Abstract
Contractile tails are composed of an inner tube wrapped by an outer sheath assembled in an extended, metastable conformation that stores mechanical energy necessary for its contraction. Contraction is used to propel the rigid inner tube towards target cells for DNA or toxin delivery. Although recent studies have revealed the structure of the contractile sheath of the type VI secretion system, the mechanisms by which its polymerization is controlled and coordinated with the assembly of the inner tube remain unknown. Here we show that the starfish-like TssA dodecameric complex interacts with tube and sheath components. Fluorescence microscopy experiments in enteroaggregative Escherichia coli reveal that TssA binds first to the type VI secretion system membrane core complex and then initiates tail polymerization. TssA remains at the tip of the growing structure and incorporates new tube and sheath blocks. On the basis of these results, we propose that TssA primes and coordinates tail tube and sheath biogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
Coordinates and structure factors have been deposited in the Protein Data Bank under accession numbers 4YO3 and 4YO5 for TssANt2 and TssACt respectively. Electron microscopy map for full-length TssA has been deposited in the Electron Microscopy Data Bank under accession code EMD-3282.
References
Leiman, P. G. & Shneider, M. M. Contractile tail machines of bacteriophages. Adv. Exp. Med. Biol. 726, 93–114 (2012)
Ge, P. et al. Atomic structures of a bactericidal contractile nanotube in its pre- and postcontraction states. Nature Struct. Mol. Biol. 22, 377–382 (2015)
Bönemann, G., Pietrosiuk, A. & Mogk, A. Tubules and donuts: a type VI secretion story. Mol. Microbiol. 76, 815–821 (2010)
Yang, G., Dowling, A. J., Gerike, U. ffrench-Constant, R. H. & Waterfield, N. R. Photorhabdus virulence cassettes confer injectable insecticidal activity against the wax moth. J. Bacteriol. 188, 2254–2261 (2006)
Shikuma, N. J. et al. Marine tubeworm metamorphosis induced by arrays of bacterial phage tail-like structures. Science 343, 529–533 (2014)
Heymann, J. B. et al. Three-dimensional structure of the toxin-delivery particle antifeeding prophage of Serratia entomophila . J. Biol. Chem. 288, 25276–25284 (2013)
Kube, S. & Wendler, P. Structural comparison of contractile nanomachines. AIMS Biophysics. 2, 88–115 (2015)
Leiman, P. G. et al. Morphogenesis of the T4 tail and tail fibers. Virol. J. 7, 355 (2010)
Ferguson, P. L. & Coombs, D. H. Pulse-chase analysis of the in vivo assembly of the bacteriophage T4 tail. J. Mol. Biol. 297, 99–117 (2000)
Rybakova, D. et al. Role of antifeeding prophage (Afp) protein Afp16 in terminating the length of the Afp tailocin and stabilizing its sheath. Mol. Microbiol. 89, 702–714 (2013)
Aschtgen, M. S., Gavioli, M., Dessen, A., Lloubès, R. & Cascales, E. The SciZ protein anchors the enteroaggregative Escherichia coli Type VI secretion system to the cell wall. Mol. Microbiol. 75, 886–899 (2010)
Felisberto-Rodrigues, C. et al. Towards a structural comprehension of bacterial type VI secretion systems: characterization of the TssJ-TssM complex of an Escherichia coli pathovar. PLoS Pathog. 7, e1002386 (2011)
Durand, E. et al. Biogenesis and structure of the Type VI secretion membrane core complex. Nature 523, 555–560 (2015)
Zoued, A. et al. Architecture and assembly of the Type VI secretion system. Biochim. Biophys. Acta 1843, 1664–1673 (2014)
Brunet, Y. R., Hénin, J., Celia, H. & Cascales, E. Type VI secretion and bacteriophage tail tubes share a common assembly pathway. EMBO Rep. 15, 315–321 (2014)
Leiman, P. G. et al. Type VI secretion apparatus and phage tail-associated protein complexes share a common evolutionary origin. Proc. Natl Acad. Sci. USA 106, 4154–4159 (2009)
Basler, M., Pilhofer, M., Henderson, G. P., Jensen, G. J. & Mekalanos, J. J. Type VI secretion requires a dynamic contractile phage tail-like structure. Nature 483, 182–186 (2012)
English, G., Byron, O., Cianfanelli, F. R., Prescott, A. R. & Coulthurst, S. J. Biochemical analysis of TssK, a core component of the bacterial Type VI secretion system, reveals distinct oligomeric states of TssK and identifies a TssK-TssFG subcomplex. Biochem. J. 461, 291–304 (2014)
Brunet, Y. R., Zoued, A., Boyer, F., Douzi, B. & Cascales, E. The Type VI secretion TssEFGK-VgrG phage-like baseplate is recruited to the TssJLM membrane complex via multiple contacts and serves as assembly platform for tail tube/sheath polymerization. PLoS Genet. 11, e1005545 (2015)
Basler, M., Ho, B. T. & Mekalanos, J. J. Tit-for-tat: type VI secretion system counterattack during bacterial cell-cell interactions. Cell 152, 884–894 (2013)
Brunet, Y. R., Espinosa, L., Harchouni, S., Mignot, T. & Cascales, E. Imaging type VI secretion-mediated bacterial killing. Cell Rep. 3, 36–41 (2013)
Kube, S. et al. Structure of the VipA/B type VI secretion complex suggests a contraction-state-specific recycling mechanism. Cell Rep. 8, 20–30 (2014)
Kudryashev, M. et al. Structure of the type VI secretion system contractile sheath. Cell 160, 952–962 (2015)
Kapitein, N. et al. ClpV recycles VipA/VipB tubules and prevents non-productive tubule formation to ensure efficient type VI protein secretion. Mol. Microbiol. 87, 1013–1028 (2013)
Zoued, A. et al. TssK is a trimeric cytoplasmic protein interacting with components of both phage-like and membrane anchoring complexes of the type VI secretion system. J. Biol. Chem. 288, 27031–27041 (2013)
King, J. Assembly of the tail of bacteriophage T4. J. Mol. Biol. 32, 231–262 (1968)
Vegge, C. S. et al. Structural characterization and assembly of the distal tail structure of the temperate lactococcal bacteriophage TP901–1. J. Bacteriol. 187, 4187–4197 (2005)
Pell, L. G. et al. The X-ray crystal structure of the phage λ tail terminator protein reveals the biologically relevant hexameric ring structure and demonstrates a conserved mechanism of tail termination among diverse long-tailed phages. J. Mol. Biol. 389, 938–951 (2009)
Fokine, A. et al. The molecular architecture of the bacteriophage T4 neck. J. Mol. Biol. 425, 1731–1744 (2013)
Yonekura, K. et al. The bacterial flagellar cap as the rotary promoter of flagellin self-assembly. Science 290, 2148–2152 (2000)
Brunet, Y. R., Bernard, C. S., Gavioli, M., Lloubès, R. & Cascales, E. An epigenetic switch involving overlapping fur and DNA methylation optimizes expression of a type VI secretion gene cluster. PLoS Genet. 7, e1002205 (2011)
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000)
Chaveroche, M. K., Ghigo, J. M. & d'Enfert, C. A rapid method for efficient gene replacement in the filamentous fungus Aspergillus nidulans . Nucleic Acids Res. 28, E97 (2000)
van den Ent, F. & Löwe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J. Biochem. Biophys. Methods 67, 67–74 (2006)
Karimova, G., Pidoux, J., Ullmann, A. & Ladant, D. A bacterial two-hybrid system based on a reconstituted signal transduction pathway. Proc. Natl Acad. Sci. USA 95, 5752–5756 (1998)
Battesti, A. & Bouveret, E. The bacterial two-hybrid system based on adenylate cyclase reconstitution in Escherichia coli . Methods 58, 325–334 (2012)
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nature Methods 9, 676–682 (2012)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. & Svergun, D. I. PRIMUS: a Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003)
Konarev, P. V., Petoukhov, M. V., Volkov, V. V. & Svergun, D. I. ATSAS 2.1, a program package for small-angle scattering data analysis. J. Appl. Crystallogr. 39, 277–286 (2006)
Guinier, A. La diffraction des rayons X aux très petits angles; application à l'étude de phénomènes ultramicroscopiques. Ann. Phys. (Paris) 12, 161–237 (1939)
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992)
Franke, D. & Svergun, D. I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J. Appl. Crystallogr. 42, 342–346 (2009)
Volkov, V. V. & Svergun, D. I. Uniqueness of ab initio shape determination in small-angle scattering. J. Appl. Crystallogr. 36, 860–864 (2003)
Kozin, M. B. & Svergun, D. I. Automated matching of high- and low-resolution structural models. J. Appl. Crystallogr. 34, 33–41 (2001)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D 60, 2210–2221 (2004)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
DeLano, W.L. The PyMOL Molecular Graphics System, Version 1.8 Schrödinger, LLC
Pettersen, E. F. et al. UCSF Chimera – a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Gerc, A. J. et al. Visualization of the Serratia Type VI secretion system reveals unprovoked attacks and dynamic assembly. Cell Rep. 12, 2131–2142 (2015)
Douzi, B. et al. Crystal structure and self-interaction of the type VI secretion tail-tube protein from enteroaggregative Escherichia coli . PLoS ONE 9, e86918 (2014)
Acknowledgements
This work was funded by the Centre National de la Recherche Scientifique, the Aix-Marseille Université, and grants from the Agence Nationale de la Recherche to E.C. (ANR-10-JCJC-1303-03), to E.C. and C.C. (ANR-14-CE14-0006-02) and from the Fondation pour la Recherche Médicale to C.C. (FRM DEQ2011-0421282) and supported by the French Infrastructure for Integrated Structural Biology (FRISBI, ANR-10-INSB-05-01). A.Z. and Y.R.B. were supported by doctoral fellowships from the French Ministry of Research. A.Z. and E.D. were supported by end-of-thesis (FDT20140931060) and post-doctoral (SPF20101221116) fellowships from the Fondation pour la Recherche Médicale, respectively. We gratefully acknowledge the Soleil synchrotron radiation facility for beamtime allocation. We thank R. Lloubès, J. Sturgis and A. Galinier for constant support, the members of the Cascales, Cambillau, Lloubès, Sturgis and Bouveret research groups for helpful discussions, E. Bouveret for providing vectors, protocols and advice for the bacterial two-hybrid assay, C. Bebeacua for preliminary electron microscopy analyses, L. Espinosa for help regarding statistical analyses, R. Lebrun and S. Lignon (proteomic platform, IMM) for mass spectrometry analyses, Y. Cully for the Supplementary Video, O. Uderso, I. Bringer and A. Brun for technical assistance, and J. D. Barras-Elatable for encouragement.
Author information
Authors and Affiliations
Contributions
A.Z., E.D., C.C. and E.C. designed and conceived the experiments. C.C. and E.C supervised the execution of the experiments. A.Z., E.D., Y.R.B., S.S., B.D. and M.G. performed the experiments. A.Z. performed the in vivo experiments (BACTH, fluorescence microscopy) with the help of Y.R.B., M.G., N.F., L.J. and T.M. E.D. performed the in vitro experiments (protein purification and characterization, SAXS, electron microscopy and X-ray analyses) with the help of S.S., P.L. and R.F. B.D. performed the SPR experiments. P.L., R.F., T.M., C.C. and E.C. provided tools. E.C. wrote the paper with contributions of A.Z., E.D. and C.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Hcp and TssC interact with TssA, a cytoplasmic protein required for sheath assembly and Hcp release.
a, Schematic representation of the architecture of the bacterial type VI secretion system. The scheme highlights the membrane complex anchoring the tail structure composed of the assembly baseplate, the spike, the tube and the sheath (cyto, cytoplasm; IM, inner membrane; PG, peptidoglycan layer; OM, outer membrane). b, Bacterial two-hybrid assay. BTH101 reporter cells producing the indicated proteins or domains (TssLc, cytoplasmic domain of the TssL protein; TssMc and TssMp, cytoplasmic and periplasmic domain of the TssM protein respectively) fused to the T18 or T25 domain of the Bordetella adenylate cyclase were spotted on plates supplemented with IPTG and the chromogenic substrate X-Gal. Interaction between the two fusion proteins is attested by the dark blue colour of the colony. The TolB–Pal interaction serves as a positive control. c, The absence of TssA prevents T6SS sheath dynamics. Fluorescence microscopy time-lapse recordings showing sheath dynamics using the chromosomally-encoded tssB-mCherry fusion in wild-type (WT) (tssB–mCherry pBAD33), ∆tssA (∆tssA tssB–mCherry pBAD33) and complemented ∆tssA (∆tssA tssB–mCherry pBAD33-TssAVSV-G) cells. Individual images were taken every 30 s. Assembly and contraction/disassembly events are indicated above the time-lapse images. The scale bars are 1 μm. d, The absence of TssA prevents Hcp release. Hcp release was assessed by separating whole cells (C) and supernatant (SN) fractions from 17-2 (WT), ∆tssA (∆tssA pBAD33, tssA) and complemented ∆tssA (∆tssA pBAD33-TssAVSV-G, tssAWT) cells producing Flag-epitope-tagged Hcp. A total of 1 × 108 cells and the TCA-precipitated material from the supernatant of 5 × 108 cells were analysed by western blot using anti-Flag monoclonal antibody (lower panel) and anti-TolB polyclonal antibodies as a lysis control (upper panel). The molecular weight markers (in kDa) are indicated on the left. The uncropped scans of the western blots are provided in the Supplementary Figure. e, TssA co-fractionates with cytoplasmic and membrane proteins. A fractionation procedure was applied to EAEC ∆tssA cells producing Flag-tagged TssA. Whole cells (T) were fractionated to isolate the supernatant (SN), periplasmic (P), cytoplasmic (C) and total membrane (M) fractions. Extracts from 109 (T) or 2 × 109 (SN, P, C, M) cells were separated by SDS–PAGE and immunodetected with anti-Flag monoclonal (TssA), anti-EF-Tu (cytoplasmic marker) and TolB (periplasmic marker) antibodies. The molecular weight markers (in kDa) are indicated on the left. The uncropped scans of the western blots are provided in Supplementary Figure 1.
Extended Data Figure 2 Purification and negative-stain electron microscopy analyses of the TssJLM–TssA complex.
a, TssA interacts with the TssJM complex. The total solubilized membrane extract (T) of 4 × 109 cells producing the indicated proteins was subjected to affinity chromatography using streptactin resin. Bound proteins (E) were separated by SDS–PAGE and immunodetected with anti-Flag (TssA and TssL), anti-strep tag (TssJ) and anti-5×His (TssM) monoclonal antibodies. The molecular weight markers are indicated on the left. The uncropped scans of the western blots are provided in Supplementary Figure 1. b, Superose 6 10/300 gel-filtration profile of the purified TssJLM–TssA complex. The asymmetry of the peak probably reflects the co-purification of different complexes or the dissociation of TssA from the TssJLM complex. c, Examples of representative raw particles observed for the purified TssJLM–TssA complex sample using negative-stain electron microscopy. A typical TssJLM complex is shown in red (number of particles observed n = 240) whereas a TssA-bound TssJLM complex is shown in white (n = 95). Scale bar is 10 nm. d, Magnification of the two complexes shown in c. Scale bars, 10 nm.
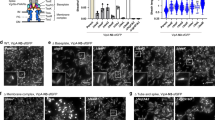
Extended Data Figure 3 TssA localization and dynamics.
a, Mean square displacement (MSD; in arbitrary units (a.u.)) of a representative sfGFP–TssA clusters in a wild-type strain (red line) or its ∆tssBC isogenic derivative (black line) were measured by sub-pixel tracking of fluorescent foci and plotted over time (in sec). b, Kymographic analysis reporting representative sfGFP–TssA (green) and TssB–mCherry (red) positions within the cell as a function of time. c, Representative fluorescence lifetime imaging microscopy (FLIM) of sfGFP–TssA clusters in the sfGFP–TssA/TssB–mCherry strain. A membrane-associated sfGFP–TssA cluster was chosen to define the bleached area (red circle). The laser (488 nm) was set to maximum power and focused for 3 s to ensure complete bleaching of the GFP diffusible pool. Images were taken every 30 s to follow recovery dynamics. The scale bar is 1 μm. d, Quantification of sfGFP–TssA fluorescence dynamics over time after bleaching. The dynamics of fluorescence intensity is shown over time for n = 10 independent sfGFP–TssA foci after FLIM (blue line). The fluorescence intensity of the bleached focus was also followed over time (FRAP, red line). As a control for laser focusing and intensity, membrane-associated clusters were systematically bleached in these experiments and showed no recovery suggesting the total intracellular sfGFP–TssA has been bleached by the laser. e, Representative fluorescence microscopy time-lapse recordings of the indicated ∆tssK, ∆tssE, ∆tssF, ∆tssG or ∆hcp cells producing sfGFP–TssA. Individual images were taken every 30 s. Red arrowheads indicate the localizations of TssA foci. The scale bar is 1 μm. f, Representative large fields of fluorescence microscopy analyses showing localization of sfGFP–TssA in the indicated strains. The scale bars are 1 μm. g, Box-and-whisker plots of the measured number of sfGFP–TssA foci per cell for each indicated strain. The lower and upper boundaries of the boxes correspond to the 25% and 75% percentiles respectively. The black bold horizontal bar represents the median values for each strain and the whiskers represent the 10% and 90% percentiles. Outliers are shown as open circles. A Student’s t-test was used to report significant differences (ns, not significant; ***P < 0.0001). The number of cells studied per strain (n) is indicated on top. h, Statistical analyses of sfGFP–TssA dynamics. sfGFP–TssA dynamics were categorized as ‘fixed’, ‘mobile with unidirectional trajectory’ and ‘mobile with random dynamics’ and the number of sfGFP–TssA (n, on top) foci in each category is represented as a percentage for each indicated strain. Kymographs for the two first categories are shown at the bottom of the panel. i, Schematic representation of the assembly pathway of the T6SS based on this study and available data13,15,19,24,25,55. The biogenesis starts with the initial positioning of the TssJ outer membrane lipoprotein and the sequential recruitment of the indicated subunits (from left to right). The recruitment of TssA is dependent on TssM, and that of TssK is dependent on both TssL and TssA. The exact positions of VgrG and TssE (blue) in the pathway are not known but these two subunits are not required for TssA recruitment but necessary for Hcp and TssBC polymerization.
Extended Data Figure 4 TssA interacts with tail and baseplate components.
a, TssA interaction network identified by bacterial two-hybrid analysis (see legend to Extended Data Fig. 1b). b–e, Surface plasmon resonance interaction study of TssA with its partners identified by BACTH. Sensorgrams (variation of plasmon resonance in arbitrary units (∆RU) as a function of reaction time (in sec)) were recorded upon injection of the purified native TssA protein (concentrations of 3.125 (dark grey), 6.25, 12.5, 25, 50 and 100 (light grey) μM) on HC200m chips coated with the purified N-terminal domain of VgrG (b), purified TssE (c), Hcp (d) or TssBC complex (e) (upper panels). The graph reporting ∆RU as a function of the TssA concentration (lower panel) was used to estimate the indicated apparent dissociation constants (Kd). Off-rates (percentage of dissociation 400 s after ligand injection) are indicated.
Extended Data Figure 5 TssA oligomerization and SAXS and EM structural models.
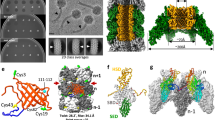
a, 10 μg of purified TssA were analysed by SDS–PAGE and Coomassie blue staining. The molecular weight markers (in kDa) are indicated on the left, and TssA and its theoretical size are indicated on the right. b, Superose 6 10/300 gel filtration profile of purified TssA (black line) and protein markers of known size (coloured lines). c, MALS/QELS/UV/RI analysis of purified TssA. The molecular mass of the TssA complex is indicated. d–j, Low-resolution SAXS model of TssA. d, Experimental scattering data calculated from an ab initio model of TssA. The square root χ value of the ‘best representative’ model is indicated. e, Representation of the Guinier plot calculated from the experimental curve. f, Pair distance distribution. g, Kratky plot representative of a multi-domain protein with flexible linkers. h–j, SAXS envelope of the ‘best representative’ model of TssA, with top (h), side (i) and tilted (j) views. The scale bar is 100 Å. k–r, Low-resolution EM model of TssA. k, Representative micrograph of the data set used for image processing. White circles indicate isolated TssA dodecamers. l. Representative selected TssA particles. m, n, Gallery of representative top (m) and side (n) class averages generated after reference-free 2D classification using Relion39. o, Fourier shell correlation (FSC) curve of the TssA reconstruction. The gold standard FSC curve was calculated in Relion39 using the masked reconstruction of TssA. p–r, Top (p), side (q) and tilted (r) views of the three-dimensional reconstruction model of the TssA dodecamer obtained by electron microscopy (accession number: EMD-3282). The scale bar is 50 Å. Whereas the SAXS model allows to better visualize the arm length compared to the EM reconstruction, its low resolution impairs the visual separation of the dimeric arms.
Extended Data Figure 6 Identification, oligomerization and interaction analysis of TssA domains.
a, Limited proteolysis of purified TssA. The purified full-length TssA protein (first lane) was submitted to proteinase K limited proteolysis for the time indicated on top of each lane and analysed by SDS–PAGE and Coomassie blue staining. Stable fragments are indicated on the right with their boundaries (numbers identified in the sequence in b) and the corresponding fragment. The uncropped scan of the Coomassie blue stained gel is provided in Supplementary Figure 1. b, TssA protein sequence. The localization of the boundaries of the stable fragments obtained after Proteinase K limited proteolysis and electrospray mass spectrometry analyses are arrowed. The secondary structures observed in the crystal structures (Fig. 2a and Extended Data Fig. 6f, g) are indicated on top of the corresponding sequence. c, Bacterial two-hybrid analysis of TssANt and TssACt interactions (see legend to Extended Data Fig. 1b). d, e, MALS/QELS/UV/RI analysis of the purified TssANt (d) and TssACt (e) fragments. f, g, X-ray structure of the TssANt2 domain (PDB: 4YO3). The rainbow coloured ribbon representation of the TssANt monomer is shown (f, consecutive α-helices numbered α1 to α7) whereas the dimeric structure (g) highlights the helices at the interface (α1, α2 and α6). h, The TssA central core interacts with Hcp and VgrG whereas the TssA arms interact with TssE and TssC. Bacterial two-hybrid analysis of TssANt and TssACt interactions (see legend to Extended Data Fig. 1b). i, j, Surface plasmon resonance interaction study of the purified TssACt (i) or TssANt (j) domains with the Hcp protein (i) or the TssBC complex (j). Sensorgrams (variation of plasmon resonance in arbitrary unit (∆RU) as a function of reaction time (in seconds)) were recorded upon injection of the purified TssA C-terminal (i) or TssANt (j) domains (concentrations of 3.125 (dark grey), 6.25, 12.5, 25, 50 and 100 (light grey) μM) on HC200m chips coated with the purified Hcp protein (i) or the TssBC complex (j) (upper panels). The graph reporting ∆RU as a function of the TssA domain concentration (lower panel) was used to estimate the indicated apparent dissociation constants (Kd).
Extended Data Figure 7 Comparison of the SAXS, EM and X-ray structures of TssA.
a, Schematic representation and colour code of the constructs used for SAXS (grey), electron microscopy (light blue) and X-ray (TssANt, dark blue; TssACt, red) analyses. The epitopes and theoretical molecular masses of the domains are indicated. TRX, thioredoxine; N, N terminus; C, C terminus. b, Fit between the experimental data (green dots) and the calculated scattering curves for TssANt2 and TssACt generated by CRYSOL (red line). c–f, SAXS/X-ray comparison. Top (c), side (d) and bottom (f) views of the fitting of TssANt2 (blue ribbon) and TssACt (red ribbon) X-ray structures into the TssA SAXS envelope (transparent grey surface). Scale bars are 10 nm. e, Magnification of a cut-away section of the fitting shown in d. Scale bar is 5 nm. g–i, SAXS/EM/X-ray comparison. Top (g) and side (h) views of the superimposition of SAXS (grey surface), EM (transparent light-blue surface) and X-ray structures of TssA. Scale bars are 10 nm. i, Magnification of a cut-away section of the superimposition shown in h. j–n, EM/X-ray comparison. Top (j), side (k) and bottom (l) views of the fitting of TssANt2 (blue ribbon) and TssACt (red ribbon) X-ray structures into the TssA EM envelope (transparent grey surface). Scale bars are 10 nm. m, n, Magnifications of the top and bottom views of the docking of the TssA domain X-ray structures into the TssA EM map highlighting the interface between the TssA central core (TssACt, red ribbon) and arms (TssANt2, blue ribbon). The C-terminal helix of TssANt2 (ends at position 377) and N-terminal helix of TssACt (starts at position 395) are shown in yellow. o, Top view of the fitting of the X-ray structure of EAEC Hcp (green ribbon, PDB 4HKH56) into the TssA SAXS envelope (grey surface). The scale bar is 10 nm.
Extended Data Figure 8 Models of tail-sheath TssA complexes and comparison between the bacteriophage T4 gp15 and T6SS TssA subunits.
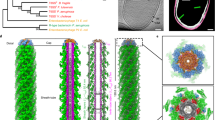
a, b, Surface (a) and cross-section (b) views of the complex of TssA (EM map, blue) with the extended tail sheath model (the last four rows shown in different colours). In the cut-away section, four stacks of Hcp rings are visible. As shown by bacterial two-hybrid and SPR analyses, Hcp contacts the TssACt central core whereas TssBC contacts the TssANt arms. The TssA arms fit between the TssBC monomers of the last row. c, Surface view of the complex of TssA (EM map, blue) with the contracted tail sheath model (the last four rows shown in different colours), highlighting the loose packing between TssA and the tail sheath in this conformation, suggesting that TssA might dissociate after sheath contraction. d–f, Comparison between the bacteriophage T4 gp15 and T6SS TssA subunits. d, e, Schematic representations of the bacteriophage T4 tail distal end comprising the gp19 tube (grey) and gp18 sheath (blue) proteins and the gp3 (green) and gp15 (red) neck proteins (d) and the T6SS tail distal end comprising the Hcp tube (grey) and TssBC sheath (blue) proteins and the TssA dodecamer (red) (e). The possibility that a functional homologue of bacteriophage T4 gp3 exists is shown by the question mark. f, Fitting of the model of the gp15 structure in complex with the last row of the gp18 sheath (in purple)29 in the TssA SAXS envelope (grey surface).
Supplementary information
Supplementary Table
This file shows the strains, plasmids and oligonucleotides used in this study. (PDF 294 kb)
Supplementary Figure
Uncropped scans of western blots and gels presented in Extended Data Fig. 1d, 1e, 2a & 6a. Molecular weight markers (in kDa) are indicated. (PDF 617 kb)
Dynamic representation of T6SS assembly highlighting the TssA mode of action
The TssA protein (red ring) bound to the membrane and baseplate complexes (TssJLM shown in yellow/orange; VgrG spike, purple; baseplate, orange) initiates tail polymerization. Polymerization is pursued by the TssA-dependent sequential and processive recruitement of Hcp rings (green rings) and TssBC strands (blue rings). Once assembled, the sheath contracts and propels the Hcp/VgrG needle to the cell exterior. (MOV 1732 kb)
Rights and permissions
About this article
Cite this article
Zoued, A., Durand, E., Brunet, Y. et al. Priming and polymerization of a bacterial contractile tail structure. Nature 531, 59–63 (2016). https://doi.org/10.1038/nature17182
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17182
This article is cited by
-
Bacterial type VI secretion system (T6SS): an evolved molecular weapon with diverse functionality
Biotechnology Letters (2023)
-
Abiotic factors modulate interspecies competition mediated by the type VI secretion system effectors in Vibrio cholerae
The ISME Journal (2022)
-
Pseudomonas fluorescens F113 type VI secretion systems mediate bacterial killing and adaption to the rhizosphere microbiome
Scientific Reports (2021)
-
In silico comparative analysis of Aeromonas Type VI Secretion System
Brazilian Journal of Microbiology (2021)
-
TssA–TssM–TagA interaction modulates type VI secretion system sheath-tube assembly in Vibrio cholerae
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.