Abstract
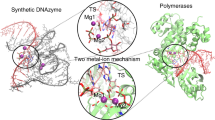
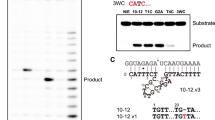
Catalysis in biology is restricted to RNA (ribozymes) and protein enzymes, but synthetic biomolecular catalysts can also be made of DNA (deoxyribozymes)1 or synthetic genetic polymers2. In vitro selection from synthetic random DNA libraries identified DNA catalysts for various chemical reactions beyond RNA backbone cleavage3. DNA-catalysed reactions include RNA and DNA ligation in various topologies4,5, hydrolytic cleavage6,7 and photorepair of DNA8, as well as reactions of peptides9,10 and small molecules11,12. In spite of comprehensive biochemical studies of DNA catalysts for two decades, fundamental mechanistic understanding of their function is lacking in the absence of three-dimensional models at atomic resolution. Early attempts to solve the crystal structure of an RNA-cleaving deoxyribozyme resulted in a catalytically irrelevant nucleic acid fold13. Here we report the crystal structure of the RNA-ligating deoxyribozyme 9DB1 (ref. 14) at 2.8 Å resolution. The structure captures the ligation reaction in the post-catalytic state, revealing a compact folding unit stabilized by numerous tertiary interactions, and an unanticipated organization of the catalytic centre. Structure-guided mutagenesis provided insights into the basis for regioselectivity of the ligation reaction and allowed remarkable manipulation of substrate recognition and reaction rate. Moreover, the structure highlights how the specific properties of deoxyribose are reflected in the backbone conformation of the DNA catalyst, in support of its intricate three-dimensional organization. The structural principles underlying the catalytic ability of DNA elucidate differences and similarities in DNA versus RNA catalysts, which is relevant for comprehending the privileged position of folded RNA in the prebiotic world and in current organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Breaker, R. R. & Joyce, G. F. A DNA enzyme that cleaves RNA. Chem. Biol. 1, 223–229 (1994)
Taylor, A. I. et al. Catalysts from synthetic genetic polymers. Nature 518, 427–430 (2015)
Schlosser, K. & Li, Y. Biologically inspired synthetic enzymes made from DNA. Chem. Biol. 16, 311–322 (2009)
Coppins, R. L. & Silverman, S. K. A DNA enzyme that mimics the first step of RNA splicing. Nature Struct. Mol. Biol. 11, 270–274 (2004)
Sreedhara, A., Li, Y. & Breaker, R. R. Ligating DNA with DNA. J. Am. Chem. Soc. 126, 3454–3460 (2004)
Chandra, M., Sachdeva, A. & Silverman, S. K. DNA-catalyzed sequence-specific hydrolysis of DNA. Nature Chem. Biol. 5, 718–720 (2009)
Gu, H., Furukawa, K., Weinberg, Z., Berenson, D. F. & Breaker, R. R. Small, highly active DNAs that hydrolyze DNA. J. Am. Chem. Soc. 135, 9121–9129 (2013)
Chinnapen, D. J. & Sen, D. A deoxyribozyme that harnesses light to repair thymine dimers in DNA. Proc. Natl Acad. Sci. USA 101, 65–69 (2004)
Walsh, S. M., Sachdeva, A. & Silverman, S. K. DNA catalysts with tyrosine kinase activity. J. Am. Chem. Soc. 135, 14928–14931 (2013)
Chandrasekar, J. & Silverman, S. K. Catalytic DNA with phosphatase activity. Proc. Natl Acad. Sci. USA 110, 5315–5320 (2013)
Li, Y. & Sen, D. A catalytic DNA for porphyrin metallation. Nature Struct. Biol. 3, 743–747 (1996)
Chandra, M. & Silverman, S. K. DNA and RNA can be equally efficient catalysts for carbon–carbon bond formation. J. Am. Chem. Soc. 130, 2936–2937 (2008)
Nowakowski, J., Shim, P. J., Prasad, G. S., Stout, C. D. & Joyce, G. F. Crystal structure of an 82-nucleotide RNA–DNA complex formed by the 10-23 DNA enzyme. Nature Struct. Biol. 6, 151–156 (1999)
Purtha, W. E., Coppins, R. L., Smalley, M. K. & Silverman, S. K. General deoxyribozyme-catalyzed synthesis of native 3′–5′ RNA linkages. J. Am. Chem. Soc. 127, 13124–13125 (2005)
Büttner, L., Seikowski, J., Wawrzyniak, K., Ochmann, A. & Höbartner, C. Synthesis of spin-labeled riboswitch RNAs using convertible nucleosides and DNA-catalyzed RNA ligation. Bioorg. Med. Chem. 21, 6171–6180 (2013)
Vicens, Q. & Cech, T. R. A natural ribozyme with 3′,5′ RNA ligase activity. Nature Chem. Biol. 5, 97–99 (2009)
Bartel, D. P. & Szostak, J. W. Isolation of new ribozymes from a large pool of random sequences [see comment]. Science 261, 1411–1418 (1993)
Ekland, E. H., Szostak, J. W. & Bartel, D. P. Structurally complex and highly active RNA ligases derived from random RNA sequences. Science 269, 364–370 (1995)
Jaeger, L., Wright, M. C. & Joyce, G. F. A complex ligase ribozyme evolved in vitro from a group I ribozyme domain. Proc. Natl Acad. Sci. USA 96, 14712–14717 (1999)
Wachowius, F., Javadi-Zarnaghi, F. & Höbartner, C. Combinatorial mutation interference analysis reveals functional nucleotides required for DNA catalysis. Angew. Chem. Int. Ed. Engl. 49, 8504–8508 (2010)
Wachowius, F. & Höbartner, C. Probing essential nucleobase functional groups in aptamers and deoxyribozymes by nucleotide analogue interference mapping of DNA. J. Am. Chem. Soc. 133, 14888–14891 (2011)
Ikawa, Y., Tsuda, K., Matsumura, S. & Inoue, T. De novo synthesis and development of an RNA enzyme. Proc. Natl Acad. Sci. USA 101, 13750–13755 (2004)
Pitt, J. N. & Ferré-D’Amaré, A. R. Structure-guided engineering of the regioselectivity of RNA ligase ribozymes. J. Am. Chem. Soc. 131, 3532–3540 (2009)
Coppins, R. L. & Silverman, S. K. Rational modification of a selection strategy leads to deoxyribozymes that create native 3′–5′ RNA linkages. J. Am. Chem. Soc. 126, 16426–16432 (2004)
Doudna, J. A. & Cech, T. R. The chemical repertoire of natural ribozymes. Nature 418, 222–228 (2002)
Fedor, M. J. & Williamson, J. R. The catalytic diversity of RNAs. Nature Rev. Mol. Cell Biol. 6, 399–412 (2005)
Shechner, D. M. & Bartel, D. P. The structural basis of RNA-catalyzed RNA polymerization. Nature Struct. Mol. Biol. 18, 1036–1042 (2011)
Davies, D. R. et al. Unique motifs and hydrophobic interactions shape the binding of modified DNA ligands to protein targets. Proc. Natl Acad. Sci. USA 109, 19971–19976 (2012)
Shechner, D. M. et al. Crystal structure of the catalytic core of an RNA-polymerase ribozyme. Science 326, 1271–1275 (2009)
Robertson, M. P. & Scott, W. G. The structural basis of ribozyme-catalyzed RNA assembly. Science 315, 1549–1553 (2007)
Pitsch, S., Weiss, P. A., Jenny, L., Stutz, A. & Wu, X. Reliable chemical synthesis of oligoribonucleotides (RNA) with 2′-O-[(Triisopropylsilyl)oxy]methyl(2′-O-TOM)-protected phosphoramidites. Helv. Chim. Acta 84, 3773–3795 (2001)
Kumar, R. K., Olsen, P. & Ravikumar, V. T. An alternative advantageous protocol for efficient synthesis of phosphorothioate oligonucleotides utilizing phenylacetyl disulfide (PADS). Nucleosides Nucleotides Nucleic Acids 26, 181–188 (2007)
Frederiksen, J. K. & Piccirilli, J. A. Separation of RNA phosphorothioate oligonucleotides by HPLC. Methods Enzymol. 468, 289–309 (2009)
Koch, M. et al. Role of a ribosomal RNA phosphate oxygen during the EF-G-triggered GTP hydrolysis. Proc. Natl Acad. Sci. USA 112, E2561–E2568 (2015)
Ludwig, J. & Eckstein, F. Rapid and efficient synthesis of nucleoside 5′-O-(1-thiotriphosphates), 5′-triphosphates and 2′,3′-cyclophosphorothioates using 2-chloro-4h-1,3,2-benzodioxaphosphorin-4-one. J. Org. Chem. 54, 631–635 (1989)
Goldeck, M., Tuschl, T., Hartmann, G. & Ludwig, J. Efficient solid-phase synthesis of pppRNA by using product-specific labeling. Angew. Chem. Int. Ed. Engl. 53, 4694–4698 (2014)
Kabsch, W. X. D. S. Acta Crystallogr. D 66, 125–132 (2010)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Terwilliger, T. C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. D 65, 582–601 (2009)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
Keating, K. S. & Pyle, A. M. RCrane: semi-automated RNA model building. Acta Crystallogr. D 68, 985–995 (2012)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012)
Sun, G., Voigt, J. H., Filippov, I. V., Marquez, V. E. & Nicklaus, M. C. PROSIT: pseudo-rotational online service and interactive tool, applied to a conformational survey of nucleosides and nucleotides. J. Chem. Inf. Comput. Sci. 44, 1752–1762 (2004)
Ren, A. et al. In-line alignment and Mg2+ coordination at the cleavage site of the env22 twister ribozyme. Nat. Commun. 5, 5534 (2014)
Acknowledgements
This work was supported by the Max Planck Society. We thank J. Ludwig for a gift of 5′-triphosphorylated RNAs, J. Seikowski for assistance with RNA and DNA synthesis, F. Wachowius and B. Samanta for initial crystallization samples, P. Afonine and members of the Pena and Höbartner laboratories for discussions, and the beamline staff at the Swiss Light Source, Villigen, Switzerland for assistance with data collection.
Author information
Authors and Affiliations
Contributions
Crystallographic work was performed by A.P.-S. under the supervision of V.P., biochemical experiments were performed by A.P.-S., K.W.-T. and C.H. U.S. obtained initial crystals. A.P.-S., V.P. and C.H. designed the experiments and all authors discussed the results and contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Semi-continuous helix in the crystal lattice.
Nucleotides important for the crystal contacts are labelled accordingly.
Extended Data Figure 3 Distribution of pseudorotation phase angels of 9DB1 deoxyribozyme in comparison with twister46 and L1 ligase ribozyme30.
Extended Data Figure 4 Active sites of RNA-ligating nucleic acid enzymes.
Left: deoxyribozyme 9DB1; middle: class I ribozyme; right: L1 ligase ribozyme. Figure was generated using Protein Data Bank (PDB) accession numbers 5CKK (9DB1), 3R1L (class I) and 2OIU (L1).
Extended Data Figure 5 Positioning of the reactive nucleotides in the active centre of ligase deoxyribozyme 9DB1 in comparison to ribozymes.
A schematic of the three enzymes is shown before and after the crystal structure determination (left and central columns, respectively). Strands that were deliberately chosen are depicted in red and blue, while the ones that resulted from random selection are shown in black. Nucleotides at the ligation junction are coloured in cyan, stacking nucleotides are depicted in light red. Nucleotides shown as white-filled circles are indicated for orientation purposes. Recognition between reactive and pairing nucleotides is shown as dashed lines (left column). Structures of product-bound nucleic acid enzymes are compared (PDB accessions 5CKK (9DB1), 3HHN (class I) and 2OIU (L1), right column).
Rights and permissions
About this article
Cite this article
Ponce-Salvatierra, A., Wawrzyniak-Turek, K., Steuerwald, U. et al. Crystal structure of a DNA catalyst. Nature 529, 231–234 (2016). https://doi.org/10.1038/nature16471
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16471
This article is cited by
-
Structure of a 10-23 deoxyribozyme exhibiting a homodimer conformation
Communications Chemistry (2023)
-
Structure and mechanism of the methyltransferase ribozyme MTR1
Nature Chemical Biology (2022)
-
Time-resolved structural analysis of an RNA-cleaving DNA catalyst
Nature (2022)
-
Caging-Decaging Strategies to Realize Spatiotemporal Control of DNAzyme Activity for Biosensing and Bioimaging
Chemical Research in Chinese Universities (2022)
-
DNA nano-pocket for ultra-selective uranyl extraction from seawater
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.