Abstract
Plasmodium falciparum antigens expressed on the surface of infected erythrocytes are important targets of naturally acquired immunity against malaria, but their high number and variability provide the pathogen with a powerful means of escape from host antibodies1,2,3,4. Although broadly reactive antibodies against these antigens could be useful as therapeutics and in vaccine design, their identification has proven elusive. Here we report the isolation of human monoclonal antibodies that recognize erythrocytes infected by different P. falciparum isolates and opsonize these cells by binding to members of the RIFIN family. These antibodies acquired broad reactivity through a novel mechanism of insertion of a large DNA fragment between the V and DJ segments. The insert, which is both necessary and sufficient for binding to RIFINs, encodes the entire 98 amino acid collagen-binding domain of LAIR1, an immunoglobulin superfamily inhibitory receptor encoded on chromosome 19. In each of the two donors studied, the antibodies are produced by a single expanded B-cell clone and carry distinct somatic mutations in the LAIR1 domain that abolish binding to collagen and increase binding to infected erythrocytes. These findings illustrate, with a biologically relevant example, a novel mechanism of antibody diversification by interchromosomal DNA transposition and demonstrate the existence of conserved epitopes that may be suitable candidates for the development of a malaria vaccine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
NCBI Reference Sequence
Data deposits
The VH and VL sequences of the antibodies have been deposited in GenBank under accession numbers from KU058438 to KU058491 (Supplementary Table 1).
Change history
06 January 2016
The email address for corresponding author Pe.B. was corrected.
References
Chan, J.-A., Fowkes, F. J. I. & Beeson, J. G. Surface antigens of Plasmodium falciparum-infected erythrocytes as immune targets and malaria vaccine candidates. Cell. Mol. Life Sci. 71, 3633–3657 (2014)
Scherf, A., Lopez-Rubio, J. J. & Riviere, L. Antigenic variation in Plasmodium falciparum . Annu. Rev. Microbiol. 62, 445–470 (2008)
Goel, S. et al. RIFINs are adhesins implicated in severe Plasmodium falciparum malaria. Nature Med. 21, 314–317 (2015)
Bull, P. C. et al. Parasite antigens on the infected red cell surface are targets for naturally acquired immunity to malaria. Nature Med. 4, 358–360 (1998)
Traggiai, E. et al. An efficient method to make human monoclonal antibodies from memory B cells: potent neutralization of SARS coronavirus. Nature Med. 10, 871–875 (2004)
Meyaard, L. The inhibitory collagen receptor LAIR-1 (CD305). J. Leukoc. Biol. 83, 799–803 (2008)
Le Roch, K. G. et al. Discovery of gene function by expression profiling of the malaria parasite life cycle. Science 301, 1503–1508 (2003)
Florens, L. et al. A proteomic view of the Plasmodium falciparum life cycle. Nature 419, 520–526 (2002)
Messier, T. L., O’Neill, J. P., Hou, S.-M., Nicklas, J. A. & Finette, B. A. In vivo transposition mediated by V(D)J recombinase in human T lymphocytes. EMBO J. 22, 1381–1388 (2003)
Vaandrager, J. W., Schuuring, E., Philippo, K. & Kluin, P. M. V. V(D)J recombinase-mediated transposition of the BCL2 gene to the IGH locus in follicular lymphoma. Blood 96, 1947–1952 (2000)
Küppers, R., Klein, U., Hansmann, M. L. & Rajewsky, K. Cellular origin of human B-cell lymphomas. N. Engl. J. Med. 341, 1520–1529 (1999)
Teng, G. et al. RAG represents a widespread threat to the lymphocyte genome. Cell 162, 751–765 (2015)
Hu, J. et al. Chromosomal loop domains direct the recombination of antigen receptor genes. Cell 163, 947–959 (2015)
Onozawa, M. et al. Repair of DNA double-strand breaks by templated nucleotide sequence insertions derived from distant regions of the genome. Proc. Natl Acad. Sci. USA. 111, 7729–7734 (2014)
Reynaud, C.-A., Aoufouchi, S., Faili, A. & Weill, J.-C. What role for AID: mutator, or assembler of the immunoglobulin mutasome? Nature Immunol. 4, 631–638 (2003)
Robbiani, D. F. et al. Plasmodium infection promotes genomic instability and AID-dependent B cell lymphoma. Cell 162, 727–737 (2015)
Wilson, P. C. et al. Somatic hypermutation introduces insertions and deletions into immunoglobulin V genes. J. Exp. Med. 187, 59–70 (1998)
Kepler, T. B. et al. Immunoglobulin gene insertions and deletions in the affinity maturation of HIV-1 broadly reactive neutralizing antibodies. Cell Host Microbe 16, 304–313 (2014)
Brondijk, T. H. C. et al. Crystal structure and collagen-binding site of immune inhibitory receptor LAIR-1: unexpected implications for collagen binding by platelet receptor GPVI. Blood 115, 1364–1373 (2010)
Trager, W. & Jensen, J. B. Human malaria parasites in continuous culture. Science 193, 673–675 (1976)
Midega, J. T. et al. Wind direction and proximity to larval sites determines malaria risk in Kilifi District in Kenya. Nat. Commun. 3, 674 (2012)
Tiller, T. et al. Efficient generation of monoclonal antibodies from single human B cells by single cell RT-PCR and expression vector cloning. J. Immunol. Methods 329, 112–124 (2008)
Lefranc, M.-P. et al. IMGT, the international ImMunoGeneTics information system. Nucleic Acids Res. 37, D1006–D1012 (2009)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Merelli, I. et al. RSSsite: a reference database and prediction tool for the identification of cryptic recombination signal sequences in human and murine genomes. Nucleic Acids Res. 38, W262–W267 (2010)
Kepler, T. B. Reconstructing a B-cell clonal lineage. I. Statistical inference of unobserved ancestors. F1000 Res. 2, 103 (2013)
Liao, H.-X. et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496, 469–476 (2013)
Pappas, L. et al. Rapid development of broadly influenza neutralizing antibodies through redundant mutations. Nature 516, 418–422 (2014)
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nature Biotechnol. 26, 1367–1372 (2008)
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011)
Acknowledgements
We thank M. Nussenzweig for providing reagents for antibody cloning and expression. This work was supported by the European Research Council (grant no. 250348 IMMUNExplore and 670955 BROADimmune), the Swiss National Science Foundation (grant no. 160279), the Swiss Vaccine Research Institute and the Wellcome Trust (grant no. 084535, 077092, 084538, 084113/Z/07/Z, 084378/Z/07/A, 092741 and 099811). A.L. is supported by the Helmut Horten Foundation. This paper is published with the permission of the Director of Kenya Medical Research Institute (KEMRI).
Author information
Authors and Affiliations
Contributions
J.T. performed all experiments involving P. falciparum; K.P. characterized genomic DNA; L.P. produced mutant antibodies; J.T., K.P. and L.P. analysed the data and wrote the manuscript; A.A. and C.M.T. performed initial parasite work; M.F. performed bioinformatics analysis; R.G. analysed MS data; D.J. and C.S.F. performed cell sorting and antibody isolation; F.M.N., J.W. and Ph.B. provided cohort samples; B.F.-R. and So.B. produced antibodies; Si.B. performed immunoprecipitation experiments; K.M., V.T., D.C. and F.S. provided supervision; A.L. and Pe.B. provided overall supervision and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.L. is the scientific founder and shareholder of Humabs BioMed. F.S. is a shareholder of Humabs BioMed. Si.B. and D.C. are employees of Humabs BioMed, a company that commercializes human monoclonal antibodies.
Extended data figures and tables
Extended Data Figure 1 Nucleotide sequence alignments of VH regions of antibodies isolated from donor C.
Dots indicate positions where the nucleotide of a mature antibody is identical to that of the UCA.
Extended Data Figure 2 Nucleotide sequence alignments of VH regions of antibodies isolated from donor D.
Dots indicate positions where the nucleotide of a mature antibody is identical to that of the UCA.
Extended Data Figure 3 Protein sequence alignments of VH regions of antibodies isolated from donors C and D.
a, Donor C. b, Donor D. Putative complementarity-determining regions (CDRs) are highlighted in red. Dots indicate positions where the amino acid of a mature antibody is identical to that of the UCA.
Extended Data Figure 4 Nucleotide sequence alignments of VL regions of antibodies isolated from donors C and D.
a–d, Antibodies from donor C use VL7-43/JL3 (a), VK4-1/JK2 (b), or VK1-5/JK2 (c), while antibodies from donor D use VK1-8/JK5 (d). Complementarity-determining regions (CDRs) are highlighted in red. Dots indicate positions where the nucleotide of a mature antibody is identical to that of the UCA.
Extended Data Figure 5 Genealogy trees generated from VL and LAIR1 exon sequences.
a–d, The trees were drawn based on the somatic mutations in light chain variable regions (a, b) or LAIR1 exons (c, d) of the antibodies isolated from donors C and D. In the donor C VL trees, VL(1), VL(2) and VL(3) refer to VL7-43/JL3, VK1-5/JK2 and VK4-1/JK2, respectively. Shown are the nucleotide and amino acid substitutions, with the latter in parentheses.
Extended Data Figure 6 Genomic DNA analysis of LAIR1-containing antibodies of donor C and donor D.
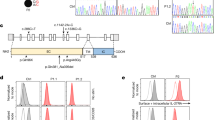
a, The sequence alignment of genomic DNA (gDNA) and cDNA of a LAIR1-containing antibody (AB) from donor C (DonC) reveals a 507 bp LAIR1 insert in chromosome 14 (Chr14) and the removal of a 160 bp fragment by RNA splicing. Splice donor and acceptor sites are highlighted in yellow. b, Schematic overview of the genomic organization of a LAIR1-containing antibody from donor D, not to scale. c, Alignment of a region of antibody-encoding DNA (chromosome 14) with the corresponding region of chromosome 13 from gDNA. The sequence maintained in the mature antibody mRNA is boxed and the splice donor site is highlighted in yellow. d, Alignment of gDNA and cDNA reveals that a part of the chromosome 13 region and the entire inserted 5′ LAIR1 intron are removed by RNA splicing. Splice donor and acceptor sites are highlighted in yellow. e, Alignment of the two repeated elements found in the inserted LAIR1 intron in chromosome 14 with the corresponding sequence in chromosome 19. The repeats are named R1 and R2, and K = G/A.
Extended Data Figure 7 LAIR1 and chromosome 13 inserts are flanked by 12/23 cryptic RSS sites.
The regions on chromosome 19 (Chr19) and chromosome 13 (Chr13) of donor-derived gDNA corresponding to the ends of the inserts were sequenced and RSS sites were identified using the RSSsite web server. The sequences shown begin from the ends of the inserts. Cryptic RSS sites are highlighted in grey, with complementary ends underlined and prediction scores shown in parentheses.
Extended Data Figure 8 Both LAIR1 alleles on chromosome 19 are intact in B cells producing LAIR1 antibodies.
Heterozygosity of the chromosome 19 LAIR1 exon in cells from donor C showing that both LAIR1 alleles are intact in B cells producing LAIR1-containing antibodies. Displayed are the chromatograms obtained for B-cell clones with or without a LAIR1 insertion (LAIR1+ or LAIR1− B cell) and for polyclonal T cells. Y = C/T
Extended Data Figure 9 Reactivity and functional assays of MGC and MGD antibodies.
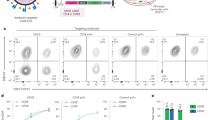
a, MGD21 staining of 3D7 IEs that were enriched or depleted of MGD21 reactivity (representative of n = 3 independent experiments). WT, wild type. b, Heat map from LC-MS analysis showing RIFIN expression levels (calculated as intensity-based absolute quantification (iBAQ) scores) in erythrocyte ghosts prepared from 3D7-MGD21+ and 3D7-MGD21− IEs (two experiments shown). Grey boxes indicate that expression levels are below the detection limit. c, Shown is the percentage of IEs (representative of n = 2 independent experiments) or of transfected CHO cells (n = 1) stained by the antibodies. RIFINs that were enriched in 3D7-MGD21+ ghosts are highlighted blue, while RIFINs that were similarly expressed or not detected in 3D7-MGD21− and 3D7-MGD21+ ghosts are shown in red. BKC3 is a negative control antibody. d, Western blot showing MGD21 binding to immunoprecipitates (IP) prepared from 9605-MGD21− and 9605-MGD21+ IEs (representative of n = 2 independent experiments). Specific bands are marked with an asterisk. Anti-human IgG was used as the secondary antibody, resulting in detection of antibodies used for immunoprecipitation alongside antigens of interest. For gel source data, see Supplementary Fig. 1. Numbers on left indicate kDa e, Percentage of 9605-MGD21− and 9605-MGD21+ IEs recognized by representative MGC and MGD antibodies (representative of n = 2 independent experiments). f, Binding of MGD21 to CHO cells transfected with RIFINs (PF3D7_1400600 and PF3D7_0100200), a RIFIN chimaera containing the constant region of PF3D7_0100200 and the variable region of PF3D7_1400600 (PF3D7_0100200c_1400600v), or the inverse chimaera (PF3D7_1400600c_0100200v) (n = 1). g, Binding of an Fc fusion protein containing the LAIR1 domain of MGD21 to CHO cells transfected with RIFINs or RIFIN chimaeras (n = 1). h, Parasitaemia of 3D7-MGD21+ in vitro culture after 2 days of incubation with various concentrations of MGD21 or an irrelevant antibody (BKC3) (n = 1). mAb, monoclonal antibody. i, Percentage of 3D7-MGD21+ IEs recognized by MGD21 after 2 days of incubation with various concentrations of MGD21 or BKC3. The antibodies were removed after 2 days (during the ring stage of the life cycle) and the parasites were allowed to grow for 24 h to the late trophozoite/schizont stage before detection with MGD21 (n = 1). j, Rosetting of 9605-MGD21+ IEs with blood group O+ or A+ uninfected erythrocytes (uEs) after incubation with MGD21 or BKC3. Shown is the mean ± s.d. from n = 4 independent experiments. Statistical significance was evaluated by the Wilcoxon signed-rank test (P > 0.1 for both blood groups). NS, not significant. k, Agglutinates of 3D7-MGD21+ or 11019-MGD21+ IEs formed by MGD21 or MGC34. Scale bar, 25 μm. l, Opsonic phagocytosis of 11019-MGD21+ IEs by monocytes (n = 2). The IEs were stained with DAPI, which was quantified in monocytes as a measure of phagocytosis.
Extended Data Figure 10 A schematic representation of interchromosomal LAIR1 transposition.
Shown is the insertion of a fragment of LAIR1 into the immunoglobulin heavy chain locus through a mechanism still to be molecularly defined, followed by the acquisition of somatic mutations that increase binding to IEs and abolish binding to collagen. Chr, chromosome.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 (Western blots for Figure 4a and Extended Data Figure 9d) and Supplementary Table 1 (accession numbers of heavy and light chain immunoglobulin sequences of donors C and D). (PDF 369 kb)
Rights and permissions
About this article
Cite this article
Tan, J., Pieper, K., Piccoli, L. et al. A LAIR1 insertion generates broadly reactive antibodies against malaria variant antigens. Nature 529, 105–109 (2016). https://doi.org/10.1038/nature16450
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16450
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.