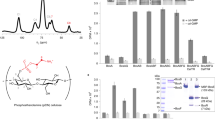
Abstract
Many biopolymers, including polysaccharides, must be translocated across at least one membrane to reach their site of biological function. Cellulose is a linear glucose polymer synthesized and secreted by a membrane-integrated cellulose synthase. Here, in crystallo enzymology with the catalytically active bacterial cellulose synthase BcsA–BcsB complex reveals structural snapshots of a complete cellulose biosynthesis cycle, from substrate binding to polymer translocation. Substrate- and product-bound structures of BcsA provide the basis for substrate recognition and demonstrate the stepwise elongation of cellulose. Furthermore, the structural snapshots show that BcsA translocates cellulose via a ratcheting mechanism involving a ‘finger helix’ that contacts the polymer’s terminal glucose. Cooperating with BcsA’s gating loop, the finger helix moves ‘up’ and ‘down’ in response to substrate binding and polymer elongation, respectively, thereby pushing the elongated polymer into BcsA’s transmembrane channel. This mechanism is validated experimentally by tethering BcsA’s finger helix, which inhibits polymer translocation but not elongation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Keegstra, K. Plant cell walls. Plant Physiol. 154, 483–486 (2010)
Serra, D. O., Richter, A. M. & Hengge, R. Cellulose as an architectural element in spatially structured Escherichia coli biofilms. J. Bacteriol. 195, 5540–5554 (2013)
Römling, U. Molecular biology of cellulose production in bacteria. Res. Microbiol. 153, 205–212 (2002)
Kimura, S., Ohshima, C., Hirose, E., Nishikawa, J. & Itoh, T. Cellulose in the house of the appendicularian Oikopleura rufescens. Protoplasma 216, 71–74 (2001)
Nishiyama, Y., Sugiyama, J., Chanzy, H. & Langan, P. Crystal structure and hydrogen bonding system in cellulose Iα from synchrotron X-ray and neutron fiber diffraction. J. Am. Chem. Soc. 125, 14300–14306 (2003)
McNamara, J., Morgan, J. L. W. & Zimmer, J. A molecular description of cellulose biosynthesis. Annu. Rev. Biochem. 84, 895–921 (2015)
Omadjela, O. et al. BcsA and BcsB form the catalytically active core of bacterial cellulose synthase sufficient for in vitro cellulose synthesis. Proc. Natl Acad. Sci. USA 110, 17856–17861 (2013)
Brown, C., Leijon, F. & Bulone, V. Radiometric and spectrophotometric in vitro assays of glycosyltransferases involved in plant cell wall carbohydrate biosynthesis. Nature Protocols 7, 1634–1650 (2012)
Somerville, C. Cellulose synthesis in higher plants. Annu. Rev. Cell Dev. Biol. 22, 53–78 (2006)
Morgan, J. L., Strumillo, J. & Zimmer, J. Crystallographic snapshot of cellulose synthesis and membrane translocation. Nature 493, 181–186 (2013)
Bokranz, W., Wang, X., Tschäpe, H. & Römling, U. Expression of cellulose and curli fimbriae by Escherichia coli isolated from the gastrointestinal tract. J. Med. Microbiol. 54, 1171–1182 (2005)
Whitney, J. C. et al. Structural basis for alginate secretion across the bacterial outer membrane. Proc. Natl Acad. Sci. USA 108, 13083–13088 (2011)
Keiski, C.-L. et al. AlgK is a TPR-containing protein and the periplasmic component of a novel exopolysaccharide secretin. Structure 18, 265–273 (2010)
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem. 77, 521–555 (2008)
Hubbard, C., McNamara, J. T., Azumaya, C., Patel, M. S. & Zimmer, J. The hyaluronan synthase catalyzes the synthesis and membrane translocation of hyaluronan. J. Mol. Biol. 418, 21–31 (2012)
Merzendorfer, H. Insect chitin synthases: a review. J. Comp. Physiol. B 176, 1–15 (2006)
Rehm, B. H. Alginate production: precursor biosynthesis, polymerization and secretion. Microbiology Monographs 13, 55–71 (2009)
Ross, P. et al. Regulation of cellulose synthesis in Acetobacter xylinum by cyclic diguanylic acid. Nature 325, 279–281 (1987)
Römling, U., Galperin, M. Y. & Gomelsky, M. Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol. Mol. Biol. Rev. 77, 1–52 (2013)
Morgan, J. L. W., McNamara, J. T. & Zimmer, J. Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP. Nature Struct. Mol. Biol. 21, 489–496 (2014)
Saxena, I. M., Brown, R. M., Jr & Dandekar, T. Structure–function characterization of cellulose synthase: relationship to other glycosyltransferases. Phytochemistry 57, 1135–1148 (2001)
Persson, K. et al. Crystal structure of the retaining galactosyltransferase LgtC from Neisseria meningitidis in complex with donor and acceptor sugar analogs. Nature Struct. Mol. Biol. 8, 166–175 (2001)
Chan, P. H. et al. Investigating the structural dynamics of α-1,4-galactosyltransferase C from Neisseria meningitidis by nuclear magnetic resonance spectroscopy. Biochemistry 52, 320–332 (2013)
Gardner, K. H. & Blackwell, J. The hydrogen bonding in native cellulose. Biochim. Biophys. Acta 343, 232–237 (1974)
Yang, H., Zimmer, J., Yingling, Y. G. & Kubicki, J. D. How cellulose elongates–A QM/MM study of the molecular mechanism of cellulose polymerization in bacterial CESA. J. Phys. Chem. B 119, 6525–6535 (2015)
Carpita, N. C. Update on mechanisms of plant cell wall biosynthesis: how plants make cellulose and other (1→4)-β-d-glycans. Plant Physiol. 155, 171–184 (2011)
Martin, J. L., Johnson, L. N. & Withers, S. G. Comparison of the binding of glucose and glucose 1-phosphate derivatives to T-state glycogen phosphorylase b. Biochemistry 29, 10745–10757 (1990)
Vocadlo, D. J., Davies, G. J., Laine, R. & Withers, S. G. Catalysis by hen egg-white lysozyme proceeds via a covalent intermediate. Nature 412, 835–838 (2001)
Qasba, P. K., Ramakrishnan, B. & Boeggeman, E. Structure and function of β-1,4-galactosyltransferase. Curr. Drug Targets 9, 292–309 (2008)
Forood, B., Feliciano, E. J. & Nambiar, K. P. Stabilization of alpha-helical structures in short peptides via end capping. Proc. Natl Acad. Sci. USA 90, 838–842 (1993)
Dirr, H. W., Little, T., Kuhnert, D. C. & Sayed, Y. A conserved N-capping motif contributes significantly to the stabilization and dynamics of the C-terminal region of class alpha glutathione S-transferases. J. Biol. Chem. 280, 19480–19487 (2005)
Scharnagl, C. et al. Side-chain to main-chain hydrogen bonding controls the intrinsic backbone dynamics of the amyloid precursor protein transmembrane helix. Biophys. J. 106, 1318–1326 (2014)
Cao, Z. & Bowie, J. U. Shifting hydrogen bonds may produce flexible transmembrane helices. Proc. Natl Acad. Sci. USA 109, 8121–8126 (2012)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
PyMol. DeLano Scientific (Sancarlos: CA, USA, )
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013)
Acknowledgements
We thank T. Rapoport for critical comments on the manuscript and J. Acheson for advice on reducing BcsA–BcsB complexes in crystallo. Diffraction data were collected at the Argonne National Laboratory’s Advanced Photon Source (APS) beam lines 23-ID-D (GM/CA-), 22-ID (SER-) and 24-ID-C (NE-CAT). GM/CA@APS has been funded in whole or in part with Federal funds from the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006). The NE-CAT beam lines are funded by the National Institute of General Medical Sciences from the National Institutes of Health (P41 GM103403). The Pilatus 6M detector on 24-ID-C beam line is funded by a NIH-ORIP HEI grant (S10 RR029205). Data for this research was also in part collected at the APS SER-CAT beam line, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by ANL under Contract No. DE-AC02-06CH11357. J.L.W.M. is supported by a National Science Foundation Graduate Research Fellowship, Grant No. DGE-1315231. M.F. thanks the Austrian Science Fund (FWF) (J3293-B21) for an Erwin Schrödinger postdoctoral fellowship. This research was primarily supported by the National Institutes of Health, Grant 1R01GM101001, awarded to J.Z.; S.G.W. thanks the Natural Sciences and Engineering Research Council of Canada for financial support.
Author information
Authors and Affiliations
Contributions
J.T.M. and J.L.W.M. purified and crystallized BcsA–BcsB and performed all crystal soaking experiments. J.T.M. cloned and analysed all BcsA cysteine mutants. J.T.M. and J.L.W.M. collected and processed diffraction data and built and refined the BcsA–BcsB models. M.F. synthesized the fluorinated and phosphonate UDP-Glc analogues and J.R. and H.-M.C. synthesized the UDP-thio-galactose analogues. J.T.M., J.L.W.M. and J.Z. analysed the data. J.Z. and J.L.W.M. wrote the paper and all authors edited the text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Conformational flexibility of the gating loop after cellulose extension.
Unbiased Sigma-A weighted Fo − Fc difference electron density of the gating loop in the pre-translocation state contoured at 2σ. The ordered part of the gating loop is shown as a thick green ribbon and two alternative backbone positions are indicated by a black dashed line. The position of the gating loop in the inserted state in the presence of a UDP molecule as observed in PDB entry 4P00 is shown as a cartoon representation coloured blue.
Extended Data Figure 2 In crystallo translocation of a 6-thio-galactose-containing cellulose polymer.
The position of the 6-thio-galactose group at the polymer’s non-reducing end was determined after polymer extension (upper panel) and upon subsequent incubation with UDP/Mg2+ (lower panel) in an anomalous difference Fourier electron density (DANO) map. DANO peaks detected at a wavelength of 1.74 Å are shown as a red mesh contoured at 3.5σ. Unbiased Sigma-A weighted Fo − Fc difference electron density for the cellulose polymer is shown as a green mesh contoured at 4σ. The cellulose polymer was extended and translocated as described in Fig. 1 with the exception that UDP-6-thio-galactose was used as substrate and Mg2+ was included during the initial soaking step. The extended DANO peak around Cys318 in the post-translocation state might arise from overlapping peaks originating from Cys318 and the thio-Gal unit in an opposite orientation. All Cys and Met residues close to BcsA’s active site are shown as sticks. UDP is shown as sticks in violet for its carbon atoms.
Extended Data Figure 3 Position of the disulfide-tethered finger helix.
The BcsA-2C–BcsB complex was crystallized as described for wild-type BcsA–BcsB. a, Unbiased Sigma-A weighted Fo − Fc difference electron density of BcsA’s finger helix contoured at 4σ (magenta mesh). Cellulose and BcsA’s Trp383 at the entrance to the transmembrane channel are shown as sticks in cyan and grey for their carbon atoms, respectively. The finger helix and IF2 are shown as cartoon helices coloured yellow and grey, respectively. b, The finger helix-tethered BcsA–BcsB complex was refined in a resolution range from 34 to 3.2Å to a final R/Rfree of 19.9/23.9% in Phenix_refine34 with Ala residues at positions 338 and 394 of BcsA. A strong difference electron density peak indicates the position of the omitted disulfide bond in a Sigma-A weighted Fo − Fc difference electron density map, (green mesh, contoured at 2σ).
Extended Data Figure 4 Comparison of the UDP conformation in the substrate and UDP-bound states of BcsA.
The substrate-bound BcsA structure was superimposed with PDB entry 4P00 by secondary structure matching in Coot. The substrate is shown as ‘balls and sticks’ in violet for the carbon atoms and the UDP molecule from PDB entry 4P00 is shown as grey sticks. BcsA’s finger helix is shown as a yellow cartoon and the cellulose polymer is shown as cyan and red ‘balls and sticks’ as observed in PDB entry 4P00. Magnesium is shown as a green sphere.
Extended Data Figure 5 UDP-Glc induced polymer translocation.
The nascent cellulose polymer was extended with a chain-terminating galactose residue upon soaking BcsA–BcsB crystals with UDP-Gal. Following dilution of the substrate as described in Fig. 1, crystals were incubated for 150 min either in the absence of a nucleotide or in the presence of UDP/Mg2+ or UDP-Glc/Mg2+, respectively. The unbiased SigmaA-weighted Fo − Fc difference electron density of the nascent polymer (green mesh) is shown at three different contour levels, indicating that UDP-Glc also induces polymer translocation.
Extended Data Figure 6 Stabilization of BcsA’s finger helix by conserved residues.
Top panel: stick representation of BcsA’s finger helix and nascent cellulose polymer shown in yellow and cyan for their carbon atoms. The finger helix is shown as a poly-glycine helix except for the labelled residues. Bottom panel: The finger helix’s ‘TEDxxT’ motif is conserved among pro- and eukaryotic cellulose synthases. Finger helix sequences are aligned for Micrasterias denticulata CesA, Physcomitrella patens CesA5, Arabidopsis thaliana CesA8, Rhodobacter sphaeroides and Escherichia coli BcsA, and Ciona savignyi CesA. The conserved threonine following the TED motif is indicated with a red box. Of note, the threonine residue is absent from the Ciona CesA sequence; however, this protein contains a serine residue at the following position, which could perform a similar function.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data and Supplementary Figures 1-4. (PDF 4118 kb)
Rights and permissions
About this article
Cite this article
Morgan, J., McNamara, J., Fischer, M. et al. Observing cellulose biosynthesis and membrane translocation in crystallo. Nature 531, 329–334 (2016). https://doi.org/10.1038/nature16966
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16966
This article is cited by
-
Structure, catalysis, chitin transport, and selective inhibition of chitin synthase
Nature Communications (2023)
-
Cationic cotton modified by 3-chloro-2-hydroxypropyl trimethyl ammonium chloride for salt-free dyeing with high levelling performance
Cellulose (2022)
-
Cellulose-synthesizing machinery in bacteria
Cellulose (2022)
-
In situ tunability of bacteria derived hierarchical nanocellulose: current status and opportunities
Cellulose (2021)
-
In silico structure prediction of full-length cotton cellulose synthase protein (GhCESA1) and its hierarchical complexes
Cellulose (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.