Abstract
Insulin/insulin-like growth factor signalling (IIS) is a critical regulator of an organism’s most important biological decisions from growth, development, and metabolism to reproduction and longevity. It primarily does so through the activity of the DAF-16 transcription factor (forkhead box O (FOXO) homologue), whose global targets were identified in Caenorhabditis elegans using whole-worm transcriptional analyses more than a decade ago1. IIS and FOXO also regulate important neuronal and adult behavioural phenotypes, such as the maintenance of memory2 and axon regeneration3 with age, in both mammals4 and C. elegans, but the neuron-specific IIS/FOXO targets that regulate these functions are still unknown. By isolating adult C. elegans neurons for transcriptional profiling, we identified both the wild-type and IIS/FOXO mutant adult neuronal transcriptomes for the first time. IIS/FOXO neuron-specific targets are distinct from canonical IIS/FOXO-regulated longevity and metabolism targets, and are required for extended memory in IIS daf-2 mutants. The activity of the forkhead transcription factor FKH-9 in neurons is required for the ability of daf-2 mutants to regenerate axons with age, and its activity in non-neuronal tissues is required for the long lifespan of daf-2 mutants. Together, neuron-specific and canonical IIS/FOXO-regulated targets enable the coordinated extension of neuronal activities, metabolism, and longevity under low-insulin signalling conditions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Data deposits
Sequencing reads are deposited at NCBI BioProject under accession number PRJNA297798.
References
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans . Nature 424, 277–283 (2003)
Kauffman, A. L., Ashraf, J. M., Corces-Zimmerman, M. R., Landis, J. N. & Murphy, C. T. Insulin signaling and dietary restriction differentially influence the decline of learning and memory with age. PLoS Biol. 8, e1000372 (2010)
Byrne, A. B. et al. Insulin/IGF1 signaling inhibits age-dependent axon regeneration. Neuron 81, 561–573 (2014)
Sjöberg, J. & Kanje, M. Insulin-like growth factor (IGF-1) as a stimulator of regeneration in the freeze-injured rat sciatic nerve. Brain Res. 485, 102–108 (1989)
Wolkow, C. A., Kimura, K. D., Lee, M. S. & Ruvkun, G. Regulation of C. elegans life-span by insulinlike signaling in the nervous system. Science 290, 147–150 (2000)
Libina, N., Berman, J. R. & Kenyon, C. Tissue-specific activities of C. elegans DAF-16 in the regulation of lifespan. Cell 115, 489–502 (2003)
Luo, S., Kleemann, G. A., Ashraf, J. M., Shaw, W. M. & Murphy, C. T. TGF-β and insulin signaling regulate reproductive aging via oocyte and germline quality maintenance. Cell 143, 299–312 (2010)
Tepper, R. G. et al. PQM-1 complements DAF-16 as a key transcriptional regulator of DAF-2-mediated development and longevity. Cell 154, 676–690 (2013)
Zhang, P., Judy, M., Lee, S.-J. & Kenyon, C. Direct and indirect gene regulation by a life-extending FOXO protein in C. elegans: roles for GATA factors and lipid gene regulators. Cell Metab. 17, 85–100 (2013)
Murakami, H., Bessinger, K., Hellmann, J. & Murakami, S. Aging-dependent and -independent modulation of associative learning behavior by insulin/insulin-like growth factor-1 signal in Caenorhabditis elegans . J. Neurosci. 25, 10894–10904 (2005)
Tank, E. M. H., Rodgers, K. E. & Kenyon, C. Spontaneous age-related neurite branching in Caenorhabditis elegans . J. Neurosci. 31, 9279–9288 (2011)
Toth, M. L. et al. Neurite sprouting and synapse deterioration in the aging Caenorhabditis elegans nervous system. J. Neurosci. 32, 8778–8790 (2012)
Chikina, M. D., Huttenhower, C., Murphy, C. T. & Troyanskaya, O. G. Global prediction of tissue-specific gene expression and context-dependent gene networks in Caenorhabditis elegans . PLOS Comput. Biol. 5, e1000417 (2009)
Ewald, C. Y., Landis, J. N., Abate, J. P., Murphy, C. T. & Blackwell, T. K. Dauer-independent insulin/IGF-1-signalling implicates collagen remodelling in longevity. Nature 519, 97–101 (2015)
Zhang, S., Banerjee, D. & Kuhn, J. R. Isolation and culture of larval cells from C. elegans . PLoS ONE 6, e19505 (2011)
Spencer, W. C. et al. Isolation of specific neurons from C. elegans larvae for gene expression profiling. PLoS ONE 9, e112102 (2014)
Schafer, J. C. et al. IFTA-2 is a conserved cilia protein involved in pathways regulating longevity and dauer formation in Caenorhabditis elegans . J. Cell Sci. 119, 4088–4100 (2006)
Apfeld, J. & Kenyon, C. Regulation of lifespan by sensory perception in Caenorhabditis elegans . Nature 402, 804–809 (1999)
Hansen, M., Hsu, A.-L., Dillin, A. & Kenyon, C. New genes tied to endocrine, metabolic, and dietary regulation of lifespan from a Caenorhabditis elegans genomic RNAi screen. PLoS Genet. 1, 119–128 (2005)
Shaw, W. M., Luo, S., Landis, J., Ashraf, J. & Murphy, C. T. The C. elegans TGF-β Dauer pathway regulates longevity via insulin signaling. Curr. Biol. 17, 1635–1645 (2007)
Lakhina, V. et al. Genome-wide functional analysis of CREB/long-term memory-dependent transcription reveals distinct basal and memory gene expression programs. Neuron 85, 330–345 (2015)
Chen, L. et al. Axon regeneration pathways identified by systematic genetic screening in C. elegans . Neuron 71, 1043–1057 (2011)
Tian, N. M., Pratt, T. & Price, D. J. Foxg1 regulates retinal axon pathfinding by repressing an ipsilateral program in nasal retina and by causing optic chiasm cells to exert a net axonal growth-promoting activity. Development 135, 4081–4089 (2008)
Chang, Y.-W. et al. Rapid induction of genes associated with tissue protection and neural development in contused adult spinal cord after radial glial cell transplantation. J. Neurotrauma 26, 979–993 (2009)
Ikeda, D. D. et al. CASY-1, an ortholog of calsyntenins/alcadeins, is essential for learning in Caenorhabditis elegans . Proc. Natl Acad. Sci. USA 105, 5260–5265 (2008)
Hoerndli, F. J. et al. A conserved function of C. elegans CASY-1 calsyntenin in associative learning. PLoS ONE 4, e4880 (2009)
Ohno, H. et al. Role of synaptic phosphatidylinositol 3-kinase in a behavioral learning response in C. elegans . Science 345, 313–317 (2014)
Ewald, C. Y. et al. Pan-neuronal expression of APL-1, an APP-related protein, disrupts olfactory, gustatory, and touch plasticity in Caenorhabditis elegans . J. Neurosci. 32, 10156–10169 (2012)
Hammarlund, M., Nix, P., Hauth, L., Jorgensen, E. M. & Bastiani, M. Axon regeneration requires a conserved MAP kinase pathway. Science 323, 802–806 (2009)
Wong, A. K., Krishnan, A., Yao, V., Tadych, A. & Troyanskaya, O. G. IMP 2.0: a multi-species functional genomics portal for integration, visualization and prediction of protein functions and networks. Nucleic Acids Res. 43, W128–W133 (2015)
Tomioka, M. et al. The insulin/PI 3-kinase pathway regulates salt chemotaxis learning in Caenorhabditis elegans . Neuron 51, 613–625 (2006)
Kodama, E. et al. Insulin-like signaling and the neural circuit for integrative behavior in C. elegans . Genes Dev. 20, 2955–2960 (2006)
Murakami, H., Bessinger, K., Hellmann, J. & Murakami, S. Aging-dependent and -independent modulation of associative learning behavior by insulin/insulin-like growth factor-1 signal in Caenorhabditis elegans . J. Neurosci. 25, 10894–10904 (2005)
Stein, G. M. & Murphy, C. T. C. elegans positive olfactory associative memory is a molecularly conserved behavioral paradigm. Neurobiol. Learn. Mem. 115, 86–94 (2014)
Acknowledgements
We thank the C. elegans Genetics Center for strains; WormBase (WS250); L. Parsons for RNA-seq data support; J. Wiggins and the Lewis-Sigler High Throughput Sequencing Core Facility for RNA-seq support; C. DeCoste and the Flow Cytometry Facility for assistance; V. Yao for tissue prediction analysis; R. DiLoreto for chemotaxis assay analysis; and Z. Gitai, the Murphy lab, and W. Mair for discussion. Funding was provided by a Keck Scholars Program fellowship (C.T.M.), National Institutes of Health Cognitive Aging R01 (C.T.M.), Ruth L. Kirschstein National Research Service Awards (R.K., R.A.), National Science Foundation (J.L.) and New Jersey Commission on Brain Injury Research (V.L.) fellowships.
Author information
Authors and Affiliations
Contributions
C.T.M., R.K., V.L. and J.L. designed experiments. R.K., V.L., R.A., J.L., J.A., and C.T.M. performed experiments and analysed data. R.K., V.L. and J.L. performed tissue isolation and RNA-seq. A.W., R.K., V.L. and J.L. performed bioinformatics analysis. R.K., V.L. and R.A. performed short-term memory experiments. V.L. performed axon regeneration experiments. R.K., V.L. and C.T.M. wrote the manuscript. R.A. and A.W. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Tissue-specific rescue of DAF-16 activity in daf-16;daf-2 mutants identifies distinct gene expression profiles.
a, DAF-16 tissue-specific transgenics; heatmap of all genes with expression differences ≥1.5-fold in ≥3 arrays. b, Significant gene ontology (GO) cluster terms from Punc-119::daf-16-regulated up- and downregulated genes (enrichment score >1). c, Pairwise Pearson correlations between arrays of DAF-16-upregulated or downregulated targets. The red box highlights the negative correlation between neuronal DAF-16 rescued targets (Punc-119::daf-16::gfp;daf-16;daf-2 vs daf-16;daf-2) and intestinal DAF-16 targets (Pges-1::daf-16::gfp;daf-16;daf-2 vs daf-16;daf-2), while the blue box shows the positive correlation between intestinal DAF-16 targets (Pges-1::daf-16::gfp;daf-16;daf-2 vs daf-16;daf-2) and whole-worm DAF-16 targets (Pdaf-16::daf-16::gfp; daf-16;daf-2 vs daf-16;daf-2). The green box shows the weak correlation between neuronal rescued and whole-worm DAF-16 targets. d, Tissue enrichment analysis (mean ± s.e.m.) of significant DAF-16-rescued up- and downregulated genes (Supplementary Table 1) (FDR <0.5). e, Significant GO terms (adjusted P value < 0.05) for DAF-16 upregulated and downregulated genes from whole worm, intestine-, neuron- and muscle-rescued DAF-16 strains. Genes used for GO analysis (Supplementary Table 2) were derived from SAM analysis of the microarrays in a and Supplementary Table 1.
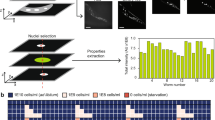
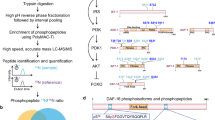
Extended Data Figure 2 Protocol for isolating neuron-specific targets using FACS followed by RNA-sequencing.
a, Pipeline for isolation of adult cells for FACS and RNA sequencing. b, Workflow for RNA-seq data analysis of isolated neurons. c, Heatmap of wild-type neuron-expressed relative to whole-worm-expressed genes. d, Actinomycin D (transcription inhibitor) treatment (100 μg ml−1) during the cell isolation process demonstrates that the neuron isolation technique induces minimal transcriptional changes in wild type animals. Gene ontology (GO) terms represent genes upregulated in the absence of actinomycin D (Fig. 1b, Supplementary Table 4). e, The 26 differentially expressed genes from actinomycin D treatment are listed. f, C. elegans tissue gene expression prediction confirms neuronal character of adult wild-type neuron-enriched genes. Neuron-enriched genes were divided among equal bins according to P value. Bin 1: FDR <0.003%; bin 2: 0.003–0.03%; bin 3: 0.03–1.3%; bin 4: 1.3–4%; bin 5: 4–10%. g, Principal component analysis (PCA) shows a clear separation between wild-type adult neuronal and whole-worm samples. h, Downsampling of wild-type neuron sequencing reads demonstrates sufficient sampling depth. The number of genes detected at the 3 counts per million threshold (for expressed genes) with different proportions of total sequencing depth analysed.
Extended Data Figure 3 Neuron-expressed genes identified by our method are confirmed to be expressed in adults and have adult neuronal functions.
a, Promoter–GFP transcriptional fusions of candidate uncharacterized neuronal genes (day 1 of adulthood). b, Gene ontology clusters were generated from the categories in Fig. 1e. Non-overlapping GO terms suggest a transition from development-related processes in embryonic and larval animals to neuronal processes involved in behaviour in adults (Supplementary Table 5). c, Venn diagram depicting the overlap between genes classified as “expressed” among embryonic and larval neurons16 and adult neurons from our RNA-seq analysis (Supplementary Table 5).
Extended Data Figure 4 Comparison of neuronal DAF-16 targets with wild-type neuronal targets and whole-worm DAF-16 targets.
a, Principal component analysis of the whole worm and isolated adult neuron samples obtained for this study. b, Venn diagram depicting the overlap of daf-2- and daf-16;daf-2-expressed genes with those expressed in wild-type adult neurons. c, Spearman correlation of whole-worm and isolated adult neuron samples. d, The DAF-16 cell-autonomous and cell-non-autonomous targets are distinct. The number of genes that overlap between neuronal DAF-16-rescued whole-worm targets (Punc-119::daf-16::gfp;daf-16;daf-2 vs daf-16;daf-2) and isolated neuron IIS targets (daf-2 vs daf-16;daf-2) is shown (Supplementary Table 8). Hypergeometric distribution analysis (P values) shows that the extent of overlap between the gene categories is not significant.
Extended Data Figure 5 Promoter analysis and gene ontology term analysis of neuronal IIS/FOXO genes.
a, The different classes of neuronal IIS/FOXO genes shown in Fig. 2b were analysed for DBE and DAE sequences in the 1 kb upstream promoter regions. The genome-wide percentage of DBE and DAE occurrences across the 1 kb promoters of all gene-encoding regions is reported. Comparison of whole-worm (Class I)8 vs neuronal-IIS/FOXO-regulated targets. P values: hypergeometric distributions. b, GO terms of Class I whole worm8 vs neuronal-IIS upregulated genes (left) and Class II whole worm8 vs neuronal-IIS downregulated genes (right) (Supplementary Table 5).
Extended Data Figure 6 Short-term associative memory phenotypes obtained upon knocking down neuronal IIS genes in daf-2 mutants and wild-type animals.
daf-2 is required for various forms of C. elegans associative learning2,27,31,32,33,34. daf-16 is required for the improvements and extensions of abilities with age of daf-2 mutants2. daf-2 mutants are defective for salt chemotaxis learning27,31,32, and daf-16 is not involved in salt chemotaxis learning27,31,32. Furthermore, salt learning utilizes a unique daf-2c isoform27 in a daf-16-independent manner31, suggesting a learning mechanism distinct from the associative memory paradigms studied here. We are specifically interested in understanding how activation of DAF-16 results in the improved and extended abilities of daf-2 mutants to carry out olfactory associative learning2, short-term associative memory2,34, and long-term associative memory2, all of which require daf-16. a, Chemotaxis index profile of wild type (N2) and daf-2 animals at time points following memory training. b, RNAi knockdown of sod-3, a non-neuronal DAF-16-regulated target that influences lifespan, has no effect on the extended short-term associative memory (STAM) of daf-2 mutants when treated with RNAi-feeding bacteria throughout the whole life (b) or only the post-developmental (adult-only) period (c, d) of the animal. daf-2 worms treated with daf-16 RNAi have defective STAM, as previously reported2. e, Knockdown of the neuronal IIS candidate genes zip-5 and best-23 does not affect STAM. Time-courses showing the chemotaxis index for each time point are shown in d and e. Learning indices are shown in b, c, f and g. b–e, Two-way repeated measures ANOVA, Bonferroni post hoc tests. f, Treatment of daf-2 worms with neuronal DAF-16 target RNAi does not affect short-term associative learning. g, Neuronal-RNAi sensitive worms (Punc-119::sid-1) in a wild-type background were treated only during adulthood with RNAi targeted against the neuronal DAF-16 target genes. Learning (0 h) and 1 h short-term associative memory time points are shown. a–g, Mean ± s.e.m., *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Figure 7 Characterization of age-dependent axon regeneration and structural defects upon fkh-9 overexpression in mechanosensory neurons.
a, Six adult mechanosensory neurons labelled by mec-4p::GFP were isolated for RNA-seq. b, Axon length from the cell body to the site of injury was measured in μm immediately after axotomy and 24 h later. Regenerative capacity of wild-type PLM axons declines from day 1 to day 5 of adulthood. c, Day 5 wild-type animals regrow axons that are significantly shorter than in day 1 animals. d, Axotomies of daf-2 mutants grown on vector control, sod-3, or daf-16 RNAi demonstrate that sod-3, a lifespan-regulating DAF-16 target, does not influence the axon regeneration capacity of daf-2 worms at day 5 of adulthood. e, fkh-9 does not affect the regenerative capacity of daf-2 axons on day 1 of adulthood. f, fkh-9 is not required for axon regeneration in day 1 adults. b–f, Mean ± s.e.m., Fisher’s exact test, *P < 0.05. g, Overexpression of the a and b isoforms of fkh-9 in wild-type animals causes axonal structural defects. Rescuing fkh-9 activity in the mechanosensory neurons of daf-2;fkh-9 mutants results in severe beading and degeneration of axons.
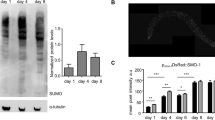
Extended Data Figure 8 WormBase gene models for fkh-9 and sod-3 are shown with modENCODE data for DAF-16 ChIP-seq experiments.
a, b, Wormbase (http://www.wormbase.org) gene models for fkh-9 (a) and sod-3 (b). Primer sets for ChIP-qPCR are depicted in a. c, Posterior intestinal FKH-9–GFP expression is only modestly increased in daf-2 compared to wild-type animals expressing fkh-9p::fkh-9::gfp. N = 25 animals.
Extended Data Figure 9 Knocking down fkh-9 via RNAi or using mutants reduces the enhanced short-term memory of daf-2 animals.
a, b, Whole-life RNAi of fkh-9 reduces daf-2 STAM. c, RNAi knockdown of fkh-9 exclusively during adulthood results in reduced daf-2 STAM comparable to daf-16 RNAi-treatment. d, e, daf-2;fkh-9 mutants have reduced learning (tested immediately following STAM training) and STAM compared to daf-2. Mean ± s.e.m., *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Time-courses showing the chemotaxis index for each time point are shown in b and e. Learning indices are shown in a, c and d.
Extended Data Figure 10 Neuronal FKH-9 is not required for the enhanced lifespan of daf-2 mutants.
a, b, Adult-only (a) or whole-life (b) fkh-9 RNAi treatment increases matricide in daf-2 worms. The cumulative percentage of animals dead as a result of bagging and/or exploding was recorded every other day. Two biological replicates were performed, with a representative experiment shown. c, Neuronal rescue of fkh-9 in daf-2;fkh-9 animals does not diminish the rate of vulval protrusions with age. N ≥ 60 per conditions for each experiment. d, Neuronal rescue of fkh-9 does not restore longevity of the daf-2;fkh-9 double mutant. daf-2 median lifespan: 41 days, daf-2;fkh-9 20 days, daf-2;fkh-9;Punc-119::fkh-9 20 days. P < 0.0001 for daf-2 vs both daf-2;fkh-9 and daf-2;fkh-9;Punc-119::fkh-9. N = 112 worms per strain. Censor rate for daf-2 19%, daf-2;fkh-9 51%, daf-2;fkh-9;Punc-119::fkh-9 56%.
Supplementary information
Supplementary Text and Tables
Supplementary Methods and legends for Supplementary Tables 1–10 (PDF 1089 kb)
Supplementary Tables 1–10
This zipped file contains Supplementary Tables 1-10 (see Supplementary Information file for legends). This zipped file was replaced on 6 January 2016 to update the file names, no content was altered. (ZIP 10606 kb)
Rights and permissions
About this article
Cite this article
Kaletsky, R., Lakhina, V., Arey, R. et al. The C. elegans adult neuronal IIS/FOXO transcriptome reveals adult phenotype regulators. Nature 529, 92–96 (2016). https://doi.org/10.1038/nature16483
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16483
This article is cited by
-
Intermediate filaments associate with aggresome-like structures in proteostressed C. elegans neurons and influence large vesicle extrusions as exophers
Nature Communications (2023)
-
Femtosecond laser microdissection for isolation of regenerating C. elegans neurons for single-cell RNA sequencing
Nature Methods (2023)
-
Regulation of fatty acid desaturase- and immunity gene-expression by mbk-1/DYRK1A in Caenorhabditis elegans
BMC Genomics (2022)
-
GAIT-GM integrative cross-omics analyses reveal cholinergic defects in a C. elegans model of Parkinson’s disease
Scientific Reports (2022)
-
Molecular encoding and synaptic decoding of context during salt chemotaxis in C. elegans
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.