Abstract
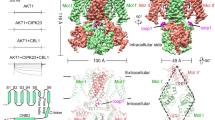
Two-pore channels (TPCs) comprise a subfamily (TPC1–3) of eukaryotic voltage- and ligand-gated cation channels1,2 with two non-equivalent tandem pore-forming subunits that dimerize to form quasi-tetramers. Found in vacuolar3 or endolysosomal4 membranes, they regulate the conductance of sodium5 and calcium3,6 ions, intravesicular pH5, trafficking7 and excitability8,9. TPCs are activated by a decrease in transmembrane potential1,3,9,10 and an increase in cytosolic calcium concentrations1,10, are inhibited by low luminal pH and calcium11, and are regulated by phosphorylation12,13. Here we report the crystal structure of TPC1 from Arabidopsis thaliana at 2.87 Å resolution as a basis for understanding ion permeation3,4,10, channel activation1,5,10, the location of voltage-sensing domains1,9,10 and regulatory ion-binding sites11,14. We determined sites of phosphorylation3,4 in the amino-terminal and carboxy-terminal domains that are positioned to allosterically modulate cytoplasmic Ca2+ activation. One of the two voltage-sensing domains (VSD2) encodes voltage sensitivity and inhibition by luminal Ca2+ and adopts a conformation distinct from the activated state observed in structures of other voltage-gated ion channels15,16. The structure shows that potent pharmacophore trans-Ned-19 (ref. 17) acts allosterically by clamping the pore domains to VSD2. In animals, Ned-19 prevents infection by Ebola virus and other filoviruses, presumably by altering their fusion with the endolysosome and delivery of their contents into the cytoplasm7.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hedrich, R. & Neher, E. Cytoplasmic calcium regulates voltage-dependent ion channels in plant vacuoles. Nature 329, 833–836 (1987)
Calcraft, P. J. et al. NAADP mobilizes calcium from acidic organelles through two-pore channels. Nature 459, 596–600 (2009)
Hedrich, R. & Marten, I. TPC1 – SV Channels Gain Shape. Mol. Plant 4, 428–441 (2011)
Patel, S. Function and dysfunction of two-pore channels. Sci. Signal. 8, re7 (2015)
Wang, X. et al. TPC proteins are phosphoinositide- activated sodium-selective ion channels in endosomes and lysosomes. Cell 151, 372–383 (2012)
Pitt, S. J. et al. TPC2 is a novel NAADP-sensitive Ca2+ release channel, operating as a dual sensor of luminal pH and Ca2+. J. Biol. Chem. 285, 35039–35046 (2010)
Sakurai, Y. et al. Two-pore channels control Ebola virus host cell entry and are drug targets for disease treatment. Science 347, 995–998 (2015)
Hedrich, R., Flügge, U. I. & Fernandez, J. M. Patch-clamp studies of ion transport in isolated plant vacuoles. FEBS Lett. 204, 228–232 (1986)
Cang, C., Bekele, B. & Ren, D. The voltage-gated sodium channel TPC1 confers endolysosomal excitability. Nature Chem. Biol. 10, 463–469 (2014)
Guo, J. et al. Structure of the voltage-gated two-pore channel TPC1 from Arabidopsis thaliana. Nature http://dx.doi.org/10.1038/nature16446 (this issue)
Beyhl, D. et al. The fou2 mutation in the major vacuolar cation channel TPC1 confers tolerance to inhibitory luminal calcium. Plant J. 58, 715–723 (2009)
Cang, C. et al. mTOR regulates lysosomal ATP-sensitive two-pore Na+ channels to adapt to metabolic state. Cell 152, 778–790 (2013)
Morgan, A. J., Davis, L. C., Ruas, M. & Galione, A. TPC: the NAADP discovery channel? Biochem. Soc. Trans. 43, 384–389 (2015)
Schulze, C., Sticht, H., Meyerhoff, P. & Dietrich, P. Differential contribution of EF-hands to the Ca2+-dependent activation in the plant two-pore channel TPC1. Plant J. 68, 424–432 (2011)
Long, S. B., Campbell, E. B. & Mackinnon, R. Crystal structure of a mammalian voltage-dependent Shaker family K+ channel. Science 309, 897–903 (2005)
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011)
Naylor, E. et al. Identification of a chemical probe for NAADP by virtual screening. Nature Chem. Biol. 5, 220–226 (2009)
Kamb, A., Iverson, L. E. & Tanouye, M. A. Molecular characterization of Shaker, a Drosophila gene that encodes a potassium channel. Cell 50, 405–413 (1987)
Papazian, D. M., Schwarz, T. L., Tempel, B. L., Jan, Y. N. & Jan, L. Y. Cloning of genomic and complementary DNA from Shaker, a putative potassium channel gene from Drosophila. Science 237, 749–753 (1987)
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350, aad2395 (2015)
Catterall, W. A., Perez-Reyes, E., Snutch, T. P. & Striessnig, J. International Union of Pharmacology. XLVIII. Nomenclature and structure-function relationships of voltage-gated calcium channels. Pharmacol. Rev. 57, 411–425 (2005)
Brailoiu, E. et al. Essential requirement for two-pore channel 1 in NAADP-mediated calcium signaling. J. Cell Biol. 186, 201–209 (2009)
Peterson, B. Z., Tanada, T. N. & Catterall, W. A. Molecular determinants of high affinity dihydropyridine binding in L-type calcium channels. J. Biol. Chem. 271, 5293–5296 (1996)
Pitt, S. J., Lam, A. K. M., Rietdorf, K., Galione, A. & Sitsapesan, R. Reconstituted human TPC1 is a proton-permeable ion channel and is activated by NAADP or Ca2+. Sci. Signal. 7, ra46 (2014)
Tang, L. et al. Structural basis for Ca2+ selectivity of a voltage-gated calcium channel. Nature 505, 56–61 (2014)
Armstrong, C. M. Time course of TEA+-induced anomalous rectification in squid giant axons. J. Gen. Physiol. 50, 491–503 (1966)
Holmgren, M., Smith, P. L. & Yellen, G. Trapping of organic blockers by closing of voltage-dependent K+ channels: evidence for a trap door mechanism of activation gating. J. Gen. Physiol. 109, 527–535 (1997)
Perozo, E., Cortes, D. M. & Cuello, L. G. Three-dimensional architecture and gating mechanism of a K+ channel studied by EPR spectroscopy. Nature Struct. Mol. Biol . 5, 459–469 (1998)
Clayton, G. M., Altieri, S., Heginbotham, L., Unger, V. M. & Morais-Cabral, J. H. Structure of the transmembrane regions of a bacterial cyclic nucleotide-regulated channel. Proc. Natl Acad. Sci. USA 105, 1511–1515 (2008)
Zhou, Y., Morais-Cabral, J. H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by a K+ channel–Fab complex at 2.0 Å resolution. Nature 414, 43–48 (2001)
Mumberg, D., Müller, R. & Funk, M. Regulatable promoters of Saccharomyces cerevisiae: comparison of transcriptional activity and their use for heterologous expression. Nucleic Acids Res. 22, 5767–5768 (1994)
Hattori, M., Hibbs, R. E. & Gouaux, E. A fluorescence-detection size-exclusion chromatography-based thermostability assay for membrane protein precrystallization screening. Structure 20, 1293–1299 (2012)
Gourdon, P. et al. HiLiDe—systematic approach to membrane protein crystallization in lipid and detergent. Cryst. Growth Des. 11, 2098–2106 (2011)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 103, 8060–8065 (2006)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Abrahams, J. P. & Leslie, A. G. W. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
Cowtan, K. ‘dm’: an automated procedure for phase improvement by density modification, in Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography. 31, 34–38 http://www.ccp4.ac.uk/newsletters/newsletter36/07_dm.html (1994)
Pedersen, B. P., Morth, J. P. & Nissen, P. Structure determination using poorly diffracting membrane-protein crystals: the H+-ATPase and Na+,K+-ATPase case history. Acta Crystallogr. D 66, 309–313 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004)
Bond, C. S. & Schüttelkopf, A. W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D 65, 510–512 (2009)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. HOLE: A program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996)
Jones, D. T. Protein secondary structure prediction based on position-specific scoring matrices1. J. Mol. Biol. 292, 195–202 (1999)
Cuff, J. A., Clamp, M. E., Siddiqui, A. S., Finlay, M. & Barton, G. J. JPred: a consensus secondary structure prediction server. Bioinformatics 14, 892–893 (1998)
Blom, N., Sicheritz-Pontén, T., Gupta, R., Gammeltoft, S. & Brunak, S. Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4, 1633–1649 (2004)
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013)
Paulsen, C. E., Armache, J.-P., Gao, Y., Cheng, Y. & Julius, D. Structure of the TRPA1 ion channel suggests regulatory mechanisms. Nature 520, 511–517 (2015)
Acknowledgements
We are grateful to J. M. Holton and G. Meigs at ALS beamline 8.3.1 and A. Gonzalez at SSRL beamline 12-2 for help with data collection and processing, B. P. Pedersen, B. H. Schmidt and J. S. Finer-Moore for critical analysis of the manuscript. This work was supported by NIH grant GM24485 to R.M.S. Beamline 8.3.1 at the Advanced Light Source is operated by the University of California Office of the President, Multicampus Research Programs and Initiatives grant MR-15-328599 and Program for Breakthrough Biomedical Research, which is partially funded by the Sandler Foundation. Figures were made with Chimera developed by the Resource for Biocomputing, Visualization and Informatics at the University of California, San Francisco (supported by NIGMS P41-GM103311), and PyMOL developed by W. DeLano. Mass spectrometry was carried out by the UCSF mass spectrometry facility (supported by NIGMS P41-RR001614). The Berkeley Center for Structural Biology is supported in part by the National Institutes of Health, National Institute of General Medical Sciences, and the Howard Hughes Medical Institute. The Advanced Light Source is supported by the Director, Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under Contract No. DE-AC02-05CH11231. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIGMS or NIH.
Author information
Authors and Affiliations
Contributions
A.F.K. performed the experiments, collected and processed the data, determined, refined, and analysed the structure. R.M.S. supervised the project and aided in structure determination and analysis. A.F.K. and R.M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of TPCs with TPC1 experimental and predicted secondary structure.
A sequence alignment based on seven human and plant TPC orthologues with observed TPC1 (black), and secondary structure predictions by Psipred48 (red) and Jpred49 (blue). Helices are shown as cylinders, β sheets as planks, coils as solid lines, and unstructured regions as dashed lines. Level of conservation is indicated by colour (>50% yellow; >80% red). Blue dots mark arginines in S4; red dots mark charge-transfer anions; green dots mark Ca2+-binding sites in the EF-hand. Solid and dashed green lines represent observed and absent peptides from mass spectrometry experiments. Coloured stars indicate observed phosphorylation sites using electrospray ionization mass spectrometry (magenta; Extended Data Fig. 3a), potential phosphorylation sites identified by truncation constructs (orange; Extended Data Fig. 3e), predicted phosphorylation sites by NetPhosK50 (cyan), non-phosphorylated sites (blue), unlikely sites owing to solvent inaccessibility (black), and unknown or unidentified regions (grey).
Extended Data Figure 2 Crystal packing of TPC1 and experimental electron densities.
a, Views of the TPC1 C2221 crystal lattice viewed along two-fold axes parallel to a (left) and b (right). Unit cell boundaries are shown. The asymmetric unit is shown in yellow. b, Transverse view of TPC1 is shown with overlayed FOM-weighted experimental electron density calculated using native amplitudes (native 1) and heavy-atom phases contoured at 1σ. Density-modified phases from non-dehydrated derivatives (left; Extended Data Table 1), dehydrated derivatives (middle; Extended Data Table 2), and combined phases with solvent flattening, histogram matching, phase extension to high resolution native (native 2), and cross-crystal averaging (right; see Methods, Extended Data Table 3). c, Transverse views of TPC1 with overlayed heavy-atom electron densities calculated from isomorphous differences.
Extended Data Figure 3 Determination of phosphorylation sites in TPC1.
a, Electrospray ionization mass spectrometry peptide sequence coverage from an in-gel digest of wild-type (WT) TPC1 by four enzyme combinations (Trypsin/Asp-N, Trypsin/Glu-C, Lys-C or Chymotrypsin). Measured peptides (top, red highlight), predicted (middle, black star) and observed (middle, red star) phosphorylation sites are shown. Observed phosphopeptides are listed with the sites of phosphorylation coloured red. b–e, Polyacrylamide gels of purified TPC1 (10 μg) stained with phosphoprotein-specific probe ProDiamond-Q, SyproRuby or Coomassie (left to right). Molecular weights of standards are indicated in kilodaltons. The first two lanes are PrecisionPlus protein molecular weight standards and PeppermintStick phosphoprotein molecular weight standards. b, Wild-type TPC1 and crystallographic TPC1. c, Untreated and treated wild-type TPC1 with lambda phosphatase for 1 h at 25 °C. d, Schematic of TPC1 truncations (NΔ1; 2–11, NΔ2; 2–21, NΔ3; 2–30, CΔ1; 682–733, CΔ2; 693–733, CΔ3; 709–733, CΔ4; 724–733). e, Analysis of TPC1 N- and C-terminal truncations for binding to ProDiamond-Q. NΔ1CΔ1 was unstable during purification.
Extended Data Figure 4 Sequence alignment of TPC1 subdomains with hTPC1, hTPC2, human Cav 1.1–1.4, Nav 1.1, NavAb and hCaM.
a, S4 voltage-sensing segments. Conserved arginines are highlighted in cyan. Stars mark potential voltage-sensing residues. b, S2 segments. Stars mark conserved charge-transfer residues. c, S6 segments, pore gate and poly-E motif. Conserved hydrophobic residues are highlighted in cyan. Dihydropyridine-binding residues in the Cav S6 domain 3, domain 4, and corresponding residues in TPC1 are highlighted in red and green, respectively. Stars mark the position of phenylalkylamine drug-binding residues in Cavs. Conserved residues in the gate and poly-E motif are highlighted in magenta and purple, respectively. d, Pore loops. Conserved residues are highlighted in red. e, Alignment of hCaM EF-hand domains (EF-hand 1, residues 1–71; EF-hand 2, residues 72–149) and AtTPC1 (residues 322–398). Stars mark calcium binding motifs and orange dots mark the interaction site with CTD.
Extended Data Figure 5 Comparison of VSDs between TPC1 and symmetrical ion channels.
a–e, Structural alignment of S11–S12 segments of TPC1 domain 2 (blue) with S5–S6 of TPC1 domain 1 (a; red), NavAb16 (b; PDB ID 3RVY, grey); Kv1.2 (c; PDB ID 2A79, green)15, TRPV1 (d; PDB ID 3J5P, orange)51, and TRPA1 (e; PDB ID 3J9P, yellow)52. Angles between S4 segments with respect to S10 of TPC1 domain 2 are shown.
Extended Data Figure 6 Electrostatic surfaces and hydrophobic surfaces.
a, b, Top from the luminal side (left) and bottom from the cytoplasmic side (right) views of an electrostatic surface representation (a) and surface representation, coloured according to Kyte–Doolittle hydropathy of TPC1 (b). Notable domains and residues are labelled. Electrostatic potential and Kyte–Doolittle hydropathy without any bound ions were generated using Chimera46. The EF-hand and NTD domains are negatively charged and bind cations. The poly-R motif accounts for the positively charged region in the CTD.
Extended Data Figure 7 Mechanism for TPC gating.
A schematic summarizing structural features of TPC1 that suggest mechanisms for voltage-sensing, ion permeation, inhibition by Ned-19, luminal Ca2+ inhibition, cytosolic Ca2+-activation (EF-hand), and phosphoregulation (NTD/CTD). Ca2+ (green) and lanthanide (red) ion binding sites are shown. An ion permeation pathway through the putative selectivity filter, gate, and poly-E and poly-R motifs are summarized. M+ represents a general cation (Na+, K+, Ca2+) and ‘+’ signs are gating charges.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary Table 1 and Supplementary References. (PDF 227 kb)
Rights and permissions
About this article
Cite this article
Kintzer, A., Stroud, R. Structure, inhibition and regulation of two-pore channel TPC1 from Arabidopsis thaliana. Nature 531, 258–264 (2016). https://doi.org/10.1038/nature17194
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17194
This article is cited by
-
Two-pore channel blockade by phosphoinositide kinase inhibitors YM201636 and PI-103 determined by a histidine residue near pore-entrance
Communications Biology (2022)
-
Salinity tolerance mechanisms and their breeding implications
Journal of Genetic Engineering and Biotechnology (2021)
-
Parallel adaptation in autopolyploid Arabidopsis arenosa is dominated by repeated recruitment of shared alleles
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.