Abstract
The gut is home to trillions of microorganisms that have fundamental roles in many aspects of human biology, including immune function and metabolism1,2. The reduced diversity of the gut microbiota in Western populations compared to that in populations living traditional lifestyles presents the question of which factors have driven microbiota change during modernization. Microbiota-accessible carbohydrates (MACs) found in dietary fibre have a crucial involvement in shaping this microbial ecosystem, and are notably reduced in the Western diet (high in fat and simple carbohydrates, low in fibre) compared with a more traditional diet3. Here we show that changes in the microbiota of mice consuming a low-MAC diet and harbouring a human microbiota are largely reversible within a single generation. However, over several generations, a low-MAC diet results in a progressive loss of diversity, which is not recoverable after the reintroduction of dietary MACs. To restore the microbiota to its original state requires the administration of missing taxa in combination with dietary MAC consumption. Our data illustrate that taxa driven to low abundance when dietary MACs are scarce are inefficiently transferred to the next generation, and are at increased risk of becoming extinct within an isolated population. As more diseases are linked to the Western microbiota and the microbiota is targeted therapeutically, microbiota reprogramming may need to involve strategies that incorporate dietary MACs as well as taxa not currently present in the Western gut.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Accessions
Sequence Read Archive
Data deposits
The 16S sequence data have been deposited in the Sequence Read Archive (SRA) under the accession PRJNA303185.
References
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012)
Karlsson, F., Tremaroli, V., Nielsen, J. & Backhed, F. Assessing the human gut microbiota in metabolic diseases. Diabetes 62, 3341–3349 (2013)
Sonnenburg, E. D. & Sonnenburg, J. L. Starving our microbial self: the deleterious consequences of a diet deficient in microbiota-accessible carbohydrates. Cell Metab. 20, 779–786 (2014)
Schnorr, S. L. et al. Gut microbiome of the Hadza hunter-gatherers. Nature Commun. 5, 3654 (2014)
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012)
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl Acad. Sci. USA 107 , 14691–14696 (2010)
Clemente, J. C. et al. The microbiome of uncontacted Amerindians. Science Advances 1 , e1500183 (2015)
Obregon-Tito, A. J. et al. Subsistence strategies in traditional societies distinguish gut microbiomes. Nature Commun. 6, 6505 (2015)
Martinez, I. et al. The gut microbiota of rural Papua New Guineans: composition, diversity patterns, and ecological processes. Cell Rep. 11, 527–538 (2015)
McGill, C. R., Fulgoni, V. L. III & Devareddy, L. Ten-year trends in fiber and whole grain intakes and food sources for the United States population: National Health and Nutrition Examination Survey 2001–2010. Nutrients 7, 1119–1130 (2015)
King, D. E., Mainous, A. G. III & Lambourne, C. A. Trends in dietary fiber intake in the United States, 1999–2008. J. Acad. Nutr. Diet. 112, 642–648 (2012)
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012)
Kashyap, P. C. et al. Genetically dictated change in host mucus carbohydrate landscape exerts a diet-dependent effect on the gut microbiota. Proc. Natl Acad. Sci. USA 110, 17059–17064 (2013)
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014)
Tikhonov, M., Leach, R. W. & Wingreen, N. S. Interpreting 16S metagenomic data without clustering to achieve sub-OTU resolution. ISME J. 9, 68–80 (2015)
El Kaoutari, A., Armougom, F., Gordon, J. I., Raoult, D. & Henrissat, B. The abundance and variety of carbohydrate-active enzymes in the human gut microbiota. Nature Rev. Microbiol. 11, 497–504 (2013)
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010)
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005)
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature Methods 7, 335–336 (2010)
Kashyap, P. C. et al. Complex interactions among diet, gastrointestinal transit, and gut microbiota in humanized mice. Gastroenterology 144, 967–977 (2013)
Turnbaugh, P. J. et al. The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 1, 6ra14 (2009)
Bokulich, N. A. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nature Methods 10, 57–59 (2013)
Yin, Y. et al. dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 40, W445–W451 (2012)
Marchler-Bauer, A. et al. CDD: Specific functional annotation with the Conserved Domain Database. Nucleic Acids Res. 37, D205–D210 (2009)
Aspeborg, H., Coutinho, P. M., Wang, Y., Brumer, H. & Henrissat, B. Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 5 (GH5). BMC Evol. Biol. 12, 186 (2012)
Stam, M. R., Danchin, E. G. J., Rancurel, C., Coutinho, P. M. & Henrissat, B. Dividing the large glycoside hydrolase family 13 into subfamilies: towards improved functional annotations of α-amylase-related proteins. Protein Eng. Des. Sel. 19, 555–562 (2006)
Acknowledgements
We thank M. St. Onge for technical assistance. This work was funded by a grant from National Institutes of Health NIDDK (R01-DK085025 to J.L.S.), an NSF graduate fellowship (to S.A.S.), a Stanford Graduate Fellowship (to S.A.S.), and the Simons Foundation (to M.T.). J.L.S. holds an Investigators in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund.
Author information
Authors and Affiliations
Contributions
E.D.S. and J.L.S. conceived and designed the project. E.D.S., J.L.S. and S.K.H. designed and supervised the experiments. E.D.S. and S.K.H. performed the experiments. E.D.S., S.A.S. and M.T. analysed the experimental data. N.S.W. designed and supervised data analysis. E.D.S. and S.A.S. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Collating data from studies of the microbiota of hunter–gatherers in Tanzania, agrarians from Malawi and Venezuela, and Westerners from the United States reveals that Western populations have depleted alpha-diversity from birth through childbearing years and are missing bacterial taxa present in the traditional groups.
a, Scatterplot of faecal microbiota of individuals plotted by phylogenetic diversity against age of the Hadza hunter–gatherers from Tanzania (n = 16, green), agrarians from Malawi (n = 81, red) and Venezuela (n = 78, purple) and Americans (n = 213, blue) b, Individuals plotted by unweighted UniFrac PC1 versus phylogenetic diversity. c, Individuals plotted by unweighted UniFrac PC1 versus age. d, Line plot of unique OTUs from faecal microbiota across populations (Americans, n = 315; Malawi and Venezuela, n = 213; Tanzania, n = 27). OTUs (x axis; black, present; white, absent) are considered present if represented by ≥0.001% of reads within each population. OTUs were sorted along the x axis by their relative abundance in the US and Tanzanian populations and further subdivided by their distributions within a population into tracks (red >0.05%, yellow ≤0.05%, and green ≤0.01%, relative abundance). The opacity of the line is the proportion of that population that meets the criteria for that respective track.
Extended Data Figure 2 Comparison of human donor and humanized mice.
a, Taxa summary plot of the relative abundance of taxa from humanized mice faeces (mice) (n = 10) and human donor faeces (human) (n = 1). b, Alpha-diversity of the faecal microbiota from humanized mice (mice) and human donor (human) expressed as number of OTUs (top) and phylogenetic diversity (bottom). Error bars are s.e.m.
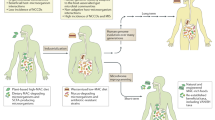
Extended Data Figure 3 Detailed schematic of multigeneration experiment.
Generation one: humanized mice were fed a high-MAC diet for 4 weeks then switched to a low-MAC diet. One week after diet switch, the mice were bred to generate a litter of pups. After three additional weeks on the low-MAC diet, generation-two pups were born and remained in the cage with their mother, suckling for 3 weeks (generation one still consuming the low-MAC diet). After pups were weaned, generation-one mice were returned to the high-MAC diet for 6 weeks. Generation two: pups were weaned from their mother at 3 weeks old onto a low-MAC diet, which they consumed for 10 weeks. Breeding pairs for generation-two mice were set-up at 7 weeks old. After three additional weeks on the low-MAC diet, generation-three pups were born and remained in the cage with their mother, suckling for 3 weeks (generation-two mice still consuming the low-MAC diet). After pups were weaned, generation-two mice were returned to the high-MAC diet for 6 weeks. Generations three and four followed the same protocol as generation two described above.
Extended Data Figure 4 Microbiota diversity is not regained after direct weaning the diet-switching group onto the high-MAC diet.
a, Alpha-diversity as measured by Shannon index of faecal microbiota from generation-five mice from the high-MAC-diet control (control) (n = 6), generation-five diet-switching group that was weaned directly onto the high-MAC diet (Gen 5 diet switching) (n = 6), and generation-four mice from the diet-switching group after weaning and maintenance on the low-MAC diet for 13 weeks and returned to the high-MAC diet for 4 weeks (Gen 4 diet switching) (n = 5). Error bars are s.e.m. and P values are from a two-tailed Student’s t-test b, Principal coordinate analysis of unweighted UniFrac distance for 16S rRNA amplicon profiles from faecal samples collected from first-generation control mice on a high-MAC diet (green), fourth-generation diet-switching mice (purple), and fifth-generation mice from the diet-switching lineage weaned directly onto the high-MAC diet (orange). Control is plotted as weeks post-humanization and generations four and five are plotted as age.
Extended Data Figure 5 Fraction of high-confidence OTUs from the Clostridiales order increases and from the Bacteroidales order decreases over several generations in the low-MAC-consuming mice.
a, Percentage of high-confidence OTUs, grouped by order, detected in mice faeces over four generations in the diet-switching lineage on the low-MAC diet (lo) and high-MAC diet (hi) (n = 5 for Gen 1; n = 6 for Gen 2–4). b, Percentage of high-confidence OTUs, grouped by order, detected in mice faeces over four generations in the control high-MAC diet lineage at the equivalent time points to the high-MAC diet (a) and low-MAC diet (b) of the diet-switching group (n = 5 for Gen 1; n = 6 for Gen 2–4). c, Imputed gycloside hydrolase (GH) family members that show significant differences (at least twofold change and P < 0.05, Bonferroni-corrected t-test) between generation-four diet-switching mice after 4 weeks on the high-MAC diet (teal) (n = 5) and the starting generation-one mice (salmon) (n = 10). Error bars depict s.e.m. No glycoside hydrolase families showed significant changes in the control group.
Extended Data Figure 6 Inefficient inter-generational transfer of taxa driven to low abundance by low dietary MACs.
Heat map of abundance of high-confidence sub-OTUs (number of sequencing reads, columns) from faeces of the diet-switching (top) and control (bottom) group. Each row represents an individual mouse faecal microbiota from 4 weeks post-humanization (initial), while consuming the low-MAC diet (week 9, lo, shaded yellow), and 4 weeks after switching to the high-MAC diet (week 15, hi, shaded grey). Corresponding time points from controls are also shaded. Top row shows the taxonomic assignment for the OTUs plotted: Bacteroidetes are green, Firmicutes are orange, and others are grey.
Extended Data Figure 7 Reintroduction of lost taxa and a high-MAC diet restores microbiota diversity and composition with Clostridiales order decreasing and Bacteroidales order increasing in low-MAC-consuming mice that receive a faecal transplant.
a, Plot of percentage representation of high-confidence OTUs from generation-four mice faeces in the diet-switching group at day 0 before the FMT (starting) (n = 6) and then 3–14 days no-FMT control (n = 3) or post-FMT (n = 3). FMT donor is plotted on the right. b, Heat map of abundance of high-confidence sub-OTUs (number of sequencing reads, columns) from the faeces of the diet-switching group at day 0 (Gen 4), days 3–14 that did not receive an FMT (control) (n = 3 for each day), days 3–14 that received an FMT (+FMT), and the FMT donor. Each row represents an individual mouse faecal microbiota. Top row shows the taxonomic assignment for the OTUs plotted: Bacteroidetes are green, Firmicutes are orange, and others are grey.
Supplementary information
Supplementary Table 1
A table of high-confidence OTUs that dropped in abundance by at least two-fold in the first generation diet-switching mice after switch to the low-MAC diet and the control high-MAC diet mice. (XLSX 15 kb)
Supplementary Table 2
A table of high-confidence OTUs that dropped in abundance by at least two-fold in the first generation diet-switching mice after return to the high-MAC diet and the control high-MAC diet mice. (XLSX 12 kb)
Supplementary Table 3
A table of high-confidence OTUs lost through generation four while consuming the low MAC diet and after switch to the high MAC diet. (XLSX 9 kb)
Supplementary Table 4
A comparison of glycoside hydrolases from metagenomic data and imputed from 16S rRNA amplicon sequencing data. (XLSX 48 kb)
Supplementary Table 5
Table of fold change of glycoside hydrolase families from generation one to generation four in the diet-switching and control groups. (XLSX 43 kb)
Supplementary Table 6
Table of unique glycoside hydrolase sub-families in diet-switching mice between generations one and four by sampling depth. (XLSX 49 kb)
Supplementary Table 7
Table of low abundance taxa that were passed or not passed between each of four generations of mice. (XLSX 11 kb)
Supplementary Table 8
Table of number of high-confidence sub-OTUs present at the start of the experiment and the number of high-confidence sub-OTUs lost by the fourth generation and between the third and fourth generation. (XLSX 9 kb)
Supplementary Table 9
Table of high-confidence OTUs that were no longer detectable after four generations on the low MAC and return after FMT. (XLSX 11 kb)
Supplementary Table 10
Table of high-confidence sub-OTUs that were no longer detectable after four generations on the low MAC and return after FMT. (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Sonnenburg, E., Smits, S., Tikhonov, M. et al. Diet-induced extinctions in the gut microbiota compound over generations. Nature 529, 212–215 (2016). https://doi.org/10.1038/nature16504
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16504
This article is cited by
-
Stable composition of gut microbiome in the Asian ladybeetle Coccinella septempunctata reared on natural and artificial diets
Scientific Reports (2024)
-
Gut microbial CAZymes markers for depression
Translational Psychiatry (2024)
-
Differential peripheral immune signatures elicited by vegan versus ketogenic diets in humans
Nature Medicine (2024)
-
The diet rapidly and differentially affects the gut microbiota and host lipid mediators in a healthy population
Microbiome (2023)
-
Gut microbiome transitions across generations in different ethnicities in an urban setting—the HELIUS study
Microbiome (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.