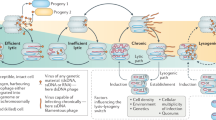
Abstract
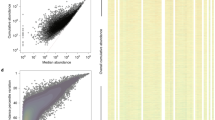
Microbial viruses can control host abundances via density-dependent lytic predator–prey dynamics. Less clear is how temperate viruses, which coexist and replicate with their host, influence microbial communities. Here we show that virus-like particles are relatively less abundant at high host densities. This suggests suppressed lysis where established models predict lytic dynamics are favoured. Meta-analysis of published viral and microbial densities showed that this trend was widespread in diverse ecosystems ranging from soil to freshwater to human lungs. Experimental manipulations showed viral densities more consistent with temperate than lytic life cycles at increasing microbial abundance. An analysis of 24 coral reef viromes showed a relative increase in the abundance of hallmark genes encoded by temperate viruses with increased microbial abundance. Based on these four lines of evidence, we propose the Piggyback-the-Winner model wherein temperate dynamics become increasingly important in ecosystems with high microbial densities; thus ‘more microbes, fewer viruses’.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
The viromes and microbiomes used in this paper are accessible at MG-RAST (http://metagenomics.anl.gov/) under the Piggyback-the-Winner project. Virome accession numbers: 4683670.3, 4683674.3, 4683677.3, 4683680.3, 4683683.3, 4683684.3, 4683686.3, 4683690.3, 4683702.3, 4683703.3, 4683704.3, 4683706.3, 4683712.3, 4683720.3, 4683739.3, 4683744.3, 4683745.3, 4683746.3, 4683747.3, 4683731.3, 4683733.3, 4683734.3, 4683718.3, 4684617.3. Microbiome accession numbers: 4683666.3, 4683667.3, 4683668.3, 4683669.3, 4683671.3, 4683672.3, 4683673.3, 4683675.3, 4683676.3, 4683678.3, 4683679.3, 4683681.3, 4683682.3, 4683685.3, 4683687.3, 4683688.3, 4683689.3, 4683691.3, 4683692.3, 4683693.3, 4683694.3, 4683695.3, 4683696.3, 4683697.3, 4683698.3, 4683699.3, 4683700.3, 4683701.3, 4683705.3, 4683707.3, 4683708.3, 4683709.3, 4683710.3, 4683711.3, 4683713.3, 4683714.3, 4683715.3, 4683716.3, 4683717.3, 4683719.3, 4683721.3, 4683722.3, 4683723.3, 4683724.3, 4683725.3, 4683726.3, 4683727.3, 4683728.3, 4683729.3, 4683732.3, 4683735.3, 4683736.3, 4683737.3, 4683738.3, 4683740.3, 4683741.3, 4683742.3, 4683743.3, 4683748.3, 4683749.3, 4683750.3, 4683751.3, 4683752.3, 4683753.3, 4683754.3, 4684616.3.
Change history
24 August 2016
A Correction to this paper has been published: https://doi.org/10.1038/nature19335
References
Suttle, C. A. Marine viruses — major players in the global ecosystem. Nature Rev. Microbiol. 5, 801–812 (2007)
Proctor, L. M. & Fuhrman, J. A. Viral mortality of marine bacteria and cyanobacteria. Nature 343, 60–62 (1990)
Wilcox, R. M. & Fuhrman, J. A. Bacterial viruses in coastal seawater: lytic rather than lysogenic production. Mar. Ecol. Ser . 114, 35–45 (1994)
Payet, J. & Suttle, C. A. To kill or not to kill: the balance between lytic and lysogenic viral infection is driven by trophic status. Limnol. Oceanogr. 58, 465–474 (2013)
Wommack, K. E. & Colwell, R. R. Virioplankton: viruses in aquatic ecosystems. Microbiol. Mol. Biol. Rev. 64, 69–114 (2000)
Thingstad, T. F. Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol. Oceanogr. 45, 1320–1328 (2000)
Rodriguez-Brito, B. et al. Viral and microbial community dynamics in four aquatic environments. ISME J. 4, 739–751 (2010)
Weitz, J. S. & Dushoff, J. Alternative stable states in host–phage dynamics. Theor. Ecol . 1, 13–19 (2008)
Thingstad, T. F., Våge, S., Storesund, J. E., Sandaa, R.-A. & Giske, J. A theoretical analysis of how strain-specific viruses can control microbial species diversity. Proc. Natl Acad. Sci. USA 111, 7813–7818 (2014)
Weinbauer, M. G. & Höfle, M. G. Significance of viral lysis and flagellate grazing as factors controlling bacterioplankton production in a eutrophic lake. Appl. Environ. Microbiol. 64, 431–438 (1998)
Bratbak, G., Egge, J. K. & Heldal, M. Viral mortality of the marine alga Emiliania huxleyi (Haptophyceae) and termination of algal blooms. Mar. Ecol. Prog. Ser. 93, 39–48 (1993)
Evans, C. & Brussaard, C. P. D. Viral lysis and microzooplankton grazing of phytoplankton throughout the Southern Ocean. Limnol. Oceanogr. 57, 1826–1837 (2012)
Needham, D. M. et al. Short-term observations of marine bacterial and viral communities: patterns, connections and resilience. ISME J. 7, 1274–1285 (2013)
Jiang, S. C. & Paul, J. H. Significance of lysogeny in the marine environment: studies with isolates and a model of lysogenic phage production. Microb. Ecol. 35, 235–243 (1998)
Paul, J. H. Prophages in marine bacteria: dangerous molecular time bombs or the key to survival in the seas? ISME J. 2, 579–589 (2008)
Paul, J. H. & Weinbauer, M. Detection of lysogeny in marine environments. Man. Aquat. Viral Ecol . 4, 30–33 (2010)
Maurice, C. F., Bouvier, C., de Wit, R. & Bouvier, T. Linking the lytic and lysogenic bacteriophage cycles to environmental conditions, host physiology and their variability in coastal lagoons. Environ. Microbiol. 15, 2463–2475 (2013)
Dinsdale, E. A. et al. Microbial ecology of four coral atolls in the Northern Line Islands. PLoS One 3, e1584 (2008)
Smith, J. E. et al. Indirect effects of algae on coral: algae-mediated, microbe-induced coral mortality. Ecol. Lett. 9, 835–845 (2006)
Thurber, R. V. et al. Metagenomic analysis of stressed coral holobionts. Environ. Microbiol. 11, 2148–2163 (2009)
Kelly, L. W. et al. Black reefs: iron-induced phase shifts on coral reefs. ISME J. 6, 638–649 (2012)
McDole, T. et al. Assessing coral reefs on a Pacific-wide scale using the microbialization score. PLoS One 7, e43233 (2012)
Barott, K. L. & Rohwer, F. L. Unseen players shape benthic competition on coral reefs. Trends Microbiol. 20, 621–628 (2012)
Alongi, D. M. et al. Phytoplankton, bacterioplankton and virioplankton structure and function across the southern Great Barrier Reef shelf. J. Mar. Syst . 142, 25–39 (2015)
Thingstad, T. F. & Lignell, R. Theoretical models for the control of bacterial growth rate, abundance, diversity and carbon demand. Aquat. Microb. Ecol. 13, 19–27 (1997)
Giovannoni, S., Temperton, B. & Zhao, Y. Giovannoni et al. reply. Nature 499, E4–E5 (2013)
James, C. E. et al. Lytic activity by temperate phages of Pseudomonas aeruginosa in long-term cystic fibrosis chronic lung infections. ISME J. 9, 1391–1398 (2015)
Muck, S. et al. Fracture zones in the Mid Atlantic Ridge lead to alterations in prokaryotic and viral parameters in deep-water masses. Front. Microbiol . 5, 264 (2014)
Bongiorni, L., Magagnini, M., Armeni, M., Noble, R. & Danovaro, R. Viral production, decay rates, and life strategies along a trophic gradient in the North Adriatic Sea. Appl. Environ. Microbiol. 71, 6644–6650 (2005)
Williamson, S. J., Houchin, L. A., McDaniel, L. & Paul, J. H. Seasonal variation in lysogeny as depicted by prophage induction in Tampa Bay, Florida. Appl. Environ. Microbiol. 68, 4307–4314 (2002)
Hennes, K. P., Suttle, C. A. & Chan, A. M. Fluorescently labeled virus probes show that natural virus populations can control the structure of marine microbial communities. Appl. Environ. Microbiol. 61, 3623–3627 (1995)
McDaniel, L. D., Rosario, K., Breitbart, M. & Paul, J. H. Comparative metagenomics: natural populations of induced prophages demonstrate highly unique, lower diversity viral sequences. Environ. Microbiol. 16, 570–585 (2014)
Fuhrman, J. A. Microbial community structure and its functional implications. Nature 459, 193–199 (2009)
Weitz, J. S. et al. A multitrophic model to quantify the effects of marine viruses on microbial food webs and ecosystem processes. ISME J. 9, 1352–1364 (2015)
Breitbart, M. et al. Metagenomic analyses of an uncultured viral community from human feces. J. Bacteriol. 185, 6220–6223 (2003)
Reyes, A. et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466, 334–338 (2010)
Angly, F. E. et al. The marine viromes of four oceanic regions. PLoS Biol. 4, e368 (2006)
Breitbart, M. et al. Diversity and population structure of a near-shore marine-sediment viral community. Proc. R. Soc. Lond. B 271, 565–574 (2004)
Brum, J. R., Hurwitz, B. L., Schofield, O., Ducklow, H. W. & Sullivan, M. B. Seasonal time bombs: dominant temperate viruses affect Southern Ocean microbial dynamics. ISME J. 10, 437–449 (2015)
Våge, S., Storesund, J. E. & Thingstad, T. F. Adding a cost of resistance description extends the ability of virus-host model to explain observed patterns in structure and function of pelagic microbial communities. Environ. Microbiol. 15, 1842–1852 (2013)
Avrani, S. & Lindell, D. Convergent evolution toward an improved growth rate and a reduced resistance range in Prochlorococcus strains resistant to phage. Proc. Natl Acad. Sci. USA 112, E2191–E2200 (2015)
Kelly, L. W. et al. Local genomic adaptation of coral reef-associated microbiomes to gradients of natural variability and anthropogenic stressors. Proc. Natl Acad. Sci. USA 111, 10227–10232 (2014)
Silveira, C. B. et al. Microbial and sponge loops modify fish production in phase-shifting coral reefs. Environ. Microbiol. 17, 3832–3846 (2015)
Barr, J. J. et al. Bacteriophage adhering to mucus provide a non-host-derived immunity. Proc. Natl Acad. Sci. USA 110, 10771–10776 (2013)
Whiteson, K. L. et al. The upper respiratory tract as a microbial source for pulmonary infections in cystic fibrosis. Parallels from island biogeography. Am. J. Respir. Crit. Care Med. 189, 1309–1315 (2014)
Willner, D. et al. Metagenomic analysis of respiratory tract DNA viral communities in cystic fibrosis and non-cystic fibrosis individuals. PLoS One 4, e7370 (2009)
Haas, A. F. et al. Unraveling the unseen players in the ocean - a field guide to water chemistry and marine microbiology. J. Vis. Exp . 93, e52131 (2014)
Brussaard, C. P. D., Payet, J. P., Winter, C. & Weinbauer, M. G. Quantification of aquatic viruses by flow cytometry. Man. Aquat. Viral Ecol . 11, 102–109 (2010)
Murray, J. D. Mathematical biology: I. an introduction . (Springer, 2002)
Amossé, J. et al. The flows of nitrogen, bacteria and viruses from the soil to water compartments are influenced by earthworm activity and organic fertilization (compost vs. vermicompost). Soil Biol. Biochem. 66, 197–203 (2013)
Bettarel, Y., Bouvy, M., Dumont, C. & Sime-Ngando, T. Virus-bacterium interactions in water and sediment of West African inland aquatic systems. Appl. Environ. Microbiol. 72, 5274–5282 (2006)
Bouvier, T. & Maurice, C. F. A single-cell analysis of virioplankton adsorption, infection, and intracellular abundance in different bacterioplankton physiologic categories. Microb. Ecol. 62, 669–678 (2011)
Glud, R. N. & Middelboe, M. Virus and bacteria dynamics of a coastal sediment: implication for benthic carbon cycling. Limnol. Oceanogr. 49, 2073–2081 (2004)
Furlan, M. Viral and microbial dynamics in the human respiratory tract . (San Diego State Univ., 2009)
Hewson, I., O’Neil, J. M., Fuhrman, J. A. & Dennison, W. C. Virus-like particle distribution and abundance in sediments and overlying waters along eutrophication gradients in two subtropical estuaries. Limnol. Oceanogr. 46, 1734–1746 (2001)
Kim, M.-S., Park, E.-J., Roh, S. W. & Bae, J.-W. Diversity and abundance of single-stranded DNA viruses in human feces. Appl. Environ. Microbiol. 77, 8062–8070 (2011)
Lisle, J. T. & Priscu, J. C. The occurrence of lysogenic bacteria and microbial aggregates in the lakes of the McMurdo Dry Valleys, Antarctica. Microb. Ecol. 47, 427–439 (2004)
Laybourn-Parry, J., Marshall, W. A. & Madan, N. J. Viral dynamics and patterns of lysogeny in saline Antarctic lakes. Polar Biol. 30, 351–358 (2006)
Madan, N. J., Marshall, W. a. & Laybourn-Parry, J. Virus and microbial loop dynamics over an annual cycle in three contrasting Antarctic lakes. Freshw. Biol. 50, 1291–1300 (2005)
Maurice, C. F., Bouvier, T., Comte, J., Guillemette, F. & Del Giorgio, P. A. Seasonal variations of phage life strategies and bacterial physiological states in three northern temperate lakes. Environ. Microbiol. 12, 628–641 (2010)
Maurice, C. F. et al. Disentangling the relative influence of bacterioplankton phylogeny and metabolism on lysogeny in reservoirs and lagoons. ISME J. 5, 831–842 (2011)
Mei, M. L. & Danovaro, R. Virus production and life strategies in aquatic sediments. Limnol. Oceanogr. 49, 459–470 (2004)
Parsons, R. J., Breitbart, M., Lomas, M. W. & Carlson, C. A. Ocean time-series reveals recurring seasonal patterns of virioplankton dynamics in the northwestern Sargasso Sea. ISME J. 6, 273–284 (2012)
Parsons, R. J. et al. Marine bacterioplankton community turnover within seasonally hypoxic waters of a subtropical sound: Devil’s Hole, Bermuda. Environ. Microbiol. 17, 3481–3499 (2015)
Rinta-Kanto, J. M., Lehtola, M. J., Vartiainen, T. & Martikainen, P. J. Rapid enumeration of virus-like particles in drinking water samples using SYBR green I-staining. Water Res. 38, 2614–2618 (2004)
Schapira, M. et al. Distribution of heterotrophic bacteria and virus-like particles along a salinity gradient in a hypersaline coastal lagoon. Aquat. Microb. Ecol. 54, 171–183 (2009)
Payet, J. P., McMinds, R., Burkepile, D. E. & Vega Thurber, R. L. Unprecedented evidence for high viral abundance and lytic activity in coral reef waters of the South Pacific Ocean. Front. Microbiol . 5, 493 (2014)
Patten, N. L., Harrison, P. L. & Mitchell, J. G. Prevalence of virus-like particles within a staghorn scleractinian coral (Acropora muricata) from the Great Barrier Reef. Coral Reefs 27, 569–580 (2008)
Duhaime, M. B., Deng, L., Poulos, B. T. & Sullivan, M. B. Towards quantitative metagenomics of wild viruses and other ultra-low concentration DNA samples: a rigorous assessment and optimization of the linker amplification method. Environ. Microbiol. 14, 2526–2537 (2012)
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011)
Schmieder, R., Lim, Y. W., Rohwer, F. & Edwards, R. TagCleaner: Identification and removal of tag sequences from genomic and metagenomic datasets. BMC Bioinformatics 11, 341 (2010)
Schmieder, R. & Edwards, R. Fast identification and removal of sequence contamination from genomic and metagenomic datasets. PLoS One 6, e17288 (2011)
Silva, G. G. Z., Cuevas, D. A., Dutilh, B. E. & Edwards, R. A. FOCUS: an alignment-free model to identify organisms in metagenomes using non-negative least squares. PeerJ 2, e425 (2014)
Chevreux, B., Wetter, T. & Suhai, S. Genome sequence assembly using trace signals and additional sequence information. in German conference on bioinformatics 45–56 (1999)
Rho, M., Tang, H. & Ye, Y. FragGeneScan: predicting genes in short and error-prone reads. Nucleic Acids Res. 38, e191 (2010)
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012)
Angly, F. E. et al. The GAAS metagenomic tool and its estimations of viral and microbial average genome size in four major biomes. PLOS Comput. Biol. 5, e1000593 (2009)
Dutilh, B. E. et al. Reference-independent comparative metagenomics using cross-assembly: crAss. Bioinformatics 28, 3225–3231 (2012)
Sanchez, S. E. et al. Phage Phenomics: physiological approaches to characterize novel viral proteins. J. Vis. Exp . 100, e52854 (2015)
Acknowledgements
This paper is dedicated to the memory of Mike Furlan, mentor, friend, and colleague. We are grateful to the National Oceanographic and Atmospheric Administration Coral Reef Ecosystem Division for supporting this research, and to the captains and crews of the NOAA ship Hi’ialakai and the Hanse Explorer. Thanks to J. Payet for providing viral and microbial abundance data. Sampling was carried out under research permits from the US Fish and Wildlife Service, Palmyra Atoll National Wildlife Refuge, the Environment and Conservation Division of the Republic of Kiribati (n. 021/13) and ICMBio, Brazil (n. 27147-2). This work was funded by the Canadian Institute for Advanced Research Integrated Microbial Biodiversity Program Fellowship Award 141679 (to F.R.) and National Science Foundation grants OISE-1243541 and DEB-1046413 (to F.R.), CNS-1305112 and MCB-1330800 (to R.A.E.), DUE-1323809 (to E.A.D.), Gordon and Betty Moore Foundation Investigator Award GBMF-3781 (to F.R.), and the Brazilian National Research Council (CNPq; to F.T.) and Brazilian National Research Council Science Without Borders Program (CNPq/CAPES; to C.B.S.).
Author information
Authors and Affiliations
Contributions
F.R., B.K., C.B.S., and F.T. conceptualized the project; B.K., F.R., C.B.S., and M.Y. wrote the manuscript; B.K., C.B.S., V.A.C., A.G.C.-G., K.T.G, K.M., G.G.Z.S., S.D.Q., Y.W.L., S.E.S., F.H.C., E.R.H. , N.L.R., B.A.B., B.F., A.L., P.S., J.N., C.Y., E.E.G., M.L., K.A.F., L.S.O., T.M.-S., J.M.H., B.Z., A.F.H., M.J.A.V., K.B., C.S., R.A.E., and F.R. performed sample collection, processing, experiments, and analysis; N.H. provided graphics and GIS analysis; E.A.D., L.W.K., S.S., J.S., R.B., C.T., G.B.G., J.N., E.S., R.A.E., F.T., and F.R. provided intellectual guidance and funding during the development of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 The observed decline in virus to microbe ratio with increasing host density is not supported by horizontal transfer (for example, of resistance genes) under conditions where strain diversity is predicted to rise.
a, Host competence gene composition likely does not facilitate the expected rise in resistance to viral infection (n = 66; m = −0.25, t = −2.40, d.f. = 64, P = 0.02; R2 = 0.08; microbial abundance log-transformed; linear regression). b, Lysogeny may provide strain diversification similar to the co-evolutionary diversification predicted by Thingstad et al. (2014)9 nested-infection chemostat model.
Extended Data Figure 2 Meta-analysis of the frequency of lysogenic cells (FLC) from mitomycin C induction experiments yields ambiguous results.
FLC from four published studies is plotted against total cell abundance. Although a sometimes-significant negative relationship exists at a within-study level (microbial abundance log-transformed; Muck et al. (2014)28, n = 9, m = −10.79, t = −1.76, d.f. = 7, P = 0.12; R2 = 0.31; Bongiorni et al. (2005)29, n = 4, m = −17.23, t = −1.91, d.f. = 2, P = 0.20; R2 = 0.65; Payet and Suttle (2013)4, n = 9, m = −48.31, t = −4.80, d.f. = 7, P = 1.96 × 10−3; R2 = 0.77; Williamson et al. (2002)30, n = 5, m = −26.08, t = −1.08, d.f. = 3, P = 0.36; R2 = 0.28; linear regression of each data set examined independently), when examined altogether across the full range of host abundances studied, no significant slope was observed (microbial abundance log-transformed; n = 27, m = −0.11, t = −0.04, d.f. = 25, P = 0.97; R2 = 5.94 × 10−5; linear regression of pooled data).
Extended Data Figure 3 Decline in virus to microbe ratio (VMR) observed in incubations with elevated host density over time, contrasted with published values and viral decay.
a, Log-transformed VLP density in experimental incubations is plotted against microbial host density over time (dot size) with VMR indicated by dot colour. Data from Mission Bay (MB) and Palmyra (Pal) water with DOC added (+ DOC) or not (− DOC) is complemented by the nutrient-added ‘lytic’ system of Hennes et al. (1995)31 (H + Nutrients) as well as the ‘non-lytic’ dilutions (3%, 10%, 20%, and 30% final concentration seawater diluted by 0.02 μm filtered seawater) of Wilcox and Fuhrman (1994)3; WF 3% SW, WF 10% SW, WF 20% SW, WF 30% SW). n = 1 all incubations and published mean values. b, Significant viral decay was not observed in cell-free viral decay controls in incubation experiments (Palmyra: n = 4, m = 1.64 × 10−3, t = 1.48, d.f. = 2, P = 0.28; R2 = 0.52; Mission Bay: n = 6, m = 4.53 × 10−3, t = 1.87, d.f. = 4, P = 0.14; R2 = 0.47; linear regression with log-transformed viral density).
Extended Data Figure 4 Temperateness of viral communities increases with host density and viral functional composition change.
a, The relative composition of provirus-like reads, normalized by total sequences in each sample, increases with host density in viral metagenomes (host density log-transformed; n = 24 independent measures). The linear equations and line of best fit from robust regression and bootstrapped 95% and 90% confidence intervals (CIs) for the slope are shown. Goodness of fit metrics are inappropriate for robust regression and are omitted. b, Viromes clustered by functional similarity (crAss cross-assembly), showing higher host density Pacific viromes (*) grouped away from lower host density Atlantic viromes (†); site names coloured by host density.
Rights and permissions
About this article
Cite this article
Knowles, B., Silveira, C., Bailey, B. et al. Lytic to temperate switching of viral communities. Nature 531, 466–470 (2016). https://doi.org/10.1038/nature17193
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17193
This article is cited by
-
Biogeographic patterns and drivers of soil viromes
Nature Ecology & Evolution (2024)
-
Virus diversity and activity is driven by snowmelt and host dynamics in a high-altitude watershed soil ecosystem
Microbiome (2023)
-
Viral predation pressure on coral reefs
BMC Biology (2023)
-
Presence and role of viruses in anaerobic digestion of food waste under environmental variability
Microbiome (2023)
-
Soil viral diversity, ecology and climate change
Nature Reviews Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.