Abstract
G protein-coupled receptors (GPCRs) are physiologically important transmembrane signalling proteins that trigger intracellular responses upon binding of extracellular ligands. Despite recent breakthroughs in GPCR crystallography1,2,3, the details of ligand-induced signal transduction are not well understood owing to missing dynamical information. In principle, such information can be provided by NMR4, but so far only limited data of functional relevance on few side-chain sites of eukaryotic GPCRs have been obtained5,6,7,8,9. Here we show that receptor motions can be followed at virtually any backbone site in a thermostabilized mutant of the turkey β1-adrenergic receptor (β1AR)10,11,12. Labelling with [15N]valine in a eukaryotic expression system provides over twenty resolved resonances that report on structure and dynamics in six ligand complexes and the apo form. The response to the various ligands is heterogeneous in the vicinity of the binding pocket, but gets transformed into a homogeneous readout at the intracellular side of helix 5 (TM5), which correlates linearly with ligand efficacy for the G protein pathway. The effect of several pertinent, thermostabilizing point mutations was assessed by reverting them to the native sequence. Whereas the response to ligands remains largely unchanged, binding of the G protein mimetic nanobody NB80 and G protein activation are only observed when two conserved tyrosines (Y227 and Y343) are restored. Binding of NB80 leads to very strong spectral changes throughout the receptor, including the extracellular ligand entrance pocket. This indicates that even the fully thermostabilized receptor undergoes activating motions in TM5, but that the fully active state is only reached in presence of Y227 and Y343 by stabilization with a G protein-like partner. The combined analysis of chemical shift changes from the point mutations and ligand responses identifies crucial connections in the allosteric activation pathway, and presents a general experimental method to delineate signal transmission networks at high resolution in GPCRs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rasmussen, S. G. F. et al. Crystal structure of the human β2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007)
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011)
Ghosh, E., Kumari, P., Jaiman, D. & Shukla, A. K. Methodological advances: the unsung heroes of the GPCR structural revolution. Nature Rev. Mol. Cell Biol. 16, 69–81 (2015)
Grzesiek, S. & Sass, H.-J. From biomolecular structure to functional understanding: new NMR developments narrow the gap. Curr. Opin. Struct. Biol. 19, 585–595 (2009)
Bokoch, M. P. et al. Ligand-specific regulation of the extracellular surface of a G-protein-coupled receptor. Nature 463, 108–112 (2010)
Kofuku, Y. et al. Efficacy of the β2-adrenergic receptor is determined by conformational equilibrium in the transmembrane region. Nat. Commun. 3, 1045–1049 (2012)
Liu, J. J., Horst, R., Katritch, V., Stevens, R. C. & Wüthrich, K. Biased signaling pathways in β2-adrenergic receptor characterized by 19F-NMR. Science 335, 1106–1110 (2012)
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013)
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015)
Warne, T. et al. Structure of a β1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008)
Miller, J. L. & Tate, C. G. Engineering an ultra-thermostable β1-adrenoceptor. J. Mol. Biol. 413, 628–638 (2011)
Miller-Gallacher, J. L. et al. The 2.1 Å resolution structure of cyanopindolol-bound β1-adrenoceptor identifies an intramembrane Na+ ion that stabilises the ligand-free receptor. PLoS One 9, e92727 (2014)
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995)
Deupi, X. & Standfuss, J. Structural insights into agonist-induced activation of G-protein-coupled receptors. Curr. Opin. Struct. Biol. 21, 541–551 (2011)
Dror, R. O. et al. Pathway and mechanism of drug binding to G-protein-coupled receptors. Proc. Natl Acad. Sci. USA 108, 13118–13123 (2011)
González, A., Perez-Acle, T., Pardo, L. & Deupi, X. Molecular basis of ligand dissociation in β-adrenergic receptors. PLoS One 6, e23815 (2011)
Baker, J. G., Proudman, R. G. W. & Tate, C. G. The pharmacological effects of the thermostabilising (m23) mutations and intra and extracellular (β36) deletions essential for crystallisation of the turkey β-adrenoceptor. Naunyn Schmiedebergs Arch. Pharmacol. 384, 71–91 (2011)
Rasmussen, S. G. F. et al. Structure of a nanobody-stabilized active state of the β2 adrenoceptor. Nature 469, 175–180 (2011)
Manglik, A. & Kobilka, B. The role of protein dynamics in GPCR function: insights from the β2AR and rhodopsin. Curr. Opin. Cell Biol. 27, 136–143 (2014)
Grzesiek, S., Cordier, F., Jaravine, V. & Barfield, M. Insights into biomolecular hydrogen bonds from hydrogen bond scalar couplings. Prog Nucl Mag Res Sp 45, 275–300 (2004)
Tate, C. G. & Schertler, G. F. Engineering G protein-coupled receptors to facilitate their structure determination. Curr. Opin. Struct. Biol. 19, 386–395 (2009)
Goncalves, J. A. et al. Highly conserved tyrosine stabilizes the active state of rhodopsin. Proc. Natl Acad. Sci. USA 107, 19861–19866 (2010)
Park, J. H., Scheerer, P., Hofmann, K. P., Choe, H.-W. & Ernst, O. P. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature 454, 183–187 (2008)
Dror, R. O. et al. Activation mechanism of the β2-adrenergic receptor. Proc. Natl Acad. Sci. USA 108, 18684–18689 (2011)
Warne, T. et al. The structural basis for agonist and partial agonist action on a β1-adrenergic receptor. Nature 469, 241–244 (2011)
Brueckner, F. et al. Structure of β-adrenergic receptors. Methods Enzymol. 520, 117–151 (2013)
Alexandrov, A. I., Mileni, M., Chien, E. Y. T., Hanson, M. A. & Stevens, R. C. Microscale fluorescent thermal stability assay for membrane proteins. Structure 16, 351–359 (2008)
Vajpai, N. et al. Backbone NMR resonance assignment of the Abelson kinase domain in complex with imatinib. Biomol. NMR Assign. 2, 41–42 (2008)
Bayburt, T. H., Grinkova, Y. V. & Sligar, S. G. Self-assembly of discoidal phospholipid bilayer nanoparticles with membrane scaffold proteins. Nano Lett. 2, 853–856 (2002)
Ernst, O. P., Bieri, C., Vogel, H. & Hofmann, K. P. Intrinsic biophysical monitors of transducin activation: fluorescence, UV-visible spectroscopy, light scattering, and evanescent field techniques. Methods Enzymol. 315, 471–489 (2000)
Han, B., Liu, Y., Ginzinger, S. W. & Wishart, D. S. SHIFTX2: significantly improved protein chemical shift prediction. J. Biomol. NMR 50, 43–57 (2011)
Acknowledgements
We acknowledge T. Sharpe for expert help with biophysical assays. We thank J. Steyaert for providing the NB80 plasmid and M. Rogowski for preparing wild-type β1AR baculovirus. F.B. was supported by Marie Curie and EMBO postdoctoral fellowships. This work was supported by Swiss National Science Foundation grants 31-132857 (S.G.), Sinergia 141898 (S.G., D.B.V., G.F.X.S.), 31-135754 (D.B.V.), 31-153145 (G.F.X.S.), 31-146520 (X.D.), European Union FP7 grant 242135 (S.G.), and COST Action CM1207 (GLISTEN) (X.D. and G.F.X.S.).
Author information
Authors and Affiliations
Contributions
S.G., G.F.X.S., D.B.V., X.D. and S.I. initiated and managed the project. S.I., X.D., F.B., D.B.V., G.F.X.S. and S.G. designed ligand response and selective labelling experiments. C.O. designed the initial TS-β1AR construct and established purification. S.I. designed and prepared all selectively labelled receptor mutants, performed ligand exchange, NB80 binding, and all NMR experiments. S.I. and S.G. designed NMR experiments, analysed and interpreted all data. F.M.H. and D.B.V. purified trimeric Gi protein, designed and performed radioligand affinity and Gi protein activation assays. C.-J.T. prepared NB80 nanobody. S.G., S.I., X.D., F.M.H., D.B.V. and G.F.X.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Position of valine residues in the thermostabilized mutant TS-β1AR.
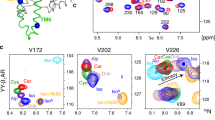
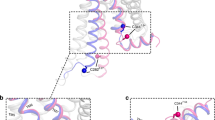
a, Schematic representation of secondary structure and amino acid sequence of TS-β1AR. As compared to the wild type, the TS-β1AR has truncations at the N and C termini and the intracellular loop (ICL3) as well as nine thermostabilizing point mutations (colour coded green) and three additional point mutations (colour coded yellow). Valines labelled with 15N are indicated by bold circles for assigned (blue) and unassigned (grey) residues. b, Structure of β1AR in complex with carvedilol (PDB code 4AMJ). The protein backbone and carvedilol are shown in ribbon and red stick representation, respectively. The individual valines are depicted as spheres (blue, assigned; grey, not assigned) labelled by residue number. c, Full 1H–15N TROSY spectra of apo TS-β1AR and all investigated ligand complexes. The ligand chemical structures are shown as inserts. Resonances are marked with assignment information (black, firm; cyan, tentative).
Extended Data Figure 2 Effect of ligand head group substitution on hydrogen bond network involving V172(4.56) in β1AR.
a, Sequence alignment of turkey β1AR and human adrenoreceptors around position 4.56 in TM4 showing the conserved proline at position 4.60. b, Schematic representation of the hydrogen bond network between TM4 and TM5 involving V172(4.56) in β1AR. The hydrogen bond network (orange dotted lines) originates at the carbonyl of V172(4.56) and connects to TM5 via two water molecules (w1 and w2, red asterisks). c, Left, phi and psi dihedral angles (averages and standard deviations in case of multiple chains) in TM4 between residues 4.48 and 4.61 for different complexes of β1AR (ligand, PDB code [chain]: carvedilol, 4AMJ [A,B]; cyanopindolol, 2VT4 [B,D] and 4BVN [A]; dobutamine, 2Y00 [A,B] and 2Y01 [A,B]; isoprenaline, 2Y03 [A,B]). V165(4.49) and V172(4.56), for which 1H–15N resonances could be observed, are shown on a green and red background, respectively. c, Right, average of the 1H–15N resonance positions for V165(4.49, green) and V172(4.56, red). The phi and psi values of V172(4.56) are distorted from the normal helical angles due to the loss of the intra-helical hydrogen bond to P176(4.60). Together with the loss of the canonical hydrogen bond, this strongly shifts both the 1H and 15N chemical shifts of V172(4.56) towards smaller p.p.m. values relative to V165(4.49), which has normal, helical phi and psi angles. d, Partial views of the crystal structures of the carvedilol complex (PDB code 4AMJ), the isoprenaline complex (PDB code 2Y03) and a docking model of the atenolol complex based on the cyanopindolol complex crystal structure (PDB code 4BVN). In the carvedilol complex, one water molecule (w2) forms a hydrogen bond network between the carbonyl oxygen of V172(4.56) and the side chains of S211(5.43) and S215(5.461). In the isoprenaline complex, the side chains of S211(5.43) and S215(5.461) are rotated and form hydrogen bonds to the catechol hydroxyl groups of isoprenaline. In this structure, no water molecule is observed at the equivalent position of w2. In the atenolol complex, the para-substituted acetamide of the ligand head ring (dashed circle) inserts between S211(5.43) and S215(5.461) and disturbs the interface between TM3 and TM5 near V172(4.56) (solid circle).
Extended Data Figure 3 Response to various ligands for all assigned valine 1H–15N resonances in TS-β1AR.
Colour coding as in Extended Data Fig. 1: cyan (atenolol), blue (isoprenaline), orange (dobutamine), purple (alprenolol), red (carvedilol), green (cyanopindolol), black (apo receptor). For clarity the centres of resonances are marked by circles. Firmly (tentatively) assigned residues are marked in black (cyan).
Extended Data Figure 4 Evidence for micro- to millisecond dynamics at the ligand entry/exit pathway.
Left, region of the 1H–15N TROSYs showing the V314(6.59) 1H–15N resonance of TS-β1AR in the apo and various ligand-bound forms. The resonances of the apo and atenolol-bound forms are severely broadened in the 15N dimension indicative of backbone dynamics in the micro- to millisecond range. The broadening is not observed for the other more tightly binding ligands. The resonances for the low affinity ligands, atenolol, isoprenaline and dobutamine, and for the high affinity ligands, alprenolol, carvedilol and cyanopindolol cluster at different positions. Interestingly, the resonance of the apo form clusters with the high affinity ligands, indicative of a similar backbone conformation. Right, 15N and 1H line widths of the V314(6.59) resonance for the apo and ligand-bound forms of TS-β1AR.
Extended Data Figure 5 Isoprenaline-induced G protein activation by β1AR mutants.
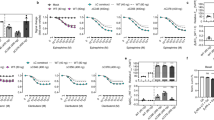
Activation was measured by the change in tryptophan fluorescence upon exchange of GDP for GTPγS in the Gα subunit of the heterotrimeric Gi protein. The figure shows the time courses of tryptophan fluorescence after the addition of different mutant forms of TS-β1AR in nanodiscs and GTPγS to the heterotrimeric G protein in the presence (red) and absence (black) of the agonist isoprenaline. The increase in fluorescence intensity at time t = 0 is caused by the additional fluorescence of the receptor, the nanodisc scaffold protein and the ligand. For the active mutant TS-β1AR(A227Y/L343Y), this initial rise is followed by an exponential fluorescence increase due to the activation and dissociation of the G protein. The rate of the Gαi activation by TS-β1AR(A227Y/L343Y) (half-life ~290 s at 6 nM receptor) is approximately 4 times slower than for the activation by rhodopsin (half-life ~400 s at 1 nM receptor) using the same assay. This assumes that the Km of Gαi for TS-β1AR(A227Y/L343Y) is of the same scale as for rhodopsin (8.6 nM) and therefore significantly below the (saturating) concentration of Gαi (100 nM). Each mutant was measured three times except for TS-β1AR(A227Y/L343Y) that was measured four times. Representative fluorescence curves from single experiments are shown.
Extended Data Figure 6 Observed NMR effects of the reverse mutations of TS-β1AR towards the native β1AR sequence.
a, 1H–15N TROSY spectra of TS-β1AR and several reverse single and double mutants in complex with either atenolol (cyan) or isoprenaline (dark blue). Resonances are marked with assignment information (black, definite; cyan, tentative). b, Enlarged regions of the 1H–15N correlation spectra showing only the resonance positions of V226(5.57) and V280(6.25) for all mutants in both ligand-receptor complexes. The black bar represents 0.1 p.p.m. in 1H and 1 p.p.m. in 15N. For all mutants, the resonances for V226(5.57) show efficacy-related chemical shift changes between atenolol (diamonds) and isoprenaline (circles). For the A227Y mutants TS-β1AR(A227Y) (green), TS-β1AR(V129I/A227Y) (orange) and TS-β1AR(A227Y/L343Y) (cyan), the 1H–15N resonances of V226(5.57) also exhibit an overall shift due to a ring current effect from the introduced Y227(5.58) side chain, which has no structural significance. However, the V129I mutants TS-β1AR(V129I) (red) and TS-β1AR(V129I/A227Y) (orange) exhibit further 1H–15N shifts towards a more active (that is, bent) state of TM5 relative to the mutants that carry the V129(3.40) residue. For all mutants, the V280(6.25) resonances fall basically in identical positions and show no changes between atenolol and isoprenaline.
Extended Data Figure 7 Binding of NB80 to agonist-activated TS-β1AR(A227Y/L343Y).
a, Sequential addition of isoprenaline (1 mM, blue), NB80 (120 μM, red), and cyanopindolol (1 mM, green) to TS-β1AR(A227Y/L343Y) (120 μM) followed by 1H–15N TROSY spectra. Addition of an equimolar amount of NB80 changes the spectrum drastically. Except for few resonances (62, 89, 102, 103, 172, 280 and 326), the assignments are not transferable from the isoprenaline complex. The addition of cyanopindolol to this mixture apparently replaces isoprenaline in the receptor and drives the spectrum to the purely cyanopindolol-bound form. These results indicate that TS-β1AR(A227Y/L343Y) can be activated, but that the fully active state requires stabilization with a G protein-like partner at the intracellular site. b, Changes in isoprenaline affinity of β1AR induced by NB80 binding. The isoprenaline affinity was assayed for TS-β1AR (orange), TS-β1AR(A227Y/L343Y) (green), and the truncated native turkey β1AR receptor (tβtrunc, blue)17 by radio-ligand competition using 3H-dihydroalprenolol in the presence of saturating amounts of NB80 (10 μM, squares) or its absence (circles). For TS-β1AR(A227Y/L343Y) and tβtrunc the apparent affinity increases by two log units in the presence of NB80, whereas no change occurs for TS-β1AR. Data points are shown as mean and standard deviation of three independent experiments for each condition. c, Combined 1H and 15N chemical shift changes (Δδ = (Δδ1H2/2 + Δδ15N2/50)1/2) between the isoprenaline-TS-β1AR(A227Y/L343Y) and the NB80-isoprenaline-TS-β1AR(A227Y/L343Y) complexes as a function of residue number. Except for the few assigned peaks in the NB80-isoprenaline-TS-β1AR(A227Y/L343Y) complex, chemical shift deviations are calculated as the minimal difference from the isoprenaline peak position to the nearest peak position in the isoprenaline–NB80 complex. Note that these minimal chemical shift deviations present a lower boundary for the deviations that could be obtained from a full assignment of the isoprenaline–NB80 complex. Peak deviations are colour-coded in red (≥0.2 p.p.m.), orange (0.2–0.05 p.p.m.), blue (<0.05 p.p.m.), and grey (ambiguity from peak overlap). d, Representation of the chemical shift response to nanobody binding given in c on the structure of β1AR. Valines in TM1 and TM7 do not show a large chemical shift response. Due to overlap the response for V280(6.25) in TM6 is unclear. Valines in TM2 show a strong response around the sodium binding region near residue D87(2.50), but valines at its extracellular side are not responsive. In general, valine resonances at the extracellular sides of TM3–TM6 are severely shifted. This indicates that the NB80 binding at the intracellular side has a long-range effect on the extracellular side near the ligand binding site.
Supplementary information
Supplementary information
This file contains Supplementary Text 1. (PDF 186 kb)
Rights and permissions
About this article
Cite this article
Isogai, S., Deupi, X., Opitz, C. et al. Backbone NMR reveals allosteric signal transduction networks in the β1-adrenergic receptor. Nature 530, 237–241 (2016). https://doi.org/10.1038/nature16577
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16577
This article is cited by
-
Programming conformational cooperativity to regulate allosteric protein-oligonucleotide signal transduction
Nature Communications (2023)
-
E. coli “Stablelabel” S30 lysate for optimized cell-free NMR sample preparation
Journal of Biomolecular NMR (2023)
-
Filling of a water-free void explains the allosteric regulation of the β1-adrenergic receptor by cholesterol
Nature Chemistry (2022)
-
Efficient production of a functional G protein-coupled receptor in E. coli for structural studies
Journal of Biomolecular NMR (2021)
-
A high-resolution description of β1-adrenergic receptor functional dynamics and allosteric coupling from backbone NMR
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.