Abstract
The carboxy-terminal domain (CTD) of the RNA polymerase II (RNAP II) subunit POLR2A is a platform for modifications specifying the recruitment of factors that regulate transcription, mRNA processing, and chromatin remodelling. Here we show that a CTD arginine residue (R1810 in human) that is conserved across vertebrates is symmetrically dimethylated (me2s). This R1810me2s modification requires protein arginine methyltransferase 5 (PRMT5) and recruits the Tudor domain of the survival of motor neuron (SMN, also known as GEMIN1) protein, which is mutated in spinal muscular atrophy. SMN interacts with senataxin, which is sometimes mutated in ataxia oculomotor apraxia type 2 and amyotrophic lateral sclerosis. Because POLR2A R1810me2s and SMN, like senataxin, are required for resolving RNA–DNA hybrids created by RNA polymerase II that form R-loops in transcription termination regions, we propose that R1810me2s, SMN, and senataxin are components of an R-loop resolution pathway. Defects in this pathway can influence transcription termination and may contribute to neurodegenerative disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Buratowski, S. Progression through the RNA polymerase II CTD cycle. Mol. Cell 36, 541–546 (2009)
Jonkers, I. & Lis, J. T. Getting up to speed with transcription elongation by RNA polymerase II. Nature Rev. Mol. Cell Biol. 16, 167–177 (2015)
Porrua, O. & Libri, D. Transcription termination and the control of the transcriptome: why, where and how to stop. Nature Rev. Mol. Cell Biol. 16, 190–202 (2015)
Venkatesh, S. & Workman, J. L. Histone exchange, chromatin structure and the regulation of transcription. Nature Rev. Mol. Cell Biol. 16, 178–189 (2015)
Mayer, A. et al. CTD tyrosine phosphorylation impairs termination factor recruitment to RNA polymerase II. Science 336, 1723–1725 (2012)
Hsin, J. P., Sheth, A. & Manley, J. L. RNAP II CTD phosphorylated on threonine-4 is required for histone mRNA 3′ end processing. Science 334, 683–686 (2011)
Sims, R. J. III et al. The C-terminal domain of RNA polymerase II is modified by site-specific methylation. Science 332, 99–103 (2011)
Yang, Y. et al. Arginine methylation facilitates the recruitment of TOP3B to chromatin to prevent R loop accumulation. Mol. Cell 53, 484–497 (2014)
Yang, Y. et al. TDRD3 is an effector molecule for arginine-methylated histone marks. Mol. Cell 40, 1016–1023 (2010)
Yang, Y. et al. PRMT9 is a type II methyltransferase that methylates the splicing factor SAP145. Nat. Commun. 6, 6428 (2015)
Hadjikyriacou, A., Yang, Y., Espejo, A., Bedford, M. T. & Clarke, S. G. Unique features of human protein arginine methyltransferase 9 (PRMT9) and its substrate RNA splicing factor SF3B2. J. Biol. Chem. 290, 16723–16743 (2015)
Licciardo, P. et al. The FCP1 phosphatase interacts with RNA polymerase II and with MEP50 a component of the methylosome complex involved in the assembly of snRNP. Nucleic Acids Res. 31, 999–1005 (2003)
Antonysamy, S. et al. Crystal structure of the human PRMT5:MEP50 complex. Proc. Natl Acad. Sci. USA 109, 17960–17965 (2012)
Friesen, W. J. et al. A novel WD repeat protein component of the methylosome binds Sm proteins. J. Biol. Chem. 277, 8243–8247 (2002)
Chen, C., Nott, T. J., Jin, J. & Pawson, T. Deciphering arginine methylation: Tudor tells the tale. Nature Rev. Mol. Cell Biol. 12, 629–642 (2011)
Bedford, M. T. & Clarke, S. G. Protein arginine methylation in mammals: who, what, and why. Mol. Cell 33, 1–13 (2009)
Sikorsky, T. et al. Recognition of asymmetrically dimethylated arginine by TDRD3. Nucleic Acids Res. 40, 11748–11755 (2012)
Suraweera, A. et al. Functional role for senataxin, defective in ataxia oculomotor apraxia type 2, in transcriptional regulation. Hum. Mol. Genet. 18, 3384–3396 (2009)
Skourti-Stathaki, K., Proudfoot, N. J. & Gromak, N. Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol. Cell 42, 794–805 (2011)
West, S., Gromak, N. & Proudfoot, N. J. Human 5′→3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature 432, 522–525 (2004)
Core, L. J., Waterfall, J. J. & Lis, J. T . Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008)
Bhatia, V. et al. BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511, 362–365 (2014)
Battle, D. J. et al. The SMN complex: an assembly machine for RNPs. Cold Spring Harb. Symp. Quant. Biol. 71, 313–320 (2006)
Burghes, A. H. & Beattie, C. E. Spinal muscular atrophy: why do low levels of survival motor neuron protein make motor neurons sick? Nature Rev. Neurosci. 10, 597–609 (2009)
Zhang, Z. et al. SMN deficiency causes tissue-specific perturbations in the repertoire of snRNAs and widespread defects in splicing. Cell 133, 585–600 (2008)
Bäumer, D. et al. Alternative splicing events are a late feature of pathology in a mouse model of spinal muscular atrophy. PLoS Genet. 5, e1000773 (2009)
Zhang, Z. et al. Dysregulation of synaptogenesis genes antecedes motor neuron pathology in spinal muscular atrophy. Proc. Natl Acad. Sci. USA 110, 19348–19353 (2013)
Muñoz, M. J. et al. DNA damage regulates alternative splicing through inhibition of RNA polymerase II elongation. Cell 137, 708–720 (2009)
Ling, S. C. et al. ALS-associated mutations in TDP-43 increase its stability and promote TDP-43 complexes with FUS/TLS. Proc. Natl Acad. Sci. USA 107, 13318–13323 (2010)
Kim, S. H., Shanware, N. P., Bowler, M. J. & Tibbetts, R. S. Amyotrophic lateral sclerosis-associated proteins TDP-43 and FUS/TLS function in a common biochemical complex to co-regulate HDAC6 mRNA. J. Biol. Chem. 285, 34097–34105 (2010)
Yamazaki, T. et al. FUS–SMN protein interactions link the motor neuron diseases ALS and SMA. Cell Rep . 2, 799–806 (2012)
Lagier-Tourenne, C. et al. Divergent roles of ALS-linked proteins FUS/TLS and TDP-43 intersect in processing long pre-mRNAs. Nature Neurosci. 15, 1488–1497 (2012)
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nature Neurosci. 14, 459–468 (2011)
Ishigaki, S. et al. Position-dependent FUS–RNA interactions regulate alternative splicing events and transcriptions. Sci. Rep. 2, 529 (2012)
Rogelj, B. et al. Widespread binding of FUS along nascent RNA regulates alternative splicing in the brain. Sci. Rep. 2, 603 (2012)
Aguilera, A. & Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–124 (2012)
Hatchi, E. et al. BRCA1 recruitment to transcriptional pause sites is required for R-loop-driven DNA damage repair. Mol. Cell 57, 636–647 (2015)
Sollier, J. et al. Transcription-coupled nucleotide excision repair factors promote R-loop-induced genome instability. Mol. Cell 56, 777–785 (2014)
Dhar, S. et al. Loss of the major type I arginine methyltransferase PRMT1 causes substrate scavenging by other PRMTs. Sci. Rep. 3, 1311 (2013)
Pal, S., Vishwanath, S. N., Erdjument-Bromage, H., Tempst, P. & Sif, S. Human SWI/SNF-associated PRMT5 methylates histone H3 arginine 8 and negatively regulates expression of ST7 and NM23 tumor suppressor genes. Mol. Cell. Biol. 24, 9630–9645 (2004)
Pal, S. & Sif, S. Interplay between chromatin remodelers and protein arginine methyltransferases. J. Cell. Physiol. 213, 306–315 (2007)
Kwak, Y. T. et al. Methylation of SPT5 regulates its interaction with RNA polymerase II and transcriptional elongation properties. Mol. Cell 11, 1055–1066 (2003)
Talbot, K. et al. Missense mutation clustering in the survival motor neuron gene: a role for a conserved tyrosine and glycine rich region of the protein in RNA metabolism? Hum. Mol. Genet. 6, 497–500 (1997)
Young, P. J. et al. The exon 2b region of the spinal muscular atrophy protein, SMN, is involved in self-association and SIP1 binding. Hum. Mol. Genet. 9, 2869–2877 (2000)
Boisvert, F. M., Chenard, C. A. & Richard, S. Protein interfaces in signaling regulated by arginine methylation. Sci. STKE 2005, re2 (2005)
Boisvert, F. M., Cote, J., Boulanger, M. C. & Richard, S. A proteomic analysis of arginine-methylated protein complexes. Mol. Cell. Proteomics 2, 1319–1330 (2003)
Uhlmann, T. et al. A method for large-scale identification of protein arginine methylation. Mol. Cell. Proteomics 11, 1489–1499 (2012)
Liu, K. et al. Crystal structure of TDRD3 and methyl-arginine binding characterization of TDRD3, SMN and SPF30. PLoS ONE 7, e30375 (2012)
Mak, A. B. et al. A lentiviral functional proteomics approach identifies chromatin remodeling complexes important for the induction of pluripotency. Mol. Cell. Proteomics 9, 811–823 (2010)
El Hage, A., French, S. L., Beyer, A. L. & Tollervey, D. Loss of topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010)
Boguslawski, S. J. et al. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 89, 123–130 (1986)
Glover-Cutter, K., Kim, S., Espinosa, J. & Bentley, D. L. RNA polymerase II pauses and associates with pre-mRNA processing factors at both ends of genes. Nature Struct. Mol. Biol . 15, 71–78 (2008)
Schmidt, D. et al. ChIP-seq: using high-throughput sequencing to discover protein–DNA interactions. Methods 48, 240–248 (2009)
Braunschweig, U. et al. Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res. 24, 1774–1786 (2014)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Ryan, K., Murthy, K. G., Kaneko, S. & Manley, J. L. Requirements of the RNA polymerase II C-terminal domain for reconstituting pre-mRNA 3′ cleavage. Mol. Cell. Biol. 22, 1684–1692 (2002)
Acknowledgements
We thank J. Manley for GST–CTD constructs; D. Reinberg for RNAP II me2a antibodies; D. Eick for wild-type and RNAP II (R1810A) constructs; and S. Leppla for purified S9.6 antibodies. We also thank D. Torti and D. Leung for Illumina library preparation and sequencing, T. Hajian for the purified PRMT5–WDR77 complex, J. Li for purified 8WG16 antibodies, and D. Durocher for constructive criticism and advice during the course of this work. This project was supported by the Ontario Research Fund from the Ontario Ministry of Research and Innovation (to J.F.G. and T.P.) and by CIHR Operating Grants to J.F.G. and B.J.B. D.Y.Z. was supported by a National Science and Engineering Research Council of Canada Studentship and an Ontario Graduate Scholarship. U.B. was supported by a long-term Postdoctoral Fellowship from HFSP. B.J.B. holds the University of Toronto Banbury Chair in Medical Research.
Author information
Authors and Affiliations
Contributions
J.F.G. supervised the project. D.Y.Z. performed the experiments. J.F.G. and D.Y.Z. wrote the manuscript. T.J.P., B.J.B., Z.N., and G.G. commented on experiments and edited the manuscript. G.G. prepared the FITC peptides. F.W.S. performed ChIP-seq experiments. U.B. and Y.L. performed computational data analysis for ChIP-seq. Vectors for shRNAs were provided by J.M., and G.Z. generated stable shRNA-mediated knockdown cell lines. Z.N. generated CRISPR knockout cell lines. J.M. and K.L. provided the purified Tudor domains. M.V. provided the purified PRMT5–WDR77 complex. W.L. performed isothermal titration calorimetry assays.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 The R1810me2s and R1810me2a modifications on POLR2A depend on PRMT5 and CARM1, respectively.
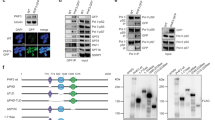
a, POLR2A carries Rme2a and Rme2s modifications. Whole-cell lysates (WCL) from HEK293 cells stably expressing Flag-tagged TDRD3 or the POLR2D subunit of RNAP II were used for immunoprecipitation using beads conjugated with M2 anti-Flag antibody, and the precipitates were western blotted with the indicated antibodies. Cells expressing Flag–GFP were used as a negative control. Precipitated TDRD3 and POLR2D contained POLR2A with the Arg-me2a modification (ASYMM24 antibody), whereas precipitated POLR2D, and not TDRD3, contained POLR2A with the Arg-me2s modification (SYMM10 and Y12 antibodies). b, Whole-cell lysate western blot controls for Figure 1b. c, Y12 and R1810me2s recognition of RNAP II CTD R1810me2s is blocked by surrounding phosphorylated residues. The detection of R1810me2s improves for both antibodies when the precipitated samples are treated with alkaline phosphatase. d, Slot blots illustrating that the Y12 and R1810me2s antibodies specifically recognize peptides containing RNAP II R1810me2s. The indicated amounts of biotin-labelled 7mer CTD peptides bracketing R1810 with no modification, Arg-me2a, and Arg-me2s were blotted before incubating with the R1810me2s or Y12 antibodies. e, Western blot confirming efficient PRMT5 knockdown, and RT–qPCR assay confirming efficient CARM1 knockdown for experiment of Fig. 1c. f, g, Whole-cell lysate western blot controls for Fig. 1c,d, respectively. h–j, Whole cell lysate western blot controls for Fig. 3c–e, respectively.
Extended Data Figure 2 Recognition of R1810me2s by SMN.
a–c, In vitro fluorescence polarization (FP) peptide binding assays. a, Recombinant SMN Tudor domain was incubated with FITC-labelled 13-mer CTD peptides bracketing either R1810 or R1603. SMN preferentially bound CTD peptides in the order R1810me2s > R1603me2s ~ R1810me2a > R1603me2a, and exhibited no detectable affinity for the unmodified peptides. b, Recombinant SMN Tudor domain was incubated with FITC-labelled CTD peptides bracketing R1810me2s also containing Y1P, S2P or both upstream of R1810me2s, showing slightly preferential binding to the peptides when the phospho-modification(s) are present. c, FITC-CTD R1810me2s or FITC-CTD R1810me2a is not recognized by other recombinant Tudor domains from SMNDC1 (also known as SPF30), TDRD1, TDRD2, TDRD9 or TDRD11 (also known as SND1). d, Isothermal titration calorimetry assays showing that the recombinant SMN Tudor domain has no enhanced binding to R1810me2s containing peptides also carrying S2P or both Y1P and S2P.
Extended Data Figure 3 Interaction of SMN and senataxin with RNAP II depends on PRMT5.
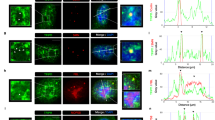
a, PRMT5 overexpression increases the R1810me2s modification and the SMN and senataxin associations with RNAP II. Left, western blots with the indicated antibodies of HEK293 whole-cell lysates overexpressing Flag-tagged PRMT5 or GFP. Overexpressing PRMT5 does not increase the amount of SMN or senataxin. Right, endogenous POLR2A was immunoprecipitated (N20 antibody) from HEK293 cell lysates with overexpressed Flag-tagged PRMT5 or GFP. b, SMN associates with phospho-isoforms of RNAP II. Immunoprecipitation with SMN antibody, but not control IgG, co-precipitated RNAP II with unmodified CTD repeats (8WG16 antibody) and CTD repeats phosphorylated on Ser5 and Ser2 as detected by western blotting. c, Requirement of PRMT5 for association of SMN and senataxin with RNAP II. Left, western blots with the indicated antibodies of HEK293 whole-cell lysates expressing siRNAs for PRMT5 or SMN. Right, endogenous POLR2A was immunoprecipitated (N20 antibody) from cells with transient siRNA-mediated knockdown of PRMT5 or SMN. Western blots were performed with the indicated antibodies. PRMT5 knockdown causes loss of R1810me2s on POLR2A, as well as reduced association of SMN and senataxin with RNAP II. SMN knockdown causes reduced association of senataxin with RNAP II. d, SMN ChIP-seq (with GFP antibody against inducible GFP–SMN, or with SMN-specific antibody in HEK293 cells). Both methods observe enriched SMN signals at promoter and termination regions; the average of both is shown.
Extended Data Figure 4 Association of SMN and senataxin with GAPDH depends on R1810 and PRMT5.
a, Chromatin immunoprecipitation (ChIP) was used to determine the distribution of SMN along the human GAPDH gene in HEK293 cells, expressed as percent input or as a ratio of SMN to RNAP II. Error bars represent technical variation in a single experiment (mean ± s.e.m., n = 3). b, SMN ChIP was performed in HEK293 cells stably expressing shRNAs for PRMT5 or GFP (as a control). With the control normalized to 1, knockdown of PRMT5 caused strong reductions of the SMN ChIP signals all along GAPDH. Error bars represent biological replicates (mean ± s.e.m., n = 3). c, SMN ChIP signals on GAPDH decrease in Raji cells expressing HA-tagged R1810A mutant POLR2A after 3 days of treatment with α-amanitin to eliminate endogenous POLR2A. ChIP results with wild-type HA-tagged POLR2A were normalized to 1. Error bars represent biological replicates (mean ± s.e.m., n = 3). d, ChIP in HEK293 cells showing the distribution of senataxin along the human GAPDH gene, expressed as percent input or as a ratio to RNAP II. Error bars represent technical variation in a single experiment (mean ± s.e.m., n = 3). e, Senataxin ChIP signals on GAPDH decrease in the POLR2A (R1810A) mutant after 3 days of treatment of Raji cells with α-amanitin to eliminate endogenous POLR2A. Results are normalized to wild-type POLR2A and expressed as the ratio of senataxin to RNAP II. Error bars represent biological replicates (mean ± s.e.m., n = 3). f, Knockdown of PRMT5 or SMN causes reductions of the senataxin ChIP signals all along GAPDH. ChIP against senataxin was performed in HEK293 cells with shRNA-mediated knock-down of PRMT5 or SMN. Results were normalized to a control knockdown of GFP and expressed as the ratio of senataxin to RNAP II. Error bars represent biological replicates (mean ± s.e.m., n = 2).
Extended Data Figure 5 The R1810 mutation causes RNAP II to accumulate in the termination region of ACTB.
Chromatin immunoprecipitation (ChIP) with three different POLR2A antibodies (8WG16, H224, 4H8), as indicated, was performed on the ACTB gene in Raji cells expressing HA-tagged wild-type or mutant (R1810A) POLR2A, 3 days after treatment with α-amanitin to eliminate endogenous POLR2A. Shown are single experiments with error bars representing technical variation (mean ± s.e.m., n = 3).
Extended Data Figure 6 R1810, PRMT5, and SMN regulate transcription termination on GAPDH.
a, Chromatin immunoprecipitation (ChIP) with the N20, 4H8, and 8WG16 antibodies to show the distribution of wild-type POLR2A along the human GAPDH gene. Error bars represent technical variation (mean ± s.e.m., n = 3). b, POLR2A ChIP along the GAPDH gene was performed in HEK293 cells after stable knockdown of PRMT5 or SMN, using stable knockdown of GFP as a negative control. RNAP II over-accumulates at the termination sites on GAPDH after knockdown of PRMT5 or SMN. Error bars represent biological replicates (mean ± s.e.m., n = 5). c, POLR2A ChIP on the GAPDH gene was performed in Raji cells that express HA-tagged wild-type or mutant (R1810A) POLR2A 3 days after α-amanitin treatment to eliminate endogenous POLR2A. R1810A mutant RNAP II over-accumulates downstream of the cleavage and polyadenylation sites where RNAP II pauses and terminates transcription on GAPDH. Error bars represent biological replicates (mean ± s.e.m., n = 4). d, Data from b, c are displayed as normalized ratios to the control (GFP knockdown, or HA wild-type POLR2A), with the ratio for the intron 5 qPCR primers at 2436 set as 1. e, Nuclear run-on experiment in which nuclei from Raji cells expressing wild-type or mutant (R1810A) POLR2A 3 days after α-amanitin treatment to eliminate endogenous POLR2A were incubated with BrUTP for 30 min, and short run-on RNAs were isolated by binding to anti-BrU antibodies. The R1810A mutation led to over-accumulation of active RNAP II in the region downstream of the poly(A) site on GAPDH. Error bars represent technical variation (mean ± s.e.m., n = 3).
Extended Data Figure 7 PRMT5 and SMN but not CARM1 regulate transcription termination by RNAP II on ACTB and GAPDH.
a, Chromatin immunoprecipitation (ChIP) for POLR2A with 4H8 antibody was performed after stable shRNA-mediated PRMT5, CARM1 or SMN knockdown in HEK293 cells to show that only PRMT5 and SMN knockdowns lead to the over-accumulation of RNAP II in the termination regions of ACTB. The graph shows a single experiment with error bars representing technical variation (mean ± s.e.m., n = 3). b, POLR2A ChIP with the 8WG16 antibody was performed after transient siRNA-mediated knockdown of PRMT5 or SMN in HEK293 cells to show that PRMT5 or SMN knockdown leads to the over-accumulation of RNAP II in the termination region of β-actin. The graph shows a single experiment with error bars representing technical variation (mean ± s.e.m., n = 3). c, ChIP for POLR2A with 4H8 antibody was performed after stable shRNA-mediated knockdown of PRMT5, SMN or GFP (as a control) in HEK293 cells to show that knockdown of PRMT5 or SMN leads to the over-accumulation of RNAP II in the termination region of GAPDH. The graph shows a single experiment with error bars representing technical variation (mean ± s.e.m., n = 3). d, POLR2A ChIP with 8WG16 antibody was performed after transient siRNA-mediated knockdown of PRMT5 or SMN in HEK293 cells to show that knockdown of PRMT5 or SMN leads to the over-accumulation of RNAP II in the termination region of GAPDH. The graph shows a single experiment with error bars representing technical variation (mean ± s.e.m., n = 3).
Extended Data Figure 8 RNAP II pausing defect and R-loop accumulation are observed in CRISPR SMN knockout cells and in SMA disease cells.
a, ChIP for POLR2A with N20 antibody was performed after stable SMN knockout (CRISPR) in HEK293 cells shows that RNAP II accumulates in the termination regions of ACTB. Scrambled guide RNA treatment was used as a negative control. Error bars represent biological variation (mean ± s.e.m., n = 3). b, Accumulation of R-loops in the termination regions of the ACTB gene after SMN knockout. A fusion protein of GFP–RNase H DNA–RNA hybrid binding domain was stably expressed in HEK293 cells. ChIP with GFP antibody (Abcam 290) was used for the detection of the R-loops (DNA–RNA hybrids), using the indicated primer positions for qPCR along the gene. Scrambled guide RNA treatment was used as a negative control. Error bars represent biological variation (mean ± s.e.m., n = 3). c, Top: live cell microscopy images showing that HEK293 cells with SMN knockout appear to be physiologically normal in comparison to the control scrambled KO. Bottom: western blot with anti-SMN antibody showing that SMN expression is knocked out by CRISPR. d, Human cell lines (3 fibroblast, 3 B lymphocyte) were obtained from the Coriell Institute for Medical Research. These include cells from two children with SMA disease and their normal parents. e, ChIP for POLR2A (N20, 8WG16 antibodies) was performed on the ACTB gene using the averaged value of the parents as control. POLR2A in the SMA disease cells (from both fibroblast and B cell lines) accumulates in the termination regions of the ACTB gene. Error bars represent biological variation (mean ± s.e.m., n = 4). f, Top: quantification of R-loops by DNA immunoprecipitation (DIP) with the S9.6 antibody in the patient cells, showing that the R-loops are sensitive to RNase H. Error bars represent technical variation (mean ± s.e.m., n = 3). Bottom: R-loop DIP with the S9.6 antibody shows that R-loops accumulate in the termination regions of the ACTB gene in the SMA disease cells. The averaged value of the parents was used as a control. Error bars represent biological variation (mean ± s.e.m., n = 5).
Extended Data Figure 9 SMN and POLR2A(R1810) effects on transcription termination by RNAP II occur on a genome-wide level.
a, Western blot using N20 antibody for POLR2A and SMN antibody to verify shSMN knockdown of SMN for the ChIP-seq experiment of Fig. 4e; shGFP was used as a control. b, Western blot using N20 and HA antibodies to verify equal expression of HA-tagged wild-type and POLR2A(R1810A) and the effect of α-amanitin treatment on cells without an HA-tagged construct for the ChIP-seq experiment of Fig. 4e. c, RNAP II ChIP-seq results for several housekeeping genes are displayed in detail (B2M, CD40, ATF4 (a short gene), JUND (an intronless gene), and TUBA1B) using the Integrative Genomics Viewer. The promoter peaks are displayed to the left, and the regions underlined in red are RNAP II termination regions. IgG ChIPseq was used as negative control. Approximately 10 million unique RNAP II ChIPseq reads (4H8, 8WG16) were obtained from GFP or SMN stable knockdown Raji cells. Approximately 10–12 million unique RNAP II ChIP-seq reads (N20) were obtained from wild-type or R1810A POLR2A Raji cells upon 3 days of amanitin treatment (2 μg ml−1).
Extended Data Figure 10 Model of the pathway that regulates R-loop accumulation to prevent DNA damage.
a, b, Quantification of γH2AX:H2AX ratio through ChIP in HEK293 (a) or Raji (b) cells, along the ACTB gene, after knocking down PRMT5, or SMN, with GFP knockdown as a negative control (a), or mutating R1810 to alanine (b). Error bars denote s.e.m. (n = 4). c, Pathway (boxed) and model showing influence of PRMT5, R1810me2s, and SMN on R-loop resolution and transcription termination.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 which shows the number of times each type of experiment described in this work was performed with the reported results) and raw western blots used in the paper. (PDF 1998 kb)
Rights and permissions
About this article
Cite this article
Yanling Zhao, D., Gish, G., Braunschweig, U. et al. SMN and symmetric arginine dimethylation of RNA polymerase II C-terminal domain control termination. Nature 529, 48–53 (2016). https://doi.org/10.1038/nature16469
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16469
This article is cited by
-
Tyrosine phosphorylation of CARM1 promotes its enzymatic activity and alters its target specificity
Nature Communications (2024)
-
Analysis of asymptomatic Drosophila models for ALS and SMA reveals convergent impact on functional protein complexes linked to neuro-muscular degeneration
BMC Genomics (2023)
-
Nucleolar reorganization after cellular stress is orchestrated by SMN shuttling between nuclear compartments
Nature Communications (2023)
-
Critical Roles of Protein Arginine Methylation in the Central Nervous System
Molecular Neurobiology (2023)
-
A small molecule antagonist of SMN disrupts the interaction between SMN and RNAP II
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.