Abstract
Genome-wide association studies (GWAS) have identified numerous genetic variants associated with complex diseases, but mechanistic insights are impeded by a lack of understanding of how specific risk variants functionally contribute to the underlying pathogenesis1. It has been proposed that cis-acting effects of non-coding risk variants on gene expression are a major factor for phenotypic variation of complex traits and disease susceptibility. Recent genome-scale epigenetic studies have highlighted the enrichment of GWAS-identified variants in regulatory DNA elements of disease-relevant cell types2,3,4,5,6. Furthermore, single nucleotide polymorphism (SNP)-specific changes in transcription factor binding are correlated with heritable alterations in chromatin state and considered a major mediator of sequence-dependent regulation of gene expression7,8,9,10. Here we describe a novel strategy to functionally dissect the cis-acting effect of genetic risk variants in regulatory elements on gene expression by combining genome-wide epigenetic information with clustered regularly-interspaced short palindromic repeats (CRISPR)/Cas9 genome editing in human pluripotent stem cells. By generating a genetically precisely controlled experimental system, we identify a common Parkinson’s disease associated risk variant in a non-coding distal enhancer element that regulates the expression of α-synuclein (SNCA), a key gene implicated in the pathogenesis of Parkinson’s disease. Our data suggest that the transcriptional deregulation of SNCA is associated with sequence-dependent binding of the brain-specific transcription factors EMX2 and NKX6-1. This work establishes an experimental paradigm to functionally connect genetic variation with disease-relevant phenotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McClellan, J. & King, M.-C. Genetic heterogeneity in human disease. Cell 141, 210–217 (2010)
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011)
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012)
Trynka, G. et al. Chromatin marks identify critical cell types for fine mapping complex trait variants. Nature Genet. 45, 124–130 (2013)
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013)
Degner, J. F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012)
Kilpinen, H. et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science (2013)
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science (2013)
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science (2013)
Leung, D. et al. Integrative analysis of haplotype-resolved epigenomes across human tissues. Nature 518, 350–354 (2015)
Singleton, A. B., Farrer, M. J. & Bonifati, V. The genetics of Parkinson’s disease: progress and therapeutic implications. Mov. Disord. 28, 14–23 (2013)
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nature Genet. 46, 989–993 (2014)
Devine, M. J., Gwinn, K., Singleton, A. & Hardy, J. Parkinson’s disease and α-synuclein expression. Mov. Disord. 26, 2160–2168 (2011)
Miller, D. W., Hague, S. M., Clarimon, J. & Baptista, M. α-synuclein in blood and brain from familial Parkinson disease with SNCA locus triplication. Neurology 62, 1835–1838 (2004)
Kim, H. J., Jeon, B. S., Yoon, M. Y. & Park, S. S. Increased expression of alpha-synuclein by SNCA duplication is associated with resistance to toxic stimuli. J. Mol. Neurosci. 47, 249–255 (2012)
Soldner, F. et al. Parkinson’s disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell 136, 964–977 (2009)
Vermunt, M. W. et al. Large-scale identification of coregulated enhancer networks in the adult human brain. Cell Rep. 9, 767–779 (2014)
Ward, L. D. & Kellis, M. Interpreting noncoding genetic variation in complex traits and human disease. Nature Biotechnol. 30, 1095–1106 (2012)
Rivera, C. M. & Ren, B. Mapping human epigenomes. Cell 155, 39–55 (2013)
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010)
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011)
Mariani, J. et al. Emx2 is a dose-dependent negative regulator of Sox2 telencephalic enhancers. Nucleic Acids Res. 40, 6461–6476 (2012)
Ligon, K. L. Loss of Emx2 function leads to ectopic expression of Wnt1 in the developing telencephalon and cortical dysplasia. Development 130, 2275–2287 (2003)
Schaffer, A. E., Freude, K. K., Nelson, S. B. & Sander, M. Nkx6 transcription factors and Ptf1a function as antagonistic lineage determinants in multipotent pancreatic progenitors. Dev. Cell 18, 1022–1029 (2010)
Farrer, M. et al. α-synuclein gene haplotypes are associated with Parkinson's disease. Hum. Mol. Genet. 10, 1847–1851 (2001)
Cronin, K. D. et al. Expansion of the Parkinson disease-associated SNCA-Rep1 allele upregulates human α-synuclein in transgenic mouse brain. Hum. Mol. Genet. 18, 3274–3285 (2009)
Chiba-Falek, O., Kowalak, J. A., Smulson, M. E. & Nussbaum, R. L. Regulation of alpha-synuclein expression by poly (ADP ribose) polymerase-1 (PARP-1) binding to the NACP-Rep1 polymorphic site upstream of the SNCA gene. Am. J. Hum. Genet. 76, 478–492 (2005)
Soldner, F. & Jaenisch, R. Medicine. iPSC disease modeling. Science 338, 1155–1156 (2012)
Soldner, F. et al. Generation of isogenic pluripotent stem cells differing exclusively at two early onset Parkinson point mutations. Cell 146, 318–331 (2011)
Lengner, C. J. et al. Derivation of pre-X inactivation human embryonic stem cells under physiological oxygen concentrations. Cell 141, 872–883 (2010)
Kim, J. et al. Direct reprogramming of mouse fibroblasts to neural progenitors. Proc. Natl Acad. Sci. USA 108, 7838–7843 (2011)
Kim, J.-H., Panchision, D., Kittappa, R. & McKay, R. Generating CNS neurons from embryonic, fetal, and adult stem cells. Methods Enzymol. 365, 303–327 (2003)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013)
Hockemeyer, D. et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nature Biotechnol. 27, 851–857 (2009)
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 29, e45 (2001)
Lee, T. I., Johnstone, S. E. & Young, R. A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nature Protocols 1, 729–748 (2006)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Sulzer, D. & Surmeier, D. J. Neuronal vulnerability, pathogenesis, and Parkinson’s disease. Mov. Disord. 28, 41–50 (2013)
Ferrer, I., Martinez, A., Blanco, R., Dalfó, E. & Carmona, M. Neuropathology of sporadic Parkinson disease before the appearance of parkinsonism: preclinical Parkinson disease. J. Neural Transm. 118, 821–839 (2011)
Irwin, D. J. et al. Neuropathologic substrates of Parkinson disease dementia. Ann. Neurol. 72, 587–598 (2012)
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008)
Lill, C. M. et al. Comprehensive research synopsis and systematic meta-analyses in Parkinson’s disease genetics: the PDGene database. PLoS Genet. 8, e1002548 (2012)
Quinlan, A. R. BEDTools: the Swiss-Army tool for genome feature analysis. Curr Protoc. Bioinformat. 47, 11.12.1–11.12.34 (2014)
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G. R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nature Genet. 44, 955–959 (2012)
Fuchsberger, C., Abecasis, G. R. & Hinds, D. A. minimac2: faster genotype imputation. Bioinformatics 31, 782–784 (2015)
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010)
Dumitriu, A. et al. Cyclin-G-associated kinase modifies α-synuclein expression levels and toxicity in Parkinson’s disease: results from the GenePD study. Hum. Mol. Genet. 20, 1478–1487 (2011)
Pankratz, N. et al. Meta-analysis of Parkinson’s disease: identification of a novel locus, RIT2. Ann. Neurol. 71, 370–384 (2012)
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Brambrink, T. et al. Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell 2, 151–159 (2008)
DeKelver, R. C. et al. Functional genomics, proteomics, and regulatory DNA analysis in isogenic settings using zinc finger nuclease-driven transgenesis into a safe harbor locus in the human genome. Genome Res. 20, 1133–1142 (2010)
Raj, A., van den Bogaard, P., Rifkin, S. A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nature Methods 5, 877–879 (2008)
Raj, A., Rifkin, S. A., Andersen, E. & van Oudenaarden, A. Variability in gene expression underlies incomplete penetrance. Nature 463, 913–918 (2010)
Faddah, D. A. et al. Brief Report. Cell Stem Cell 13, 23–29 (2013)
Acknowledgements
We acknowledge the investigators of the original Parkinson’s disease GWAS meta-analysis, PDGene and Menno Creyghton at the Hubrecht Institute for sharing data used for this study. We would like to thank B. Zaidi for helpful discussion in conceiving this work. We thank R. Alagappan, D. Fu and T. Lungjangwa for technical support and help in human ES cell culture and molecular biology and A. D’Alessio for helpful advice for performing ChIP-seq experiments. We would like to thank P. Wisniewski, M. Ly, C. Zollo and C. Araneo of the Whitehead Institute FACS facility for their help with cell sorting, T. Volkert, J. Love and S. Gupta at the Whitehead Genome Technologies Core for Solexa sequencing, W. Salmon and N. Watson from the W. M. Keck Biological Imaging Facility and T. DiCesare for help with the figure illustrations. We thank all the members of the Jaenisch laboratory for discussions and comments on the manuscript. R.J. was supported by NIH grants 1R01NS088538-01 and 2R01MH104610-15 and by Qatar National Research Fund grant number NPRP 5-531-1-094.
Author information
Authors and Affiliations
Contributions
F.S. and R.J. conceived the project, designed and supervised the experiments, interpreted results and wrote the paper with input from all authors. Y.S. assisted with ChIP experiments and contributed to the preparation of the manuscript. C.S.S. assisted with CRISPR/Cas9 genome editing. B.J.A. performed computational analysis of epigenetic datasets and prioritization of Parkinson’s disease associated SNPs. J.C.L. performed conditional GWAS and eQTL analysis. M.I.B. performed computational analysis to predict TF binding. J.G. performed single-molecule mRNA FISH analysis. R.H.M. and R.A.Y. supervised data analysis, interpreted results and contributed to the manuscript preparation. F.S. performed all other experiments.
Corresponding author
Ethics declarations
Competing interests
R.J. is an adviser to Stemgent and a cofounder of Fate Therapeutics.
Extended data figures and tables
Extended Data Figure 1 Analysis of allele-specific expression of SNCA using qRT–PCR.
a, Schematic illustration of the quantitative allele-specific SNCA expression analysis using a common primer pair and allele-specific Taqman probes conjugated with different fluorophores to specifically detect a reporter-SNP (rs356165) in the 3′ UTR of SNCA in a multiplex reaction. As indicated, 6-carboxyfluorescein (FAM) and 4,7,2′-trichloro-7′-phenyl-6-carboxyfluorescein (VIC) were used to detect the A and G allele respectively. b, Representative multiplex qRT–PCR reactions (in duplicates) measuring allele-specific SNCA expression of allele-biased samples described in Fig. 1c, d. Allele-biased samples were generated by mixing human iPS-cell-derived neurons homozygous for either the A (IPS-A) or G allele (IPS-G) at rs356165 at indicated ratios. Also included is a plot showing cDNAs synthesized in the absence of SuperScript reverse transcriptase (no RT) to control for genomic DNA contaminations. Plots are displayed as reporter dye fluorescence signal (ΔRn) in log scale as a function of run cycle.
Extended Data Figure 2 Analysis of in vitro differentiated human ES-cell-derived mixed neuronal cultures.
a–d, Immunostainings of in vitro differentiated mixed neuronal cultures (differentiation day 28) for expression of neuron- and astrocyte-specific markers. Shown are representative images for staining of neuron-specific beta-III-tubulin (TUJ1) and astrocyte-specific glial fibrillary acidic protein (GFAP) (a), neuron-specific microtubule associated protein 2 (MAP2) and glutamatergic neuron-specific glutamate vesicular transporter 1 (vGLUT1) (b), TUJ1 and dopaminergic neuron-specific tyrosine-hydroxylase (TH) (c) and the pan-neuronal marker NeuN, which was used for quantification (d). e, Quantification of a representative in vitro differentiation experiment in human ES cell line WIBR3 and BGO1. The quantification of rare TH-positive cells was estimated. Source Data for this figure are available online.
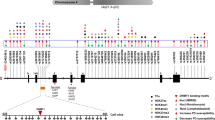
Extended Data Figure 3 Identification of Parkinson’s disease associated risk variants overlapping with distal enhancers in the SNCA locus.
a, Detailed H3K4me1 and H3K27ac ChIP-seq and DHSs-enrichment tracks for indicated brain regions in the SNCA locus. Shown are the locations of SNCA-Rep1 and Parkinson’s disease associated SNPs overlapping with two proximal enhancer elements (3′ UTR enhancer and intron-4 enhancer) highlighted by light grey boxes. On the right, enlarged view of 3′ UTR enhancer and intron-4 enhancer relative to top ranked Parkinson’s disease associated SNPs. b, Enlarged view of the intron-4 enhancer region showing H3K4me1 and H3K27ac ChIP-seq enrichment tracks for substantia nigra and DHSs enrichment tracks for fetal brain relative to location of PD-associated SNPs. Shown below is the number of predicted TF binding sites for reference (in red) and alternative SNP (minor allele) sequence (in blue) at each genomic position. Grey box indicates location of deletion described in Fig. 2b and Extended Data Fig. 4a. c, Summary of all Parkinson’s disease associated SNPs in the SNCA locus ranked by cumulative overlap with H3K4me1, H3K27ac and DHSs enhancer marks. Table summarizes the top 7 ranked SNPs with Parkinson’s disease association P values, odd ratios (OR), number of predicted differential TF binding sites (Diff TFB) and location within enhancer elements as marked in a. d, Gene tracks showing H3K4me1 and H3K27ac enrichment in the SNCA locus for in vitro human ES-cell-derived neurons (differentiation day 31). hESC, human embryonic stem cell.
Extended Data Figure 4 CRISPR/Cas9-mediated genome editing strategy for targeted insertion of Parkinson’s disease associated intron-4 enhancer elements in human ES cells.
a, Schematic illustration of the CRISPR/Cas9-mediated two-step genome editing strategy to delete and subsequently insert indicated intron-4 enhancer sequences containing the Parkinson’s disease associated risk SNPs rs356168 and rs3756054. Shown are the genomic organization of the SNCA locus, an enlarged view of wild-type and deleted (ΔE4) alleles, gRNA targeting sequences (underlined, PAM sequence in red), restriction sites, Southern blot (SB) probe and design of targeting vectors (TV; risk SNPs are highlighted in red). b, Representative Southern blot analysis of indicated targeted WIBR3 human ES cells (ΔE4/TV-A-T) compared with wild-type cells or human ES cells carrying homozygous deletions (ΔE4/ΔE4). c, Table summarizing intron-4 enhancer deletions and insertions of indicated haplotypes in WIBR3 human ES cells. Correct targeting was determined by Southern blot analysis and genomic sequencing. Cell lines with targeted enhancer elements confirmed to be in cis with A (FAM) reporter SNP (determined by genomic sequencing-based phase-reconstruction) were maintained for subsequent analysis (compare Fig. 2b).
Extended Data Figure 5 Effect of intron-4 enhancer modification on total and allele-specific expression of SNCA.
a, b, Analysis of total SNCA expression in in vitro derived neural precursors (a) (same samples analysed in Fig. 2c) and mixed neuronal cultures (b) (same samples analysed in Fig. 2d) from targeted cell lines with indicated SNP genotypes at rs356168 and rs3756054 (compare Fig. 2b) compared to human ES cells harbouring homozygous deletions of the intron-4 enhancer (ΔE4/ΔE4). qRT–PCR data are normalized to GAPDH and presented relative to the expression of ΔE4/ΔE4 cells. c, Expression analysis for the neuron-specific marker microtubule associated protein 2 (MAP2) and astrocyte specific glial fibrillary acidic protein (GFAP) by qRT–PCR in in vitro differentiated neurons described in b. Data are normalized to 60S acidic ribosomal protein P0 (RPLP0) and presented relative to the expression of ΔE4/ΔE4 neurons. Data are shown as mean ± s.d. of 3 biological replicates (each representing 3 technical replicates) for independent targeted clones as described in Fig. 2c, d (n indicates number of independently targeted clones per genotype; ΔE4/ΔE4, n = 4; A–T/ΔE4, n = 4; G–C/ΔE4, n = 3; A–C/ΔE4, n = 3; G–T/ΔE4, n = 2/3). d, Alternative presentation of data displayed in Fig. 2c as dot blot grouped according to SNPs rs356168 and rs3756054 (excluding data for cells carrying homozygous deletions (ΔE4/ΔE4) of the intron 4 enhancer). e, Table summarizing the results of two-way ANOVA analysis corresponding to d for allele-specific SNCA expression in neural precursors. f, Alternative presentation of data displayed in Fig. 2d as dot blot grouped according to SNPs rs356168 and rs3756054 (excluding data for cells carrying homozygous deletions (ΔE4/ΔE4) of the intron-4 enhancer). Each dot represents mean of 3 technical replicates, black bars indicate mean for each genotype (d, f). g, Table summarizing the results of two-way ANOVA analysis corresponding to f for allele-specific SNCA expression in mixed neuronal cultures. Allele-specific expression for each clone was analysed in 3 independent biological replicate experiments and combined according to genotypes, n indicates number of independently targeted clones per group, † indicates an additional sub-clone derived from one of the two targeted clones for this genotype. Two-way ANOVA analysis (alpha = 0.05) was calculated based on allele-specific expression of all biological replicates. *P < 0.0001. h, Schematic illustration of the experimental strategy for CRISPR/Cas9-mediated deletion of the intron-4 enhancer element. Genomic sequencing-based phase-reconstruction, to analyse the phase of the heterozygous enhancer SNP rs356168 in wild-type WIBR3 cells indicates that the functional G allele at rs356168 is in cis with the G allele of the reporter SNP rs356165. i, Relative allele-specific SNCA expression (boxplots showing median, 25th and 75th percentiles with whiskers indicating minimum and maximum) in neural precursors comparing wild-type cells to ΔE4/ΔE4 cells harbouring homozygous deletions (expression is calculated relative to ΔE4/ΔE4 neural precursors). Note that the expression of the A (FAM) allele (displayed on the left y axis) is reciprocal to the expression of the G (VIC) allele (displayed on the right y axis). Statistically significant differences between groups were calculated using unpaired two-tailed t-test. n indicates number of independent sub-clones for wild-type cells or independently deleted clones for ΔE4/ΔE4 cells; allele-specific expression for each clone was analysed in 3 independent biological replicate experiments at passage 1 and 2 respectively, each measured as 3 technical replicates. **P < 0.0001. j, qRT–PCR expression analysis for total SNCA in same samples as described in i. Data are normalized to GAPDH and displayed relative to the expression of ΔE4/ΔE4 neural precursors. Data are displayed as mean ± s.d.; Source Data and detailed statistical analysis are provided online.
Extended Data Figure 6 Conditional GWAS and eQTL analysis to assess the effect of Parkinson’s disease risk SNP rs356168.
a, Table showing baseline and conditional GWAS analysis in 5 publicly available PD GWAS cohorts (totalling 6,014 cases and 9,119 controls) to assess the extent to which rs356168 explains the observed associations in the SNCA locus of the top reported GWAS SNP rs356182 and the independent 5′ region SNP rs7681154 (ref. 12). b, qRT–PCR analysis for SNCA expression in postmortem frontal cortex tissue obtained from Parkinson’s disease patients and controls stratified by risk genotype at rs356168 (number of brain samples from Parkinson’s disease patients and controls are indicated for each genotype). Expression models were analysed including adjustment for disease status, sex, pH, age at death, as well as for the interaction between PMI and disease status. Significance was assessed using a one-sided test based on the a priori hypothesis of an association between the G allele at rs356168 and increased expression of SNCA. P values comparing each genotype are displayed in the graph. Alternatively analysis using linear regression shows a significant increase of total SNCA levels in carriers of the G allele at rs356168 (P = 0.031).
Extended Data Figure 7 Parkinson’s disease associated risk variants have little effect on enhancer-specific chromatin modifications at the intron-4 enhancer.
a, Overview of isogenic cell lines (derived from WIBR3 human ES cells) carrying distinct genotypes of the intron-4 enhancer element used for chromatin-immunoprecipitation (ChIP-qRT–PCR and ChIP-seq). b, c, ChIP-qRT–PCR analysis in neurons derived from isogenic cell lines with indicated genotypes for binding of the enhancer-specific chromatin marks H3K4me1 (b) and H3K27ac (c) at the intron-4 and 3′ UTR enhancer sequences compared with indicated negative control regions (calculated as percent of input). d, Gene tracks of ChIP-seq analysis for the active enhancer mark H3K27ac in in vitro differentiated neurons derived from isogenic cell lines with indicated genotypes. e, Quantitative read density analysis of H3K27ac ChIP-seq data (as shown in d) displaying relative read density of the intron-4 enhancer compared to the 3′ UTR enhancer to control for variability between ChIP experiments. Source Data for this figure are available online.
Extended Data Figure 8 Sequence-specific binding of brain-expressed TFs EMX2 and NKX6-1 at SNCA intron-4 enhancer.
a, ChIP-qRT–PCR for binding of indicated TFs at the intron-4 enhancer element compared with negative control region in the SNCA locus (calculated as fold enrichment compared with IgG Isotype control) in human ES cell line BGO1 and human iPS cell line IPS-PDC16 (derived from fibroblast AG20446). b, d, Western blot analysis in nuclear extracts (NE) (used for experiments displayed in Fig. 3b, c and Extended Data Fig. 8c, e) from HEK293 cells overexpressing Myc-DDK-tagged EMX2 (b) or NKX6-1 (d) using indicated antibodies against DDK-(Flag)-tag, EMX2 and NKX6-1 respectively. Lamin A was used to control for equal loading in NEs. c, e, EMSA analysis to determine TF concentration-dependent sequence-specific binding of EMX2 (c) and NKX6-1 (e) to oligonucleotides harbouring the indicated genotype at rs356168 (A/G allele). NEs from HEK293 cells overexpressing Myc-DDK(Flag)-tagged EMX2 (c) or NKX6-1 (e) were diluted with NEs from wild-type cells at indicated fractions to generate a TF concentration gradient. Relative EMX2 and NKX6-1 protein concentration in mixed samples were determined by western blot analysis using an antibody against the DDK(Flag)-tag (panel below respective EMSA). Red arrows point to oligonucleotide-specific binding of overexpressed TFs. Source Data for this figure are available online.
Extended Data Figure 9 Single-molecule mRNA FISH analysis and immunostaining for TFs EMX2 and NKX6-1 in mixed neuronal cultures.
a, Single-molecule mRNA FISH for EMX2-(Cy5) and NKX6-1-(Alexa594) (displayed in false colour) labelling EMX2 and NKX6-1 mRNA transcripts in WIBR3-derived in vitro differentiated neurons (differentiation day 21). Cultured neurons were dissociated before hybridization and attached to a glass slide before imaging. Representative image shows multiple cells, which are either single-positive for EMX2 (green arrowhead) or double-positive for EMX2 and NKX6-1 (white arrowhead). b–d, Representative images showing individual cells which are either double positive for the expression of EMX2 and NKX6-1 (b) or single positive for either NKX6-1 (c) or EMX2 (d). e, Quantification of 100 individual cells for the presence of EMX2 and NKX6-1 transcripts. f–j, Co-immunostaining for EMX2, NKX6-1, neuronal-specific beta-III-tubulin (TUJ1) and astrocyte-specific glial fibrillary acidic protein (GFAP) in mixed neuronal cultures. Source Data for this figure are available online.
Extended Data Figure 10 SNCA-Rep1 repeat length has no cis-acting effect on SNCA expression in human ES-cell-derived neurons.
a, Schematic illustration of the CRISPR/Cas9-mediated genome editing strategy to generate heterozygous deletions (ΔRep1/wild-type) in WIBR3 and BGO1 human ES cells and subsequently insert indicated SNCA-Rep1 length variants. Displayed are the genomic organization of the SNCA locus, an enlarged view of the wild-type and SNCA-Rep1 deleted (ΔRep1) allele, gRNA-targeting sequences (underlined, PAM sequence in red), restriction sites and Southern blot (SB) probe. Shown below is targeting vector (TV) design to insert the respective SNCA-Rep1 elements (Rep1-257, Rep1-259, Rep1-261 and Rep1-263) with indicated repeat sequences. Only clones with heterozygous deletion of the repeat element on the same chromosome were identified based on the genotype of two heterozygous SNPs (rs58864428 and rs10030935) upstream and downstream of SNCA-Rep1 and selected for subsequent experiments. b, Representative Southern blot analysis of wild type and targeted WIBR3 human ES cells with indicated SNCA-Rep1 genotypes (modified alleles highlighted in red; unmodified wild-type allele represents SNCA-Rep1-261 element). c, Table summarizing SNCA-Rep1 deletions and insertions in WIBR3 and BGO1 human ES cells. d, Representative fragment length analysis confirms expected SNCA-Rep-1 repeat length in targeted cell lines. Red line indicates SNCA-Rep1-261 peak at 269 bp. e, f, Analysis of relative allele-specific SNCA expression in neurons (differentiation day 25) derived from targeted cells lines with indicated SNCA-Rep1 alleles compared with untargeted controls in WIBR3 (e) and BGO1 (f) human ES cells (expression was normalized relative to wild-type cells). Shown are mean values ± s.d. of three independent biological replicate experiments for each individual clone of the indicated genotype. Differences between individual clones or combined clones by genotypes were not significant based on one-way ANOVA testing for multiple comparisons between groups (alpha = 0.05) and did not show a significant repeat length-dependent linear trend as analysed by linear regression. Source Data for this figure are available online.
Supplementary information
Supplementary Table 1
Prioritization of PD-associated risk variants in the SNCA locus based on overlap with enhancer specific epigenetic marks and predicted transcription factor binding. (XLSX 173 kb)
Supplementary Table 2
Predicted differential transcription factor binding at SNP rs356168. (XLSX 36 kb)
Supplementary Table 3
Detailed lists of gRNAs, primers, donor constructs, antibodies, Gene Expression Omnibus (GEO) identifiers for Chip-seq and DHSs datasets and CRISPR/Cas9 genome edited cell lines used in this manuscript (see Methods). (PDF 100 kb)
Rights and permissions
About this article
Cite this article
Soldner, F., Stelzer, Y., Shivalila, C. et al. Parkinson-associated risk variant in distal enhancer of α-synuclein modulates target gene expression. Nature 533, 95–99 (2016). https://doi.org/10.1038/nature17939
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17939
This article is cited by
-
Precise modulation of transcription factor levels identifies features underlying dosage sensitivity
Nature Genetics (2023)
-
Optimization of Cas9 activity through the addition of cytosine extensions to single-guide RNAs
Nature Biomedical Engineering (2023)
-
Whole-genome sequencing reveals an association between small genomic deletions and an increased risk of developing Parkinson’s disease
Experimental & Molecular Medicine (2023)
-
CRISPR: a tool with potential for genomic reprogramming in neurological disorders
Molecular Biology Reports (2023)
-
The threat of programmed DNA damage to neuronal genome integrity and plasticity
Nature Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.