Abstract
Circuits in the cerebral cortex consist of thousands of neurons connected by millions of synapses. A precise understanding of these local networks requires relating circuit activity with the underlying network structure. For pyramidal cells in superficial mouse visual cortex (V1), a consensus is emerging that neurons with similar visual response properties excite each other1,2,3,4,5, but the anatomical basis of this recurrent synaptic network is unknown. Here we combined physiological imaging and large-scale electron microscopy to study an excitatory network in V1. We found that layer 2/3 neurons organized into subnetworks defined by anatomical connectivity, with more connections within than between groups. More specifically, we found that pyramidal neurons with similar orientation selectivity preferentially formed synapses with each other, despite the fact that axons and dendrites of all orientation selectivities pass near (<5 μm) each other with roughly equal probability. Therefore, we predict that mechanisms of functionally specific connectivity take place at the length scale of spines. Neurons with similar orientation tuning formed larger synapses, potentially enhancing the net effect of synaptic specificity. With the ability to study thousands of connections in a single circuit, functional connectomics is proving a powerful method to uncover the organizational logic of cortical networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Harris, K. D. & Mrsic-Flogel, T. D. Cortical connectivity and sensory coding. Nature 503, 51–58 (2013)
Ko, H. et al. Functional specificity of local synaptic connections in neocortical networks. Nature (2011)
Li, Y. T., Ibrahim, L. A., Liu, B. H., Zhang, L. I. & Tao, H. W. Linear transformation of thalamocortical input by intracortical excitation. Nature Neurosci. 16, 1324–1330 (2013)
Lien, A. D. & Scanziani, M. Tuned thalamic excitation is amplified by visual cortical circuits. Nature Neurosci. 16, 1315–1323 (2013)
Wertz, A. et al. Single-cell-initiated monosynaptic tracing reveals layer-specific cortical network modules. Science 349, 70–74 (2015)
Niell, C. M. & Stryker, M. P. Highly selective receptive fields in mouse visual cortex. J. Neurosci. 28, 7520–7536 (2008)
Kerlin, A. M., Andermann, M. L., Berezovskii, V. K. & Reid, R. C. Broadly tuned response properties of diverse inhibitory neuron subtypes in mouse visual cortex. Neuron 67, 858–871 (2010)
Bock, D. D. et al. Network anatomy and in vivo physiology of visual cortical neurons. Nature 471, 177–182 (2011)
Hofer, S. B. et al. Differential connectivity and response dynamics of excitatory and inhibitory neurons in visual cortex. Nature Neurosci. 14, 1045–1052 (2011)
Andermann, M. L., Kerlin, A. M., Roumis, D. K., Glickfeld, L. L. & Reid, R. C. Functional specialization of mouse higher visual cortical areas. Neuron 72, 1025–1039 (2011)
Marshel, J. H., Garrett, M. E., Nauhaus, I. & Callaway, E. M. Functional specialization of seven mouse visual cortical areas. Neuron 72, 1040–1054 (2011)
Bonin, V., Histed, M. H., Yurgenson, S. & Reid, R. C. Local diversity and fine-scale organization of receptive fields in mouse visual cortex. J. Neurosci. 31, 18506–18521 (2011)
Glickfeld, L. L., Andermann, M. L., Bonin, V. & Reid, R. C. Cortico-cortical projections in mouse visual cortex are functionally target specific. Nature Neurosci. (2013)
Cossell, L. et al. Functional organization of excitatory synaptic strength in primary visual cortex. Nature 518, 399–403 (2015)
Reinhold, K., Lien, A. D. & Scanziani, M. Distinct recurrent versus afferent dynamics in cortical visual processing. Nature Neurosci. 18, 1789–1797 (2015)
Saalfeld, S., Cardona, A., Hartenstein, V. & Tomancak, P. CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25, 1984–1986 (2009)
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008)
Song, S., Sjostrom, P. J., Reigl, M., Nelson, S. & Chklovskii, D. B. Highly nonrandom features of synaptic connectivity in local cortical circuits. PLoS Biol. 3, e68 (2005)
Yoshimura, Y., Dantzker, J. L. & Callaway, E. M. Excitatory cortical neurons form fine-scale functional networks. Nature 433, 868–873 (2005)
Bopp, R., Macarico da Costa, N., Kampa, B. M., Martin, K. A. & Roth, M. M. Pyramidal cells make specific connections onto smooth (GABAergic) neurons in mouse visual cortex. PLoS Biol. 12, e1001932 (2014)
Stepanyants, A., Hof, P. R. & Chklovskii, D. B. Geometry and structural plasticity of synaptic connectivity. Neuron 34, 275–288 (2002)
Kampa, B. M., Letzkus, J. J. & Stuart, G. J. Cortical feed-forward networks for binding different streams of sensory information. Nature Neurosci. 9, 1472–1473 (2006)
Reimann, M. W., King, J. G., Muller, E. B., Ramaswamy, S. & Markram, H. An algorithm to predict the connectome of neural microcircuits. Front. Comput. Neurosci. 9, 120 (2015)
Kasthuri, N. et al. Saturated reconstruction of a volume of neocortex. Cell 162, 648–661 (2015)
Bartol, T. M. et al. Nanoconnectomic upper bound on the variability of synaptic plasticity. Elife 4, (2015)
Harris, K. M. & Stevens, J. K. Dendritic spines of CA 1 pyramidal cells in the rat hippocampus: serial electron microscopy with reference to their biophysical characteristics. J. Neurosci. 9, 2982–2997 (1989)
Matsuzaki, M. et al. Dendritic spine geometry is critical for AMPA receptor expression in hippocampal CA1 pyramidal neurons. Nature Neurosci. 4, 1086–1092 (2001)
Nusser, Z. et al. Cell type and pathway dependence of synaptic AMPA receptor number and variability in the hippocampus. Neuron 21, 545–559 (1998)
Takumi, Y., Ramirez-Leon, V., Laake, P., Rinvik, E. & Ottersen, O. P. Different modes of expression of AMPA and NMDA receptors in hippocampal synapses. Nature Neurosci. 2, 618–624 (1999)
Reid, R. C. From functional architecture to functional connectomics. Neuron 75, 209–217 (2012)
Rubinov, M. & Sporns, O. Weight-conserving characterization of complex functional brain networks. Neuroimage 56, 2068–2079 (2011)
Andermann, M. L., Kerlin, A. M. & Reid, R. C. Chronic cellular imaging of mouse visual cortex during operant behavior and passive viewing. Front. Cell. Neurosci. 4, 3 (2010)
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nature Methods 6, 875–881 (2009)
Mank, M. et al. A genetically encoded calcium indicator for chronic in vivo two-photon imaging. Nature Methods 5, 805–811 (2008)
Andermann, M. L. et al. Chronic cellular imaging of entire cortical columns in awake mice using microprisms. Neuron 80, 900–913 (2013)
Peters, A. & Kara, D. A. The neuronal composition of area 17 of rat visual cortex. I. The pyramidal cells. J. Comp. Neurol. 234, 218–241 (1985)
Ferrer, I., Fabregues, I. & Condom, E. A Golgi study of the sixth layer of the cerebral cortex. I. The lissencephalic brain of Rodentia, Lagomorpha, Insectivora and Chiroptera. J. Anat. 145, 217–234 (1986)
Hirsch, J. A., Alonso, J. M. & Reid, R. C. Visually evoked calcium action potentials in cat striate cortex. Nature 378, 612–616 (1995)
Smith, S. L., Smith, I. T., Branco, T. & Hausser, M. Dendritic spikes enhance stimulus selectivity in cortical neurons in vivo. Nature 503, 115–120 (2013)
Markram, H., Helm, P. J. & Sakmann, B. Dendritic calcium transients evoked by single back-propagating action potentials in rat neocortical pyramidal neurons. J. Physiol. (Lond.) 485, 1–20 (1995)
Larkman, A. & Mason, A. Correlations between morphology and electrophysiology of pyramidal neurons in slices of rat visual cortex. I. Establishment of cell classes. J. Neurosci. 10, 1407–1414 (1990)
Peters, A., Palay, S. L. & Webster, H. d. The fine structure of the nervous system: neurons and their supporting cells 3rd edn (Oxford Univ. Press, 1991)
Harris, K. M. & Stevens, J. K. Dendritic spines of rat cerebellar Purkinje cells: serial electron microscopy with reference to their biophysical characteristics. J. Neurosci. 8, 4455–4469 (1988)
Thévenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998)
Rubinov, M. & Sporns, O. Complex network measures of brain connectivity: uses and interpretations. Neuroimage 52, 1059–1069 (2010)
Lancichinetti, A. & Fortunato, S. Consensus clustering in complex networks. Sci. Rep. 2, 336 (2012)
Maslov, S. & Sneppen, K. Specificity and stability in topology of protein networks. Science 296, 910–913 (2002)
Acknowledgements
We thank R. Arora, E. Ashbolt, L. Bailey, L. Beaton, P. Chopra, A. Coda, M. Driesbach, R. Fang, M. Fisher, A. Giuliano, H. Godtfredsen, A. Henry, E. Holtzman, P. Hughes, K. Joines, J. Larson, S. Maillet, K. Moody, E. Nguyen, A. Orban, V. Osuiji, K. Robertson, D. Sandman, S. Schwartz, E. Sczerzenie, N. Thatra, L. Thomas, B. Titus, R. Torres for tracing and reconstruction; E. Raviola for discussions and advice; H. Kim for technical support at the beginning of the study; A. Wetzel for help with alignment and stitching; A. Cardona and S. Saalfeld for the CATMAID project being openly available; and N. da Costa for launching the EM annotation program at the Allen Institute for Brain Science, with help and support from B. Youngstrom, R. Young, C. Dang and J. Phillips. We also thank D. Brittain, W. Gray Roncal, L. Thomas, S. Yurgenson, H. Elliot, and the HMS Image and Data Analysis Core for programming; E. Benecchi and the HMS EM Core Facility for technical support; P.J. Manavalan, K. Lillaney, and R. Burns, for helping make the data freely available; and S. Druckmann, Y. Park, C. Priebe, and J. Vogelstein for discussions on statistical analyses. We are indebted to M. Andermann, S. Chatterjee, N. da Costa, L. Glickfeld, C. Harwell, D. Hildebrand, M. Histed, S. Mihalas, L. Ostroff, E. Raviola, and J. T. Vogelstein for critical reading of various versions of the manuscript. This work was supported by NIH grants to RCR (R01 EY10115 and R01 NS075436); through resources provided by the NRBSC (P41 RR06009) and MMBioS (P41 GM103712); the HMS Vision Core Grant (P30 EY12196); and the AIBS. We thank the Allen Institute founders, P. G. Allen and J. Allen, for their vision, encouragement, and support. WCL was also supported by the Bertarelli Program in Translational Neuroscience and Neuroengineering, Edward R. and Anne G. Lefler Center, and Stanley and Theodora Feldberg Fund, and the NIH (R21 NS085320). VB was also supported by Neuro-Electronics Research Flanders. The project described was partially supported by the NIH by the above named awards. Its content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
W.-C.A.L. and V.B. performed the in vivo calcium imaging and analysed it. W.-C.A.L. processed the tissue for EM, sectioned the series, and aligned the block with the in vivo imaging. W.-C.A.L. and M.R. imaged it on the TEMCA. G.H. and W.-C.A.L. aligned the EM images into a volume. W.-C.A.L., M.R., K.G. annotated the EM dataset and W.-C.A.L. and K.G. supervised segmentation efforts. B.J.G. and W.-C.A.L. generated software for visualization and analysis. W.-C.A.L. performed quantitative analysis on the tracing. W.-C.A.L., V.B., and R.C.R. designed the experiment and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
The aligned EM dataset will be publicly accessible at http://neurodata.io/lee16.
Extended data figures and tables
Extended Data Figure 1 Sensory physiology is maintained over days.
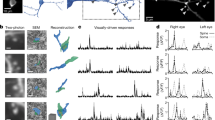
a, Visually evoked calcium responses are maintained across days. Three example neurons (rows) on the first, ninth, and twelfth day (columns) of in vivo two-photon calcium imaging. Three individual trial responses (black lines) of 8 directions, 3 spatial and 2 temporal frequencies (left column, day 1); or 16 directions, 2 spatial and 2 temporal frequencies (middle and right rows, days 9 and 12). Scale bars, 100% ΔF/F and 4 s. Standard deviation of preferred direction across days for each neuron is to the upper right of activity matrices. b, Neurons direction selectivity is stable over days. Cumulative distribution of standard deviation of peak preferred direction (4.1° ± 1.7°, median ± s.e.m.) across days for 25 neurons measured over multiple days.
Extended Data Figure 2 Deep layer apical responses are likely suprathreshold reflecting activity at the soma.
Example ΔF/F time courses from a deep layer apical dendrite optically sectioned in vivo across 6–8 planes. Note, activity is correlated across depth and relatively stable over days (Extended Data Fig. 1).
Extended Data Figure 3 Distribution of orientation and direction tuned cells.
a, b, Histograms of functionally characterized cells with measured peak orientation (a) and direction preference (b).
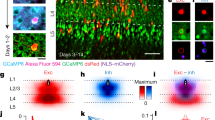
Extended Data Figure 4 In vivo to EM correspondence of neuronal targets.
Top row, volumetric projections of the aligned in vivo and EM imaged volumes. Physiology planes were acquired horizontally and EM sections cut frontally (coronally) from the brain. Interdigitated physiology planes are from 2 representative volumetrically scanned experiments stacked atop one another in space (Fig. 1c) so as to span from the border of L1 and L2 through the depth of L2/3. Scale bar, 100 μm. middle and bottom rows, re-sliced planes through the in vivo volume corresponding to EM sections. Arrowheads indicate matching cell bodies, and arrows deep layer (putative L5) apical dendrites. Small black dots mark the centres of cell bodies corresponding principally to nuclei where calcium indicator fluorescence is typically excluded. Scale bar, 50 μm.
Extended Data Figure 6 Network reconstruction.
3D rendering of dendrites and axons, cell bodies (large spheres), and synapses (small spheres) of, a, 50 functionally characterized neurons reconstructed in the EM volume. Cell bodies, dendrites, axons, and synapses colour coded by peak preferred stimulus orientation (colour key, bottom right). Axons are the thinnest processes, dendrites of L2/3 neurons are thicker, and deep layer apical dendrites are rendered with the largest calibre. Dendritic spines were traced only if they participated in connections between reconstructed neurons. b, Approximately 1,800 additional neuronal targets reconstructed in the EM volume (transparent grey). Input and output synapses are coloured cyan and red respectively when orientation selectivity was not known. Bounding box matches region in Fig. 1f. Scale bar, 150 μm.
Extended Data Figure 7 Network modularity is significantly non-random.
a, Connectivity matrix of 201 excitatory neuronal targets in our network reconstruction with multiple synaptic partners (that is, degree ≥ 2, no leaf nodes, same as Fig. 1e). Colour represents the number of synapses (colour key, c, right) between pre- and post-synaptic neurons (same neuron order on both ordinate and abscissa). Subnetworks of interconnected neurons (white boxes) detected using a consensus method of Louvain clustering17,31. b, Modularity (Q) of the reconstructed network is significantly greater than null models with degree, weight, and strength preserved. Histograms of the modularity values for the reconstructed network (dark grey, Qmean = 0.55 ± 0.003, mean ± s.d., computed 1,000 times) is significantly greater than for the Qmean of shuffled null models (light grey, Qmean = 0.50 ± 0.009, mean ± s.d., P ≈ 0, permutation test, kshuffles = 1,000). c, Example of the shuffled connectivity matrix with a Q closest to the mean of the shuffled distribution with clustering (white boxes) computed as in a. d, Null models are well-shuffled, while approximating connection input and output strengths. Histograms of correlation coefficients between the reconstructed network and the null models’ in (blue: 0.92 ± 0.02, mean ± s.d.) and out (red: 96 ± 0.01, mean ± s.d.) strength and connectivity matrix (grey: 9.1 × 10−4 ± 0.01, mean ± s.d.). e, Occurrences of three neuron connectivity motifs found in the reconstructed network between excitatory neuronal targets.
Extended Data Figure 8 Cell bodies are functionally intermingled.
a, b, Differences in peak orientation (a) and direction preference (b) between neuron pairs plotted against the distance between their cell bodies. Uniform distributions of functional versus spatial distance suggest a salt and pepper intermingling of neuronal cell bodies across functional properties.
Extended Data Figure 9 Axons and dendrites are functionally intermingled at shorter length scales.
Uniform functional diversity and prediction of connectivity at finer length scales suggest a salt-and-pepper intermingling of axons and dendrites. a–d, Same as Fig. 2c–f for s = 1 μm. Significance tests: a, P ≈ 0, permutation test, nconnected pairs = 29, nunconnected pairs = 1,951; b, Between connected (red line) and unconnected pairs (blue line, P < 0.05, permutation test, nconnected pairs = 29, nunconnected pairs = 1,951) or a model distribution based on potential synapse length (black line, P < 0.01, permutation test, nconnected pairs = 29, nunconnected pairs = 1,951). Shaded regions, a, b, and error bars, d, represent bootstrapped standard error.
Extended Data Figure 10 Connectivity is not predicted by residual signal correlation after removal of orientation preference.
a–b, Example activity (ΔF/F individual trial time courses) for connected neurons from experiments varying direction, spatial and temporal frequencies of grating stimuli. a, Presynaptic cell (top row) and two of its postsynaptic partners’ (middle and bottom rows) for 3 spatial and 2 temporal frequencies and one orientation (orientation tuning was virtually identical). b, Presynaptic cell (top) and a postsynaptic deep layer apical dendrite’s (bottom) responses to 2 spatial and 2 temporal frequencies, and 2 directions, (again, orientation tuning was virtually identical). Grey window delineates time of stimulus presentation. Scale bars, 100% ΔF/F and 4 s. c, Cumulative distribution of signal correlations from simultaneously measured cells was significantly greater between connected than unconnected pairs (P < 0.01, permutation test, nconnected pairs = 10, nunconnected pairs = 426) or a model distribution based on potential synaptic connectivity (P < 0.05, permutation test). d, After averaging over orientations, the cumulative distribution of signal correlations was similar between connected and unconnected pairs (P > 0.14, permutation test, nconnected pairs = 10, nunconnected pairs = 426) and a model distribution based on potential synaptic connectivity (P > 0.25, permutation test, nconnected pairs = 10, nunconnected pairs = 426). Shaded regions, c, d, represent bootstrapped standard error.
Supplementary information
Supplementary Data
This file contains Supplementary Data 1-3. (PDF 926 kb)
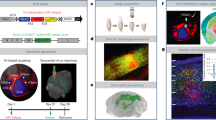
In vivo fluorescence to EM correspondence
Fluorescence in vivo two-photon microscopy data 3-D aligned with the EM volume. To visualize three imaging datasets (in vivo anatomy: fluorescence volume, in vivo physiology: fluorescence planes; and EM) first, the volume is flown-through, oriented with the EM sections (coronally, relative to the brain). Then the volume is rotated so that the perspective is above the surface and horizontal relative to the brain. Next, descending into the brain, note the correspondence of GCaMP3 (green) and functionally colored cell bodies with the nuclei in EM, the blood vessels (red) with clear vasculature in EM, and track apical dendrites in vivo (green radially oriented processes) in EM (large caliber cylindrical dendrites with spines). (MOV 20389 kb)
EM volume overview
The video shows a fly-through of the aligned EM series. Please see Author Information for directions to the publicly accessible high-resolution aligned dataset. (MOV 18428 kb)
Higher resolution EM core
The video shows fly-through of a cropped (16.4 x 16.4 µm) region traversing 400 of the aligned EM sections in the functionally imaged volume. (MOV 27726 kb)
Rights and permissions
About this article
Cite this article
Lee, WC., Bonin, V., Reed, M. et al. Anatomy and function of an excitatory network in the visual cortex. Nature 532, 370–374 (2016). https://doi.org/10.1038/nature17192
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17192
This article is cited by
-
The logic of recurrent circuits in the primary visual cortex
Nature Neuroscience (2024)
-
RoboEM: automated 3D flight tracing for synaptic-resolution connectomics
Nature Methods (2024)
-
Intracellular magnesium optimizes transmission efficiency and plasticity of hippocampal synapses by reconfiguring their connectivity
Nature Communications (2024)
-
Synaptic wiring motifs in posterior parietal cortex support decision-making
Nature (2024)
-
Local shape descriptors for neuron segmentation
Nature Methods (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.