts with 5.8S rRNA co-transcriptionally, and regulates processing of pre-rRNA by the exosome. FEBS J. 275, 4164–4178 (2008)
Thoms, M. et al. The exosome is recruited to RNA substrates through specific adaptor proteins. Cell 162, 1029–1038 (2015)
Jakovljevic, J. et al. Ribosomal proteins L7 and L8 function in concert with six A3 assembly factors to propagate assembly of domains I and II of 25S rRNA in yeast 60S ribosomal subunits. RNA 18, 1805–1822 (2012)
Mitchell, P., Petfalski, E. & Tollervey, D. The 3′ end of yeast 5.8S rRNA is generated by an exonuclease processing mechanism. Genes Dev. 10, 502–513 (1996)
Thomson, E. & Tollervey, D. The final step in 5.8S rRNA processing is cytoplasmic in Saccharomyces cerevisiae. Mol. Cell. Biol. 30, 976–984 (2010)
Zhang, J. et al. Assembly factors Rpf2 and Rrs1 recruit 5S rRNA and ribosomal proteins rpL5 and rpL11 into nascent ribosomes. Genes Dev. 21, 2580–2592 (2007)
Ulbrich, C. et al. Mechanochemical removal of ribosome biogenesis factors from nascent 60S ribosomal subunits. Cell 138, 911–922 (2009)
Sahasranaman, A. et al. Assembly of Saccharomyces cerevisiae 60S ribosomal subunits: role of factors required for 27S pre-rRNA processing. EMBO J. 30, 4020–4032 (2011)
Xu, T. et al. ProLuCID, a fast and sensitive tandem mass spectra-based protein identification program. Mol. Cell. Proteomics 5, S174 (2006)
Tabb, D. L., McDonald, W. H. & Yates, J. R., III . DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J. Proteome Res. 1, 21–26 (2002)
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nature Methods 9, 904–906 (2012)
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Shaikh, T. R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nature Protocols 3, 1941–1974 (2008)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011)
De Rijk, P., Wuyts, J. & De Wachter, R. RnaViz 2: an improved representation of RNA secondary structure. Bioinformatics 19, 299–300 (2003)
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED Protein Analysis Workbench. Nucleic Acids Res. 41, W349–W357 (2013)
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nature Methods 12, 7–8 (2015)
Schrodinger, LLC. The PyMOL molecular graphics system, version 1.3r1 (2010)
Stein, N. CHAINSAW: a program for mutating pdb files used as templates in molecular replacement. J. Appl. Cryst. 41, 641–643 (2008)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Asano, N. et al. Structural and functional analysis of the Rpf2–Rrs1 complex in ribosome biogenesis. Nucleic Acids Res. 43, 4746–4757 (2015)
Groft, C. M., Beckmann, R., Sali, A. & Burley, S. K. Crystal structures of ribosome anti-association factor IF6. Nature Struct. Biol. 7, 1156–1164 (2000)
Baßler, J. et al. A network of assembly factors is involved in remodeling rRNA elements during preribosome maturation. J. Cell Biol. 207, 481–498 (2014)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
Fernández, I. S., Bai, X. C., Murshudov, G., Scheres, S. H. & Ramakrishnan, V. Initiation of translation by cricket paralysis virus IRES requires its translocation in the ribosome. Cell 157, 823–831 (2014)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Acknowledgements
We thank the National Center for Protein Sciences (Beijing, China) for providing resource for cryo-EM data collection and computation. We also thank members of Woolford laboratory for reading the manuscript. This work was supported by the Ministry of Science and Technology of China (2013CB910404 to N.G. and 2014CB849800 to M.-Q.D.), the National Natural Science Foundation of China (31422016 and 31470722 to N.G., and 21375010 to M.-Q.D.) and National Institutes of Health grant R01GM028301 (to J.L.W.).
Author information
Authors and Affiliations
Contributions
N.G. and J.L.W. designed and directed experiments; B.K., H.B., M.G. and J.J. purified samples; D.T. and M.-Q.D. performed XL–MS; S.W. collected cryo-EM data (with J.L., Y.Y., Z.L., and C.M.), performed image processing (with Y.Z.), and analysed structures (with Y.K.). N.G., S.W. and K.Y. performed structural modelling. S.W., J.L.W. and N.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Compositional analysis of Nsa1, Nog2 and Nmd3 particles.
a, Mostly non-overlapping assembly factors Nsa1, Nog2 and Nmd3 were used to purify sequential ribosome assembly intermediates. Proteins identified by mass spectrometry analysis were marked on the gel. Orange coloured proteins are only present in Nsa1-TAP particles, green coloured proteins are present both in Nsa1-TAP and in Nog2-TAP particles, light blue coloured proteins are present in all three purified particles to varying levels, dark blue coloured proteins are present only in Nog2-particles, pink coloured proteins are present both in Nog2- and Nmd3-particles in varying levels and yellow coloured proteins are present only in Nmd3-particles. TAP-tagged proteins are indicated by white asterisks. For gel source data, see Supplementary Fig. 1. b, The lifetimes of mostly non-overlapping ribosome assembly intermediates containing assembly factors Nsa1, Nog2 and Nmd3 are indicated. Assembly factors identified in each of Nsa1-TAP, Nog2-TAP and Nmd3-TAP associated samples were colour coded. The colour scheme is identical to that used in a. *Even though this protein was identified in all three intermediates, its levels decreased more than sevenfold from Nsa1-TAP particles to Nog2-TAP particles.
Extended Data Figure 2 Cryo-EM data processing of Nog2-particles.
a, Representative 2D class averages of Nog2-particles. b, A flow-chart for 3D classification of Nog2-particles (data batch 8–10, see Methods for details).
Extended Data Figure 3 Resolution estimation and model validation.
a, Representative micrograph of Nog2-particles. b, Local resolution map of the final density map of state 1. c, FSC curve for the final density map (state 1). The nominal resolution is 3.08 Å estimated using the gold-standard (FSC = 0.143) criterion. d, Atomic model cross-validation. Three FSC curves were calculated between the refined model (against Half1 map) and the final map (black), between the refined model with Half1 map (FSCwork, red), and between the refined model with Half2 map (FSCtest, blue) (see Methods for details).
Extended Data Figure 4 Local densities of representative regions for different assembly factors.
a–l, Cryo-EM densities of representative regions of assembly factors, superimposed with respective atomic models.
Extended Data Figure 5 Interaction network of Nog2 in the pre-60S particle.
a–g, Pairwise illustration of binding partners of Nog2 in the pre-60S particle. Residues of Nog2 involved in atomic contacts are coloured red with residue numbers labelled. H and L denote helix and loop, respectively. h, Interactions between rRNA components (H43, H68, H74, H75, H86, H92, H93) and Nog2. For clarification, H69 and H71 are not shown. The N terminus of Nog2 is located in a helical junction composed of H68, H74, H75 and H93.
Extended Data Figure 6 The NTD of Nog1 interacts with Nsa2 and Nog2.
a, Nsa2, Nog2 and Nog1 collectively stabilize H89 in a distinct conformation. Nog1 interacts with Nog2 and Nsa2 through its GTPase domain and NTD, respectively. b, The CTE of Nog1 interlocks with Rlp24 by wrapping around a long helix at the C-terminal end of Rlp24 (see also Fig. 3). c, d, Comparison of the CTE of Nog1 and the CTE of Rei1 in the polypeptide tunnel. Atomic models of state 1 (c) and 60S-Arx1–Alb1–Rei1 (d) (PDB accession number 5APN)16 are aligned using the 60S subunit. For clarification, only Arx1, Nog1 and Rei1 are shown. e, Superimposition of c and d. Four major places of steric clash between Rei1 and Nog1 are marked by asterisks.
Extended Data Figure 7 Mutual interactions between factors and r-proteins in the ITS2 subcomplex.
a, An overall view of the ITS2 subcomplex. b–d, L8 interacts with three factors: Nop15 (b), Cic1 (c) and Nop7 (d). e, L27 interacts with Nop53. f–h, L25 interacts with Rlp7 (f), Nop15 (g) and Nop53 (h). Residues involved in atomic interaction sites are labelled with sequence numbers. H, L, S denote helix, loop and strand of respective structures.
Extended Data Figure 8 Restructuring of rRNA helices in the central protuberance region by Nsa2, Rpf2, Rsa4, Rrs1 and Nog2.
a, Conformation of rRNA helices from the central protuberance (H80, H82-H88, 5S rRNA) in the pre-60S particle (state 1). b, Same as a, but for the mature 60S subunit. The mature 60S subunit was aligned to state 1 structure globally. c–g, Pairwise interactions between the central protuberance helices and factors are shown in separate panels.
Extended Data Figure 9 Structures of different assembly states of the pre-60S ribosomal particles.
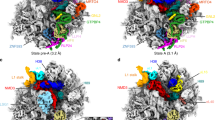
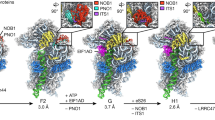
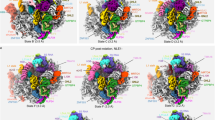
a, Cryo-EM density maps of three premature states (1–3) and the mature state are displayed in transparent surface representation, superimposed with models of the 5S RNA, H38 and associated central-protuberance-binding factors. b, Zoom-in views of the central protuberance regions in a. For clarification, only atomic models are shown. Comparison of these four states indicates that the 5S RNP rotates to a near-mature state (state 2) after Rpf2–Rrs1 leave, and further release of Rsa4 in state 3 results in a ‘mature-like’ conformation for the 5S RNP. H38 from these four states is in a series of continuous changes coupled with the 5S RNP conformational maturation. c, d, Spatial relationship of the 5S RNP, H38, Rsa4 and Cgr1 in state 1 (c) and state 2 (d). Note that repositioning of H38 from state 1 to state 2 is coupled with a dramatic conformational change on the C-terminal end of Cgr1. e–h, Additional assembly factors identified in the density map of state 2. One piece of additional density between H38 and L1 contains a characteristic HEAT repeat, which contacts the L1 stalk in an inward position (e). The atomic model of Sda1 (PDB accession number 5FL8)14 fits well with the segmented density (f). For clarification, densities immediately above Sda1 are not shown in e and f. A large piece of additional density in the map of state 2, composed of the Rix1 subcomplex and Rea1 (g, h). The density assignment was facilitated by the cryo-EM structure of Rix1–Rea1 particles14. Superimposition of the atomic model of Rea1 (PDB accession number 5FL8)14 with the segmented density map of Rea1 (h).
Supplementary information
Supplementary Figure 1
This file contains the original gel image for Extended Data Fig. 1a. (PDF 171 kb)
Supplementary Table 1
This file shows cross-linked peptides identified in CL-MS data. (XLSX 36 kb)
Supplementary Table 2
This file shows the analysis results of CL-MS data. (XLS 1144 kb)
Overall structure of Nog2-particles in State 1
The cryo-EM density (sharpened) is first shown in surface representative, with densities of factors individually colored (as in Fig. 1). The rRNA and r-proteins are colored grey and beige, respectively. Groups of factors are then highlighted in zoom-in views. (MP4 29901 kb)
Rights and permissions
About this article
Cite this article
Wu, S., Tutuncuoglu, B., Yan, K. et al. Diverse roles of assembly factors revealed by structures of late nuclear pre-60S ribosomes. Nature 534, 133–137 (2016). https://doi.org/10.1038/nature17942
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17942
This article is cited by
-
A molecular network of conserved factors keeps ribosomes dormant in the egg
Nature (2023)
-
Structure of nascent 5S RNPs at the crossroad between ribosome assembly and MDM2–p53 pathways
Nature Structural & Molecular Biology (2023)
-
Mitochondrial complexome reveals quality-control pathways of protein import
Nature (2023)
-
A single 2′-O-methylation of ribosomal RNA gates assembly of a functional ribosome
Nature Structural & Molecular Biology (2023)
-
Proteomics revealed an association between ribosome-associated proteins and amyloid beta deposition in Alzheimer's disease
Metabolic Brain Disease (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.