Abstract
Seagrasses colonized the sea1 on at least three independent occasions to form the basis of one of the most productive and widespread coastal ecosystems on the planet2. Here we report the genome of Zostera marina (L.), the first, to our knowledge, marine angiosperm to be fully sequenced. This reveals unique insights into the genomic losses and gains involved in achieving the structural and physiological adaptations required for its marine lifestyle, arguably the most severe habitat shift ever accomplished by flowering plants. Key angiosperm innovations that were lost include the entire repertoire of stomatal genes3, genes involved in the synthesis of terpenoids and ethylene signalling, and genes for ultraviolet protection and phytochromes for far-red sensing. Seagrasses have also regained functions enabling them to adjust to full salinity. Their cell walls contain all of the polysaccharides typical of land plants, but also contain polyanionic, low-methylated pectins and sulfated galactans, a feature shared with the cell walls of all macroalgae4 and that is important for ion homoeostasis, nutrient uptake and O2/CO2 exchange through leaf epidermal cells. The Z. marina genome resource will markedly advance a wide range of functional ecological studies from adaptation of marine ecosystems under climate warming5,6, to unravelling the mechanisms of osmoregulation under high salinities that may further inform our understanding of the evolution of salt tolerance in crop plants7.
Similar content being viewed by others
Main
Seagrasses are a polyphyletic assemblage of basal monocots belonging to four families in the Alismatales1,2 (Supplementary Note 1.1 and Supplementary Fig. 1.1). As a functional group, they provide the foundation of highly productive ecosystems present along the coasts of all continents except Antarctica, where they rival tropical rain forests and coral reefs in ecosystem services8,9. In colonizing sedimentary shorelines of the world’s ocean, seagrasses found a vast new habitat free of terrestrial competitors and insect pests but had to adapt to cope with new structural and physiological challenges related to full marine conditions.
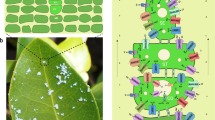
Zostera marina (Zosteraceae), or eelgrass (Fig. 1), is the most widespread species throughout the temperate northern hemisphere of the Pacific and Atlantic10. A clone of Z. marina was sequenced from the Archipelago Sea, southwest Finland, using a combination of fosmid-ends and whole-genome shotgun (WGS) approaches (Methods, Supplementary Note 2). The 202.3 Mb Z. marina genome encodes 20,450 protein-coding genes, 86.6% of which (17,511 genes, Supplementary Note 3.1) are supported by transcriptome data from leaves, roots and flowers (Extended Data Fig. 1, Supplementary Notes 3.2–3.3 and Supplementary Data 1–3). Genes are located in numerous gene-dense islands separated by stretches of repeat elements accounting for 63% of the non-gapped assembly (Extended Data Fig. 2, Supplementary Note 3.1) as compared to only 13% in the only other sequenced alismatid, the freshwater duckweek, Spirodela polyrhiza (Alismatales, Araceae)11. Gypsy-type (32%) and Copia-type (20%) transposable elements contribute to most of the repetitive DNA. Sequence divergence analysis suggests that the genome retains copies from two distinct periods of invasion by Copia elements, but only one period for Gypsy elements (Extended Data Fig. 3a–c). Genes gained by Z. marina (‘accessory’) are located closer to transposable elements than to conserved (‘single copy’) genes (Fisher’s exact test, P < 0.0001) indicating that transposable elements may have played a role in genic adaptation.
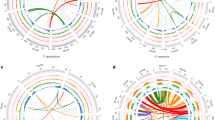
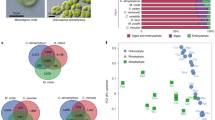
a, Gains and losses are indicated along branches and nodes. The number of gene families, orphans (single-copy gene families) and number of predicted genes is indicated next to each species. Background colours (top to bottom) are Alismatales, other monocots, dicots, mosses/algae b, Typical Zostera marina meadow, Archipelago Sea, southwest Finland (photo by C.B.).
We identified 36 conserved microRNAs with high confidence and their predicted targets (Supplementary Note 3.4, Supplementary Data 4 and 5). A novel variant of miR528 (not present in Spirodela) was found to be the only member of this miRNA family, and demonstrates that this conserved miRNA is the only one ancestral to the entire monocot lineage. Most likely, Z. marina did not take part in the subsequent birth of miRNAs that are common to several other monocots12; nor did it experience or retain traces of prominent miRNA duplications.
Analysis of synonymous substitutions per synonymous site (KS) age distributions indicates that Z. marina carries the remnants of an independent, ancient whole-genome duplication (WGD) event (Fig. 2a, Supplementary Note 4.1)13. Duplicated segments account for ~9% of the Z. marina genome, probably an underestimate due to the fragmented nature of the assembly. Zostera and Spirodela diverged somewhere between 135 and 107 million years ago (Mya)14 and phylogenomic dating13 of the Z. marina WGD suggests that it occurred 72–64 Mya (Fig. 2b), thus independently from the two WGDs reported for S. polyrhiza11. This timeframe coincides with the initial diversification of a freshwater clade that includes three of the four families of seagrasses (Supplementary Table 1.1) and with the Cretaceous–Palaeogene (K–Pg) extinction event (Fig. 2c), which provided new ecological opportunities and may have triggered seagrass adaptive radiations.
a, KS-based age distribution of the whole Z. marina paranome. The x axis shows the synonymous distance until a KS cut-off of 2, in bins of 0.04, containing the KS values that were used for mixture modelling (excluding those with a KS ≤ 0.1). The component of the Gaussian mixture model plotted in red (as identified by EMMIX) corresponds to a WGD feature based on the SiZer analysis (other components are shown in black). The transition from the blue to the red at a KS of ~0.8 in the SiZer panel (below) indicates a change in the distribution and therefore provides evidence for an ancient WGD (Supplementary Table 4.1, Supplementary Fig. 4.1). b, Absolute age distribution obtained by phylogenomic dating of Z. marina paralogues. The solid black line represents the kernel density estimate (KDE) of the dated paralogues and the vertical dashed black line represents its peak, used as the consensus WGD age estimate, at 67 Mya. Grey lines represent the density estimates from 2,500 bootstrap replicates and the vertical black dotted lines represent the corresponding 90% confidence interval for the WGD age estimate, 64–72 Mya. The original raw distribution of dated paralogues is indicated by the circles. The y axis represents the percentage of gene pairs. c, Pruned phylogenetic tree with indication of WGD events (boxes)29. The Cretaceous–Palaeogene (K–Pg) boundary is indicated by an arrow.
We mapped signatures of loss and gain of gene families (Supplementary Note 4.2) onto a phylogenetic tree (Fig. 1a). We also mapped losses and gains of Pfam domains (Supplementary Fig. 4.4, Supplementary Data 6). While many genes are shared between Zostera and Spriodela, clearly some losses and gains are unique to Zostera in relation to its marine environment, the alismatid lineage having set the stage for the subsequent freshwater–marine transition. Those unique to Z. marina include the absence of all the genes involved in stomatal differentiation (Fig. 3a, Extended Data Table 1 and Supplementary Note 5.1) and the disappearance of genes comprising entire pathways encoding volatiles synthesis and sensing (Supplementary Note 6.1), such as those for ethylene15 (Fig. 3b, Extended Data Table 2). Terpenoid genes are also drastically reduced to two (Fig. 3c), as compared with four in Spirodela, 50 in Oryza and > 100 in Eucalyptus, thus precluding synthesis of secondary volatile terpenes (Supplementary Fig. 6.2). Only aromatic acid decarboxylases (AAAD) genes were expanded (Supplementary Fig. 6.3) and these form a clade distinct from Spirodela. The loss of volatiles is also consistent with the loss of stomata, through which they are emitted for airborne communication and plant defence. The repertoire of defence-related genes such as the six groups of NBS_LRR resistance genes (Supplementary Note 6.2) is also reduced to 44 (89 in Spirodela and 100–300 in other plants), which may be linked to a lower probability of infection of Z. marina due to the absence of stomata, which are a main entry point for pests and pathogens in terrestrial plants.
a, Stomata differentiation from meristemoid mother cells (MMC) to guard mother cell (GMC) to guard cells. b, Ethylene synthesis and signalling up to EIN2 have disappeared; EIN3 and its downstream targets remain. c, Terpenoid biosynthesis in which the pathways producing volatiles are absent but those essential for primary metabolism remain. MVA, mevalonate; MEP, plastidic methylerythritol phosphate; IPP, isopentenyl pyrophosphate; DMAPP, dimethylallyl pyrophosphate; FPP, farnesyl pyrophosphate; GPP, geranyl diphosphate; GGPP, geranylgeranyl pyrophosphate; CPP, copalyl pyrophosphate; GA, gibberellic acid; PP, diphosphate; ABA, abscisic acid. d, Sporopollenin biosynthesis genes; regulatory genes in the nucleus control downstream processes (arrows) in response to signalling coming from external stimuli through receptors on the plasma membrane. All panels: genes in red are absent; blue are present; the grey line represents the plasma membrane. See Extended Data Tables 1, 2, 3.
Land and aquatic floating plants (Embryophyta) are often exposed to intense ultraviolet (UV) radiation and have developed light sensing protein receptors with protective and signalling functions. In contrast, Z. marina inhabits a light-attenuated, submarine environment where it must cope with shifted spectral composition, characterized by low penetration of UV-B, red and far-red wavelengths16. Accordingly, Z. marina has lost ultraviolet-resistance (UVR8) genes associated with sensing and responding to UV damage (Spirodela has not), as well as phytochromes associated with red/far-red receptors (Supplementary Note 7). Whereas photosystems (PSI and PSII) are similar to those of other plants including Spirodela, members of the light-harvesting complex B (LHCB) family are expanded in number, possibly in combination with non-photochemical quenching (NPQ), thereby enhancing performance at low light (Extended Data Fig. 4).
Seagrasses typically experience full marine seawater (35 g kg−1)17, whereas land plants obtain water with low osmolality (0–2 g kg−1) via the rhizosphere and aquatic plants experience fresh (0–5 g kg−1) to brackish (0.5–20 g kg−1) conditions. Although Z. marina displays a typical repertoire of Na+ and K+ antiporters (Supplementary Note 8, Supplementary Table 8.1), one of six H+-ATPase (AHA) genes (Supplementary Table 8.2, Supplementary Data 7) is strongly expressed in vegetative tissue and encodes a salt-tolerant H+-ATPase. Furthermore, Z. marina possesses three AHA genes (along with Spirodela) in a cluster unique to alismatids (Supplementary Fig. 8.1).
Uniquely, Z. marina has re-evolved new combinations of structural traits related to the cell wall. Synthesis of cutin-cuticular waxes to the outside of the leaf epidermis and suberin–lignin near the plasma membrane (Supplementary Note 9, Supplementary Table 9.1) surround a cell wall matrix of (hemi)celluloses, low-methylated pectin (zosterin) and macroalgal-like sulfated polysaccharides18 (Supplementary Note 10). The reduction in carbohydrate-related genes that modify the fine structure of cell wall hemicelluloses and pectins in Z. marina is not due to loss of pathways, but rather to the large variation within these CAZyme gene families in plants. Available genomes (including Spirodela) lack carbohydrate sulfotransferases and sulfatases, suggesting that land plants have lost these genes as a key adaptation to terrestrial as well as freshwater conditions19,20. In contrast, Z. marina has regained the ability to produce sulfated polysaccharides with an expansion of aryl sulfotransferases (12 genes) homologous to aryl sulfotransferases from land plants (Supplementary Note 10). Sulfation facilitates water and ion retention in the cell wall to cope with desiccation and osmotic stress at low tide and, likewise, low methylation of zosterin correlates with the expanded pectin carbohydrate esterase 8 (CE8) family, increasing the polyanionic character of the cell wall matrix. We speculate that several aryl sulfotransferases have evolved because carbohydrate sulfatases have been shown to be active on artificial aryl compounds such as methylumbelliferyl-sulfate21. Osmotic equilibrium is further achieved in Z. marina by organic osmolytes (mainly sucrose, trehalose and proline) in combination with a small cytoplasm:vacuole volume ratio (10%)22. Given that up to 90% of fixed carbon is stored as sucrose in the rhizomes, sucrose synthase (SuSy) and transport (SUT) genes are expanded while those for starch metabolism are greatly reduced, as expected in ‘marine sugarcane’ (Supplementary Note 7.2, Supplementary Data 8).
The repertoire of redox and other stress-resistance genes (Supplementary Note 8) is typical for angiosperms with the exception of catalase (CAT), which is reduced to a single copy in Z. marina (two in Spirodela). Late embryogenesis abundant (LEA) and dehydrins are clearly under-represented in both Zostera and Spriodela relative to other genomes. In contrast, Zostera possesses an unusual complement of metallothioneins. Aside from their role as chelators, metallothioneins may be involved in stress resistance; one of these, MT2L, is among the most highly constitutively expressed genes in Z. marina (Extended Data Fig. 5, Supplementary Note 8.2).
Sexual reproduction of Z. marina takes place underwater, involving completely submerged male and female flowers, and a unique exine-less, filiform pollen that winds around the bifurcate stigmas in a purely abiotic pollination process23. Note that freshwater alismatids (and also Spirodela)24 possess pollen with an exine layer. Exine-less pollen25 is characteristic of all seagrasses except Enhalus acoroides (which is surface pollinated). Ten genes specifically involved in biosynthesis and modification of the pollen exine coat are missing; all other genes involved in the development of viable pollen remain intact (Fig. 3d, Extended Data Table 3, Supplementary Note 11.1). Finally, MADS-box gene transcription factors are also highly reduced to 50 in Z. marina, which is most likely related to its highly reduced flowers (also a feature of Spirodela) that lack the first two whorls of specialized floral leaves, calyx and corolla (Supplementary Note 11.2, Supplementary Table 11.2).
An increasing proportion of the world population inhabits the coastal zone. This impinges multiple pressures on ecosystems including seagrass beds26,27, which in turn compromises the ecosystem services they may provide, including provisioning of harvestable fish and invertebrates, nutrient retention, carbon sequestration and erosion control. In the context of seagrass conservation, elucidating the genomic basis of Z. marina’s complex adaptations to ocean waters (Extended Data Fig. 6) will also inform the development of molecular indicators of their physiological status28, as these unique ecosystems rank, unfortunately, among the most threatened on Earth26,27.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Plant material and DNA preparation
A single genotype/clone of Zostera marina (referred to as the ‘Finnish clone’) was harvested on 26 August 2010 at 2 m depth at Fårö Island (latitude 59° 55.234′ N longitude 21° 47.766′ E) located in the northern Baltic Sea, Finland. Plant material was transported to the lab in seawater, cleaned and further processed. Care was taken to use leaf-meristem tissue harvested from the inner layer of basal shoots to minimize bacterial/diatom contamination. Tissues were immediately frozen in LN2 and stored at −80 °C for later DNA and RNA extraction. Monoclonality was verified by genotyping 40 ramets of the mega-clone with six highly polymorphic, microsatellite loci30. There was no evidence for polyploidy25,31,32 (Z. marina is 2n = 12) or somatic mutations33 as assessed by multiple peaks in the microsatellite chromatograms. Tissue was subsequently sent on dry ice to Amplicon Express for HMW DNA extraction using a CTAB isolation method modified by R. Meilan (unpublished) but available from him (rmeilan@purdue.edu), based on the original method34. Following QC according to JGI guidelines, the DNA was shipped to JGI for library and sequencing preparation.
Genome sequencing and assembly
One 35-Kb, fosmid library was generated for end sequencing. The fosmid ends were sequenced with standard Sanger sequencing protocols at the HudsonAlpha Institute for a total of 194,303 Sanger reads (0.29× coverage). Illumina libraries (two fragment libraries (6.62 Gb), one 2-Kb JGI mate-pair library (3.57 Gb), one 4-Kb JGI mate-pair library (3.41 Gb) and two 8-Kb JGI mate-pair libraries (11.94 Gb)) were sequenced with Illumina MiSeq/HiSeq genetic analysers at the Department of Energy’s Joint Genome Institute (JGI), using standard protocols. A total of 25.55 Gb of Illumina and 0.14 Gb of Sanger sequence was obtained representing 47.7× genomic coverage. Prior to assembly, all reads were screened against mitochondria, chloroplast, and Illumina controls. Reads composed of > 95% simple sequence repeats were removed. For the Illumina, paired-end libraries (2 × 250), reads <75 bp were discarded, for the 2 × 150 libraries, reads <50 bp were discarded after trimming for adaptor and quality (q < 20). An additional deduplication step was performed on the mate pairs that identified and retained only one copy of each PCR duplicate. A total of 212,101,273 reads (Supplementary Table 2.1) was assembled using our modified version of Arachne v. 20071016 (ref. 35). Subsequent directed Arachne modules were applied to collapse adjacent heterozygous contigs. The entire assembly was then run through another Arachne process starting at Stage 6 Rebuilder. This produced 15,747 scaffold sequences (30,723 contigs), with a scaffold L50 of 409.5 Kb, 613 scaffolds larger than 100 Kb, and a total genome size of 237.5 Mb (Supplementary Table 2.2).
Scaffolds were screened against bacterial proteins, organelle sequences, GenBank NR (nr_prot) and RefSeq protein databases, and removed if found to be a contaminant. Scaffolds consisting of prokaryotes, chloroplast, mitochondria and unanchored rDNA were removed. We also assembled the chloroplast and partial mitochondrial genomes (Supplementary Notes 2.2 and 2.3, Supplementary Fig. 2.1). Additionally, short (<1 Kb) scaffolds or scaffolds containing highly repetitive sequence ( > 95% 24-mers found more than four times in large scaffolds) or alternative haplotypes were also removed. Following repeat analysis and gene prediction, all scaffolds were subjected to a filtering process (based on NCBI nr_prot + NCBI taxonomy database) to eliminate remaining bacterial (and other) contaminants (Supplementary Table 2.3).
Assembly validation was performed using a set of 12 fully sequenced fosmid clones. In 4 of the 12 fosmid clones, full-length alignments were not found due to fragmentation in the region of the fosmid clone. In five of the remaining eight fosmid clones, the alignments were of high quality (<0.05% bp error). The overall base pair error rate (including marked gap bases) in the fosmid clones that aligned to full length was 0.28% (714 discrepant base pairs out of 253,332 bp). Supplementary Table 2.4 shows the individual fosmid clones and their contribution to the overall error rate. Note that two fosmid clones (16248, 16249) contributed nearly 81% of the discrepant bases. This probably occurred in polymorphic regions of the genome where the haplotype in the fosmid did not match the haplotype in the reference. There are several indels of various sizes in the clone and assembly, typical of a region of degraded transposons. Further quality analysis indicated that 90% of the set of eukaryotic core genes (CEGMA) were present and 98% were partially represented, suggesting near completeness of the euchromatin component.
Annotation of repetitive sequences
Two complementary approaches were used to identify repetitive DNA sequences in the Z. marina genome. With respect to masking repeats before gene prediction analysis, a de novo repeat identification was carried out with RepeatModeller (v. open-1.0.7; http://www.RepeatMasker.org)36 to identify repeat boundaries and build consensus models from which potential over represented, non-transposable element, protein-coding genes were removed. RepeatMasker (v. open-4.0.0, WUBlast) was used in combination with this custom repeat library to mask the assembly and prepare it for gene prediction with EuGene.
Furthermore, in order to perform a qualitative and quantitative analysis of repeats with greater resolution37 the genome assembly was processed for de novo repeat detection using the TEdenovo pipeline from the REPET package v. 2.2 (ref. 38); parameters were set to consider repeats with at least five copies. The consensus sequences generated by TEdenovo were then used as probes for whole genome annotation by the TEannot39 pipeline from the REPET package v. 2.2. The consensus repeat sequences were classified using Pastec40. Comparing the genomic positions of transposable elements (TE) to those of exons from the set of predicted genes enabled us to identify that 909 gene predictions most likely represent TEs and these were filtered from the gene set. The REPET package v. 2.2 was also used to annotate repetitive elements in the Spirodela polyrhiza genome assembly with the same parameters as for Z. marina. See Supplementary Fig. 3.1.
Transcriptome library preparation, sequencing and assembly
Leaf, root and flower tissues were separately frozen in liquid nitrogen immediately following harvest from either ambient (field collected) or experimental (mesocosm) conditions (Supplementary Note 3.2). Overall, we obtained between nine and 20 million high-quality reads from each of the flower-leaf-root replicate libraries; and for the Finnish clone library, 148.5 million high quality reads were retrieved (Supplementary Table 3.3).
The de novo assembly protocol was adapted from ref. 41. We pooled replicates of each tissue together except for the two leaf tissue libraries, which were kept separate (Supplementary Table 3.4) and performed de novo transcriptome assembly for each tissue using Trinity41(v. 2014-07-17) with digital normalization option ON to normalize input read coverage. Frame shift errors and insertion/deletion errors in the assembled transcripts were corrected by FrameDP42. Because a de novo assembly still generates many spurious transcripts, we used the transcript expression value to remove low quality contigs. We used the RSEM pipeline43 to obtain the contig expression values and removed contigs with FPKM (fragments per kilobase of transcript per million fragments mapped) value <1 and IsoPct (percentage of expression for a given transcript compared with all expression from that Trinity component) < 1. In total, we obtained between 39,000 and 53,000 assembled contigs from each library, and 52,000 contigs from the Finnish clone library (Supplementary Table 3.4). Prior to mapping the genome sequence and the predicted genes, we used the CD-HIT44 program (v. 4.6.1) to collapse redundant contigs, which resulted in 79,134 low redundant transcript contigs.
Differential gene expression analysis
High-quality RNA-seq reads were mapped to the genome assembly v.2.1 by TopHat45. Differential gene expression analysis was performed by the Cufflink pipeline45 based on the Z. marina v.2.1 gene models by converting the number of aligned reads into FPKM values. Genes with significant expression difference (log2 > 2) were selected for further investigation by GOstats46 to perform Gene Ontology (GO) term enrichment analysis with P ≤ 0.05 (Supplementary Note 3.3, Supplementary Table 3.5)
MicroRNA analysis
Genomic precursors of known miRNAs were mapped on the Z. marina genome following the procedure described in ref. 47 for the maize genome. miRNA entries from the miRBase database (release 21, 2014) were aligned to the chromosomes of the Z. marina genome. Up to three mismatches were allowed in the alignment, using SeqMap48. In parallel, novel potential DCL1/AGO1-dependent miRNAs were enriched by selecting 5′-U 20–22 nt small RNAs from three different sequenced libraries from Z. marina described in ref. 12. A subset of these small RNAs with abundance ≥10 TPM (transcripts per million) was retained and aligned to the genome with no mismatches. From every locus, we extracted two ~200-nt regions surrounding each aligned miRNA or candidate (from −30 to +160 and from −160 to +30 nucleotides relative to the putative miRNA start or end coordinate, respectively). Minimum energy RNA secondary structures were predicted for each region using the RNAfold program of the Vienna RNA 1.8.5 package (http://www.tbi.univie.ac.at/~ivo/RNA/) using default settings.
In addition, small RNAs from the three sequenced libraries were mapped on these regions, allowing no mismatches, in order to pre-select putative miRNA loci that showed evidence of expression in the three plant tissues analysed. We evaluated RNA structure and small RNA alignment in all the regions based on: (1) dominance of plus-stranded small RNAs; (2) position of the most abundant small RNAs relative to the predicted miRNA coordinates; (3) prevalence of 20–22 nt small RNAs in the predicted miRNA locus; (4) position of the putative miRNA with the stem-loop structure; and (5) absence of oversize (≥3 nt) bulges in the miRNA/miRNA* alignment. After reduction of overlapping loci to a non-redundant set and removal of stem-loop structures with the wrong orientation compared to miRNAs registered in miRBase, we manually inspected the remaining loci to further evaluate them according to the miRNA annotation criteria proposed by ref. 49. Stringency was relaxed when small RNA expression data strongly indicated the presence of miRNA loci that did not meet the whole set of criteria. Novel miRNA precursors overlapping with TEs or other repetitive elements were filtered out.
Potential miRNA targets were identified in silico using the generic small RNA-transcriptome aligner GSTAr from the CleaveLand package (v. 4)50. Predicted targets were accepted with an Allen score <4 or a MFE (minimum free energy) ratio ≥ 7.5. (Supplementary Note 3.4).
Gene prediction
Training of the gene prediction programs started with the collection of high quality ESTs. EST information was used, for example, to train the splice predictor SpliceMachine51. Detection of conserved splice sites was further investigated by RNA-seq splice junctions (count > 10) to construct a WAM model in EuGene (v. 4.1)52. Coding-potential was modelled with an interpolated Markov Model (IMM) constructed from the BLASTX alignments of proteins from the PLAZA v. 2.5 database53. An additional protein ‘monocot’ Markov Model was built based on the protein sequences from Brachypodium, maize and sorghum. Starting from EST and protein alignments, a set of 215 gene models was manually constructed and curated using the genome browser GenomeView54. The 215 models were then used as a training set for EuGene in order to optimize the different splice site and coding-potential models, as well as the weights for the extrinsic EST and homology evidence. An overall fitness score of 80.1% was achieved, which is high enough to obtain reliable results without overfitting. GeneMark55 and Augustus56 were separately trained (using the same input data as EuGene) and their predictions were integrated with EuGene using a custom script to evaluate the best gene structure at each locus. All gene models were automatically screened to highlight possible erroneous structures (for example, in-frame stop codons, deviating splice junctions) and manually curated. Transfer-RNA gene models were predicted by tRNAscan-SE (v. 1.31)57 and their structures were verified with Infernal (v. 1.1rc1, rfam11 covariant model database)58. For each gene, UTRs were assigned by identifying a set of ESTs and RNA-seq assemblies that uniquely overlapped with it. We subsequently selected the longest mapped transcript on either end of the predicted coding sequence and designated the section outside the coding sequence as the UTR. Finally, all genes were uploaded to the ORCAE platform (http://bioinformatics.psb.ugent.be/orcae)59, enabling all members of the consortium to refine and curate the gene model and assign gene function. A list of protein domains, as well as the derived Gene Ontology (GO) terms and KEGG pathway identifiers were generated using an InterProScan (v. 5.2.45)60 analysis and available in ORCAE. More specifically, gene functional descriptions were added either manually by consortium expert scientists or automatically through sequence homology searches. The automated method relies on the EC (Enzyme Commission) number reported by InterProScan to retrieve the enzyme name with BLASTP search against UniProtKB/Swiss-Prot61 to filter out hits that are below 60% identity and 70% query/hit coverage. Although such high stringency on per cent identity and sequence coverage reduced the available number of functional descriptions, it reduced the false-positive prediction rate, as desired here.
Construction of age distributions and WGD analyses
KS-based age distributions were constructed as previously described62. In brief, the KS values between genes were obtained through maximum likelihood estimation using the CODEML program63 of the PAML package (v. 4.4c)64. Gene families for which KS estimates between members did not exceed a value of 5 were subdivided into subfamilies. For each duplicated gene in the resulting phylogenetic gene tree, obtained by PhyML65, all m KS estimates between the two child clades were added to the KS distribution with a weight 1/m (where m is the number of KS estimates for a duplication event), so that the weights of all KS estimates for a single duplication event summed to one. Mixture modelling was used to confirm a WGD signature in the KS distribution (Fig. 2 and Supplementary Fig. 4.1), for which all duplicates with KS values ≤0.1 were excluded to avoid the incorporation of allelic and/or splice variants, while all duplicates with KS values > 2.0 were removed because KS saturation and stochasticity can mislead mixture modelling above this range62. For further details see Supplementary Note 4.1.
Absolute dating of the identified WGD event was performed as described previously13,29. In brief, paralogueous gene pairs located in duplicated segments (anchors) and duplicated pairs lying under the WGD peak (peak-based duplicates) were collected for phylogenetic dating. Anchors, assumed to be corresponding to the most recent WGD, were detected using i-ADHoRe 3.0 (refs 66,67). Only a low number of duplicated segments and hence anchors could be identified, most likely because of the fragmented assembly of Z. marina. However, the identified anchors did confirm the presence of a broad WGD peak between a KS of 0.8 and 1.6 (data not shown). For each WGD paralogueous pair, an orthogroup was created that included the two paralogues plus several orthologues from other plant species as identified by InParanoid (v. 4.1)68 using a broad taxonomic sampling: one representative orthologue from the order Cucurbitales, two from the Rosales, two from the Fabales, two from the Malpighiales, two from the Brassicales, one from the Malvales, one from the Solanales, two from the Poales, one orthologue from Musa acuminata69 (Zingiberales), and one orthologue from Spirodela polyrhiza11 (Alismatales). In total, about 180 orthogroups from anchor pair duplicates and peak-based duplicates were collected. The node joining the two Z. marina WGD paralogues was then dated using the BEAST v. 1.7 package70 under an uncorrelated relaxed clock model and a LG+G (four rate categories) evolutionary model. A starting tree with branch lengths satisfying all fossil prior constraints was created according to the consensus APGIII phylogeny71. Fossil calibrations were implemented using log-normal calibration priors on the following nodes: the node uniting the Malvidae based on the fossil Dressiantha bicarpellata72 with prior offset = 82.8, mean = 3.8528, and s.d. = 0.5)(ref. 73), the node uniting the Fabidae based on the fossil Paleoclusia chevalieri74 with prior offset = 82.8, mean = 3.9314, and s.d. = 0.5(ref. 75), the node uniting the Alismatales (including Z. marina and Spirodela polyrhiza) with the other monocots based on the oldest fossil monocot pollen, Liliacidites76,77 from the Trent’s Reach locality, with prior offset = 125, mean = 2.0418, and s.d. = 0.5 (refs 14,78) and the root with prior offset = 124, mean = 4.0786, and s.d. = 0.5 (ref. 79). The offsets of these calibrations represent hard minimum boundaries, while their means represent locations for their respective peak mass probabilities in accordance with some of the most recent and taxonomically complete dating studies available for these specific clades14,80. A run without data was performed to ensure proper placement of the marginal calibration prior distributions81. The Markov chain Monte Carlo (MCMC) for each orthogroup was run for 106 generations, sampling every 1,000 generations resulting in a sample size of 104. The resulting trace files of all orthogroups were evaluated manually using Tracer v. 1.570 with a burn-in of 1,000 samples to ensure proper convergence (minimum ESS for all statistics at least 200). In total, 169 orthogroups were accepted and all age estimates for the node uniting the WGD paralogous pairs were then grouped into one absolute age distribution (Fig. 2, too few anchors were available to evaluate them separately from the peak-based duplicates), for which kernel density estimation (KDE) and a bootstrapping procedure were used to find the peak consensus WGD age estimate and its 90% confidence interval boundaries, respectively.
Intra- and inter-genomic co-linearity was investigated (Supplementary Tables 4.2 and 4.3) using MCScanX82 based on a BLASTP search of all genomic protein coding genes with an E-value cut-off of e−10. Only one large duplicated segment was detected, which was most likely due to the fragmented assembly of Z. marina; only 27 scaffolds had a size larger than 1 Mb, accounting for only 23.4% of all protein-coding genes. We therefore additionally used i-ADHoRe (v. 3.0)66 to investigate genomic co-linearity by including all possible scaffolds.
Gene family comparisons
Protein sets were collected for 14 species: Z. marina (ORCAE v. 2.1), Arabidopsis thaliana (TAIR10), Thellungiella parvula (http://thellungiella.org) Populus trichocarpa (Phytozome v. 9.0), Vitis vinifera (Phytozome v. 9.0), Amborella trichopoda (http://amborella.huck.psu.edu), Oryza sativa japonica (Phytozome v. 9.0), Zea mays (Phytozome v. 9.0), Brachypodium distachyon (Phytozome v. 9.0), Spirodela polyrhiza (http://mocklerlab.org), Selaginella moellendorffii (Phytozome v. 9.0), Physcomitrella patens (Phytozome v. 9.0), Chlamydomonas reinhardtii (Phytozome v. 9.0), and Ostreococcus lucimarinus (ORCAE v. 6/3/2013). These species were selected in order to provide a phylogenetic representation traversing green algae, basal plants, monocots, and dicots. Following an ‘all-vresus-all’ TimeLogic Decypher Tera-BLASTP (Active Motif Inc.; e-value threshold 1 × e−3, max hits 500) comparison, OrthoMCL (v. 2.0; mcl inflation factor 3.0)83 was used to delineate gene families. Confidence in establishing gene losses in Zostera was enhanced by using a combination of reciprocal blast, TblastN, re-annotation of Spirodela (and other monocot genes), and careful phylogenetic analysis. OrthoMCL results and related protein resources are available in the ORCAE download section.
To further understand gene family expansion or contraction in Z. marina in comparison with other sequenced genomes, gene family sizes were calculated for all gene families (excluding orphans and species-specific families) (Supplementary Note 4.2). The number of genes per species for each family was transformed into a matrix of z-scores in order to centre and normalize the data. The first 100 families with the largest gene family size in Z. marina were selected. The z-score profile was hierarchically clustered (complete linkage clustering) using Pearson correlation as a distance measure. The functional annotation of each family was predicted based on sequence similarity to entries in the InterProScan and Pfam protein domain database where more than 30% of proteins in the family share the same protein domain. The phylogenetic profile and phylogenetic tree topology provided at PLAZA84 were used to reconstruct the most parsimonious series of gene gain and loss events. The Dollop program from the PHYLIP package85 was used to determine the minimum gene set at ancestral nodes of the phylogenetic tree. The Dollop program is based on the Dollo parsimony principle, which assumes that novel gene families arise exactly once during evolution but can be lost independently in different phylogenetic lineages.
Search for presence/absence of orthologues for specific genes and families
A dedicated search for orthologues/homologues was performed for genes and proteins involved in stomata differentiation (Supplementary Note 5.1), volatile biosynthesis and sensing with focus on ethylene and terpenes (Supplementary Note 6.1), as well as genes involved in male flower specification and pollen differentiation (Supplementary Note 11.1). To this end, queries were chosen from documented genes involved in these pathways (usually from Arabidopsis but occasionally from Oryza, Zea and tomato). Next, the search for homologues in Zostera marina, Spirodela polyrhiza, Oryza sativa japonica and Arabidopsis thaliana (when not used as a query) was performed using BLASTP. To avoid missing or poorly annotated genes a TBLASTN search was conducted using the above queries against the Zostera marina and Spirodela polyrhiza genomes. Putative orthologues were identified based on reciprocal BLASTP searches with Arabidopsis (or the other queries). Owing to species-specific duplications, this may produce some paralogous genes to appear orthologous to the query, or vice versa (see Extended Data Tables 1,2,3). To further confirm correct orthology assignments, phylogenetic trees were built using a broader sampling of protein sequences from both the query species and the three target species. Ambiguously aligned sequences (especially due to indels) were checked manually and corrected or removed.
Accession codes
Primary accessions
BioProject
Gene Expression Omnibus
NCBI Reference Sequence
Sequence Read Archive
Data deposits
Raw reads, the assembled genome sequence and annotation are accessible from NCBI under BioProject number PRJNA41721 with GenBank accession number LFYR00000000. The accession number for the Zostera marina Finnish Clone is BioSample SAMN00991190. Fosmid end sequence: GSS KG963492– KG999999; KO000001– KO144970, whole-genome shotgun data: SRA020075 and RNA-seq: GEO GSE67579. Further information on the Zostera marina project is available via the Online Resource for Community Annotation Eukaryotes (ORCA) at http://bioinformatics.psb.ugent.be/orcae/.
References
Les, D. H., Cleland, M. A. & Waycott, M. Phylogenetic studies in Alismatidae, II: evolution of marine angiosperms (seagrasses) and hydrophily. Syst. Bot. 22, 443–463 (1997)
Larkum, W. D., Orth, R. J. & Duarte, C. M. Seagrasses: Biology, Ecology and Conservation (Springer, Dordrecht, Netherlands, 2006)
Berry, J. A., Beerling, D. J. & Franks, P. J. Stomata: key players in the earth system, past and present. Curr. Opin. Plant Biol . 13, 232–239 (2010)
Aquino, R. S., Landeira-Fernandez, A. M., Valente, A. P., Andrade, L. R. & Mourao, P. A. S. Occurrence of sulfated galactans in marine angiosperms: evolutionary implications. Glycobiology 15, 11–20 (2005)
Franssen, S. U. et al. Transcriptomic resilience to global warming in the seagrass Zostera marina, a marine foundation species. Proc. Natl Acad. Sci. USA 108, 19276–19281 (2011)
Mazzuca, S. et al. Establishing research strategies, methodologies and technologies to link genomics and proteomics to seagrass productivity, community metabolism, and ecosystem carbon fluxes. Front. Plant Sci. 4, 1–19 (2013)
Duarte, C. M. et al. Will the oceans help feed humanity? Bioscience 59, 967–976 (2009)
Costanza, R. et al. The value of the world’s ecosystem services and natural capital. Nature 387, 253–260 (1997)
Fourqurean, J. W. et al. Seagrass ecosystems as a globally significant carbon stock. Nature Geosci. 5, 505–509 (2012)
Green, E. P. & Short, F. T. World Atlas of Seagrasses (University of California Press, Berkeley, CA, USA, 2003)
Wang, W. et al. The Spirodela polyrhiza genome reveals insights into its neotenous reduction fast growth and aquatic lifestyle. Nature Commun . 5, 1–13 (2014)
Chavez Montes, R. A. et al. Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs. Nature Commun . 5, 1–15 (2014)
Vanneste, K., Maere, S. & Van de Peer, Y. Tangled up in two: a burst of genome duplications at the end of the Cretaceous and the consequences for plant evolution. Philos. Trans. R. Soc. B Biol. Sci . 369, 20130353 (2014)
Nauheimer, L., Metzler, D. & Renner, S. S. Global history of the ancient monocot family Araceae inferred with models accounting for past continental positions and previous ranges based on fossils. New Phytol. 195, 938–950 (2012)
Golicz, A. A. et al. Genome-wide survejy of the seagrass Zostera muelleri suggests modification of the ethylene signalling network. J. Exp. Bot. (2015)
Kirk, J. T. O. in Light and Photosynthesis in Aquatic Ecosystems (Cambridge Univ. Press, 2011)
Touchette, B. W. Seagrass-salinity interactions: physiological mechanisms used by submersed marine angiosperms for a life at sea. J. Exp. Mar. Biol. Ecol. 350, 194–215 (2007)
Popper, Z. A. et al. Evolution and diversity of plant cell walls: from algae to flowering plants. Annu. Rev. Plant Biol. 62, 567–590 (2011)
Michel, G., Tonon, T., Scornet, D., Cock, J. M. & Kloareg, B. The cell wall polysaccharide metabolism of the brown alga Ectocarpus siliculosus: insights into the evolution of extracellular matrix polysaccharides in eukaryotes. New Phytol. 188, 82–97 (2010)
Collen, J. et al. Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the Archaeplastida. Proc. Natl Acad. Sci. USA 110, 5247–5252 (2013)
Hanson, S. R., Best, M. D. & Wong, C. H. Sulfatases: structure, mechanism, biological activity, inhibition, and synthetic utility. Angew. Chem. Int. Ed. 43, 5736–5763 (2004)
Larkum, A. W. D., Drew, E. A. & Ralph, P. J. in Seagrasses: Biology, Ecology and Conservation (eds Larkum, A. W. D., Orth, R. J. & Duarte, C. M. ) 323–345 (Springer, Dordrecht, Netherlands, 2006)
De Cock, A. W. Flowering, pollinations and fruiting in Zostera marina L. Aquat. Bot. 9, 201–220 (1980)
Furness, C. A. in Early Events in Monocot Evolution (eds Wilkin, P. & Mayo, S. J. ) 1–22 (Cambridge Univ. Press, 2013)
Kuo, J. & den Hartog, C. in Seagrasses: Biology, Ecology and Conservation (eds Larkum, A. W. D., Orth, R. J. & Duarte, C. M. ) 51–87 (Springer, 2006)
Orth, R. J. et al. A global crisis for seagrass ecosystems. Bioscience 56, 987–996 (2006)
Waycott, M. et al. Accelerating loss of seagrasses across the globe threatens coastal ecosystems. Proc. Natl Acad. Sci. USA 106, 12377–12381 (2009)
Macreadie, P. I., Schliepl, M. T., Rasheed, M. A., Chartrand, K. M. & Ralph, P. J. Molecular indicators of chronic seagrass stress: a new era in the management of seagrass ecosystems? Ecol. Indic. 38, 279–281 (2014)
Vanneste, K., Baele, G., Maere, S. & Van de Peer, Y. Analysis of 41 plant genomes supports a wave of successful genome duplications in association with the Cretaceous–Paleogene boundary. Genome Res. 24, 1334–1347 (2014)
Olsen, J. L. et al. Eelgrass Zostera marina populations in northern Norwegian fjords are genetically isolated and diverse. Mar. Ecol. Prog. Ser. 486, 121–132 (2013)
den Hartog, C., Hennen, J., Noten, T. M. P. A. & Van Wijk, R. J. Chromosome numbers of the European seagrasses. Plant Syst. Evol. 156, 55–59 (1987)
Kuo, J. Chromosome numbers of the Australian Zosteraceae. Plant Syst. Evol. 226, 155–163 (2001)
Reusch, T. B. H. & Bostrom, C. Widespread genetic mosaicism in the marine angiosperm Zostera marina is correlated with clonal reproduction. Evol. Ecol. 25, 899–913 (2010)
Doyle, J. J. & Doyle, J. L. Isolation of plant DNA from fresh tissue. Focus 12, 13–15 (1990)
Jaffe, D. B. et al. Whole-genome sequence assembly for mammalian genomes: Arachne 2. Genome Res. 13, 91–96 (2003)
Smit, A. & Hubley, R. in RepeatModeler Open-1.0 (Repeat Masker Website, http://www.repeatmasker.org/ 2010)
Maumus, F. & Quesneville, H. Deep investigation of Arabidopsis thaliana junk DNA reveals a continuum between repetitive elements and genomic dark matter. PLoS ONE 9, e94101 (2014)
Flutre, T., Duprat, E., Feuillet, C. & Quesneville, H. Considering transposable element diversification in de novo annotation approaches. PLoS ONE 6, e16526 (2011)
Quesneville, H. et al. Combined evidence annotation of transposable elements in genome sequences. PLOS Comput. Biol. 1, e22 (2005)
Hoede, C. et al. PASTEC: an automatic transposable element classification tool. PLoS ONE 9, e91929 (2014)
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnol. 29, 644–652 (2011)
Gouzy, J., Carrere, S. & Schiex, T. FrameDP: sensitive peptide detection on noisy matured sequences. Bioinformatics 25, 670–671 (2009)
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011)
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007)
Zhang, L. et al. A genome-wide characterization of microRNA genes in maize. PLoS Genet. 5, e1000716 (2009)
Jiang, H. & Wong, W. H. SeqMap: mapping massive amount of oligonucleotides to the genome. Bioinformatics 24, 2395–2396 (2008)
Meyers, B. C. et al. Criteria for annotation of plant microRNAs. Plant Cell 20, 3186–3190 (2008)
Addo-Quaye, C., Miller, W. & Axtell, M. J. CleaveLand: a pipeline for using degradome data to find cleaved small RNA targets. Bioinformatics 25, 130–131 (2009)
Degroeve, S., Saeys, Y., De Baets, B., Rouze, P. & Van de Peer, Y. SpliceMachine: predicting splice sites from high-dimensional local context representations. Bioinformatics 21, 1332–1338 (2005)
Foissac, S. et al. Genome annotation in plants and fungi: EuGene as a model platform. Curr. Bioinformatics 3, 87–97 (2008)
Van Bel, M. et al. Dissecting plant genomes with the PLAZA comparative genomics platform. Plant Physiol. 158, 590–600 (2012)
Abeel, T., Van Parys, T., Saeys, Y., Galagan, J. & Van, P. GenomeView: a next-generation genome browser. Nucleic Acids Res. 40, e12 (2012)
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 18, 1979–1990 (2008)
Stanke, M., Tzvetkova, A. & Morgenstern, B. AUGUSTUS at EGASP: using EST, protein and genomic alignments for improved gene prediction in the human genome. Genome Biol. 7, S11 (2006)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997)
Burge, S. W. et al. Rfam 11.0: 10 years of RNA families. Nucleic Acids Res. 41, D226–D232 (2013)
Sterck, L., Billiau, K., Abeel, T., Rouzé, P. & Van der Peer, Y. ORCAE: online resource for community annotation of eukaryotes. Nature Methods 9, 1041 (2012)
Mitchell, A. et al. The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res. 43, D213–D221 (2015)
The UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015)
Vanneste, K., Van de Peer, Y. & Maere, S. Inference of genome duplications from age distributions revisited. Mol. Biol. Evol. 30, 177–190 (2013)
Goldman, N. & Yang, Z. A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol. Biol. Evol. 11, 725–736 (1994)
Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007)
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010)
Proost, S. et al. i-ADHoRe 3.0-fast and sensitive detection of genomic homology in extremely large data sets. Nucleic Acids Res. 40, e11 (2012)
Fostier, J. et al. A greedy, graph-based algorithm for the alignment of multiple homologous gene lists. Bioinformatics 27, 749–756 (2011)
Ostlund, G. et al. InParanoid 7: new algorithms and tools for eukaryotic orthology analysis. Nucleic Acids Res. 38, D196–D203 (2010)
D’Hont, A. et al. The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488, 213–217 (2012)
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. 3408070; Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012)
The Angiosperm Phylogeny Group. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot. J. Linn. Soc. 161, 105–121 (2009)
Gandolfo, M., Nixon, K. & Crepet, W. A new fossil flower from the Turonian of New Jersey: Dressiantha bicarpellata gen. et sp. nov. (Capparales). Am. J. Bot. 85, 964–974 (1998)
Beilstein, M. A., Nagalingum, N. S., Clements, M. D., Manchester, S. R. & Mathews, S. Dated molecular phylogenies indicate a Miocene origin for Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 107, 18724–18728 (2010)
Crepet, W. & Nixon, K. C. Fossil Clusiaceae from the late Cretaceous (Turonian) of New Jersey and implications regarding the history of been pollination. Am. J. Bot. 85, 1122–1133 (1998)
Xi, Z. et al. Phylogenomics and a posteriori data partitioning resolve the Cretaceous angiosperm radiation Malpighiales. Proc. Natl Acad. Sci. USA 109, 17519–17524 (2012)
Doyle, J. A., Endress, P. K. & Upchurch, G. R. Early Cretaceous monocots: a phylogenetic evaluation. Acta Musei Nationalis Pragae, Series B. Historia Naturalis 64, 59–87 (2008)
Iles, W. J. D., Smith, S. Y., Gandolfo, M. A. & Graham, S. W. Monocot fossils suitable for molecular dating analyses. Bot. J. Linn. Soc. 178, 346–374 (2015)
Janssen, T. & Bremer, K. The age of major monocot groups inferred from 800+ rbcL sequences. Bot. J. Linn. Soc. 146, 385–398 (2004)
Smith, S. A., Beaulieu, J. M. & Donoghue, M. J. An uncorrelated relaxed-clock analysis suggests an earlier origin for flowering plants. Proc. Natl Acad. Sci. USA 107, 5897–5902 (2010)
Clarke, J. T., Warnock, R. C. & Donoghue, P. C. Establishing a time-scale for plant evolution. New Phytol. 192, 266–301 (2011)
Heled, J. & Drummond, A. J. Calibrated tree priors for relaxed phylogenetics and divergence time estimation. Syst. Biol. 61, 138–149 (2012)
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012)
Li, L., Stoeckert, C. J. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003)
Proost, S. et al. PLAZA 3.0: an access point for plant comparative genomics. Nucleic Acids Res. 43, D974–D981 (2015)
Felsenstein, J. in PHYLIP: Phylogenetic inference program, version 3.6 (University of Washington, 2005)
Pillitteri, L. J. & Dong, J. Stomatal development in Arabidopsis. Arabidopsis Book 11, e0162 (2013)
Lallemand, B., Erhardt, M., Heitz, T. & Legrand, M. Sporopollenin biosynthetic enzymes interact and constitute a metabolon localized to the endoplasmic reticulum of tapetum cells. Plant Physiol. 162, 616–625 (2013)
Acknowledgements
Genome sequencing, assembly and automated annotation were conducted by the US Department of Energy (DOE) Joint Genome Institute, Walnut Creek California, USA and supported by the Office of Science of the US DOE, Community Sequencing Program award (2009) contract No. DE-AC02-05CH11231 to J.L.O. Further bioinformatics and annotation was supported in part by the Ghent University Multidisciplinary Research Partnership ‘Bioinformatics: From Nucleotides to Networks’ to Y.V.d.P. Y.V.d.P also acknowledges support from the European Union Seventh Framework Programme (FP7/2007-2013) under European Research Council Advanced Grant Agreement 322739–DOUBLE-UP. RNA-seq (Finnish clone genotype) was funded by the Marine Benthic Ecology and Evolution (MarBEE) group, within the former Centre for Ecological and Evolutionary Studies (now Groningen Institute for Evolutionary Life Sciences), University of Groningen to J.L.O. RNA-seq (flower tissues) was funded by the Excellence Cluster, Future Ocean, Kiel to T.B.H.R. Participation of G.P. and E.D. was supported by the MIUR Italian Flagship project RITMARE (NRP 2011-2013). G.A.P. was supported by FCT-EXCL/AAG-GLO/0661/2012. We thank I. D. Gromicho, KAUST, for his artistry in the production of Extended Data Fig. 6. This work also benefited from discussions within the ESSEM COST action ES0906, “Seagrass productivity from genes to ecosystem management” (2009-2014), J.L.O., G.P. and T.B.H.R; and the Linnaeus Centre for Marine Evolutionary Biology (CEMEB)-Tjärnö, Gothenburg University, J.L.O. and M.T. J.L.O. especially thanks K. Johannesson (CeMEB-Tjärnö), C. Boyen (SBR-Roscoff), R. Reinhardt (MPI-Cologne) and E. Serrão (CCMAR-Faro) for their ongoing encouragement, and the more than 70 colleagues who submitted letters of support for the original proposal to the JGI-Community Sequencing Program.
Author information
Authors and Affiliations
Contributions
J.L.O., T.B.H.R., G.P. and Y.V.d.P. are the lead investigators and contributed equally to the work. J.S., J.W.J., J.G., Y.V.d.P., B.V. and Y.-C.L. coordinated the bioinformatics activities surrounding assembly, quality control, set-up and maintenance of Z. marina on the ORCAE site and deposition of the Z. marina genome resource. T.B.H.R and T.B. generated and analysed RNA-seq libraries from flowers, rhizome, roots. J.L.O., Y.-C.L. and A.J. generated and analysed RNA-seq libraries from the genome genotype and temperature stress experiments. C.B., W.T.S. and J.L.O. contributed to biological sample collection, preparation and quality control prior to DNA extraction. A.M. performed the HMW DNA extraction and quality control from the genome genotype/clone. M.A., J.G., H.T. and M.C. contributed to WGS libraries and sequencing, (fosmid)-cloning and quality control. J.G. coordinated the sequencing of FES, quality control projects. Analysis of architectural features of the genome and annotation of specific gene families, including the written contributions to the main paper and Supplementary Information sections, were performed by the following co-authors: J.W.J., the chromosome assembly analysis; B.V. and Y.-C.L., gene family clustering and comparative phylogenomics; A.R.K. and E.B.B., Pfam domains; E.D.P. and P.J.G., miRNA; R.L., K.V. and Y.V.d.P., whole-genome duplication; F.M., Y.-C.L. and Y.V.d.P., transposable elements; B.V., co-linearity and synteny comparisons; M.T., organellar genomes; P.R., stomata gene family; G.M., cell wall polysaccharides and sulfotransferases; T.T., fatty acid metabolism and its relationship to cell walls and ion homeostasis; P.R., volatiles (ethylene, terpenes); P.R., J.B. and T.B.H.R., metallothioniens; P.R., G.A.P. and C.L., osmoregulation/ion homeostasis/stress-related genes; S.D. and E.D., photosynthetic/ light-sensing genes; G.M., CAZymes; T.B., T.B.H.R. and P.R., plant defence-related; T.B. assembly and analysis of MADS box genes (flowering); P.R.; Y.V.d.P. and Y.-C.L., pollen-related and self-incompatibility genes; F.M., SLR-1gene and core eukaryotic genes analysis (CEGMA). J.L.O., Y.V.d.P., T.B.H.R., C.M.D., Y.-C.L. and P.R. wrote and edited the main manuscript (including the Methods and Extended Data), and organized and further edited the individual contributions (as listed above) for the Supplementary Information sections. J.L.O. and Y.V.d.P. provided the overall evolutionary context and T.B.H.R., G.P. and C.M.D. provided the ecological and societal context. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Number of genes expressed in five tissues of Z. marina.
a, Venn diagram of genes with expression values (FPKM) higher than 1 are considered as expressed in the tissue. b, Pairwise differential gene expression analysis between tissues. The male flower shows the highest number of differentially expressed genes.
Extended Data Figure 2 Circos plot of the ten largest scaffolds of Z. marina.
Tracks from outside to inside. GC percentage, gene density, and transposable element (TE) density (density measured in 20-Kb sliding windows and gene expression profiles from five tissues (root, leaf, male flower, female flower early and female flower late) presented as log2 FPKM values.
Extended Data Figure 3 Potential impact of transposable elements (TEs) on Z. marina evolution.
a, Frequency distribution of pairwise sequence identity values between copies of Copia- and Gypsy-type LTR retrotransposons and DNA transposons, and their cognate consensus sequences (younger repeats share higher sequence similarity). Two peaks are detectable for Copia-type elements. b, Distance to the closest TE for the set of Z. marina single-copy genes and the set of Z. marina accessory genes. TE-proximal accessory genes are more frequent than TE-proximal single-copy genes. c, Frequency of pairwise sequence identity between accessory gene-proximal Ty3-Gypsy elements and their cognate consensus sequences. A number of high-identity copies (that is, putatively young duplicate genes) is observed.
Extended Data Figure 4 Unrooted maximum likelihood tree of genes encoding light-harvesting complex A (LHCA) and LHCB proteins of Z.marina, Spirodela polyrhiza and Arabidopsis thaliana.
The analysis was carried out on protein sequences using PhyML 3 with LG substitution model and 100 bootstrap replicates. Supplementary Note 7.1, Supplementary Table 7.3.
Extended Data Figure 5 Alignment of metallothionein (MT) and half-metallothionein (HMT) genes in Z. marina as compared with other plants.
Alignments were performed in ClustalW on the Lyon PBIL web server and edited manually. The upper alignments are for type 1–3 MTs and HMTs; the lower alignment is for type 4 EcMTs where there is no Zostera homologue. Conserved residues are shown in red and residues in the same amino acid group in blue. Cys and His residues, putatively involved in binding metals, are highlighted in green and yellow, respectively. Aromatic amino acids absent in canonical animal MTs are highlighted in grey. MTs and MT-like proteins were obtained from: Arabidopsis thaliana (ARATH), Japanese rice (ORYSJ), Cicer arietinum (CICAR), banana (MUSAC), wheat (WHEAT), potato (SOLTU), Setaria Italica (SETIT), Vitis vinifera (VITVI) and the alismatids: Posidonia oceanica (POSOC) highlighted in grey, Spirodela polyrhiza (SPIPO) highlighted in blue, and Zostera marina (ZOSMA) highlighted in yellow. See Supplementary Note 8.2.
Extended Data Figure 6 Conceptual summary of physiological and structural adaptations made by Z. marina in its return to the sea.
Ecosystem services shown in blue. Physical processes related to salinity, light and CO2 availability shown in white within light-green boxes. Gene losses and gains associated with morphological and physiological processes shown in white within the dark-green box on the right.
Supplementary information
Supplementary information
This file contains Supplementary Text, Supplementary Figures and Supplementary Tables – see contents page for details. (PDF 16712 kb)
Supplementary Data
This zipped file contains Supplementary Datasets 1-8. (ZIP 20389 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Olsen, J., Rouzé, P., Verhelst, B. et al. The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea. Nature 530, 331–335 (2016). https://doi.org/10.1038/nature16548
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16548
This article is cited by
-
A two-sequence motif-based method for the inventory of gene families in fragmented and poorly annotated genome sequences
BMC Genomics (2024)
-
Population genomics unveils the century-old invasion of the Seagrass Halophila stipulacea in the Mediterranean Sea
Marine Biology (2024)
-
Mangrove species found in contrasting environments show differing phytohormonal responses to variation in soil bulk density
Plant and Soil (2024)
-
Seagrass genomes reveal ancient polyploidy and adaptations to the marine environment
Nature Plants (2024)
-
Combined transcriptome and proteome analysis reveal the key physiological processes in seed germination stimulated by decreased salinity in the seagrass Zostera marina L.
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.