Abstract
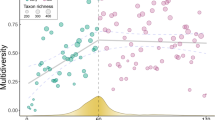
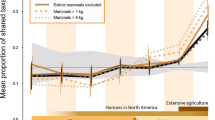
Understanding how ecological communities are organized and how they change through time is critical to predicting the effects of climate change1. Recent work documenting the co-occurrence structure of modern communities found that most significant species pairs co-occur less frequently than would be expected by chance2,3. However, little is known about how co-occurrence structure changes through time. Here we evaluate changes in plant and animal community organization over geological time by quantifying the co-occurrence structure of 359,896 unique taxon pairs in 80 assemblages spanning the past 300 million years. Co-occurrences of most taxon pairs were statistically random, but a significant fraction were spatially aggregated or segregated. Aggregated pairs dominated from the Carboniferous period (307 million years ago) to the early Holocene epoch (11,700 years before present), when there was a pronounced shift to more segregated pairs, a trend that continues in modern assemblages. The shift began during the Holocene and coincided with increasing human population size4,5 and the spread of agriculture in North America6,7. Before the shift, an average of 64% of significant pairs were aggregated; after the shift, the average dropped to 37%. The organization of modern and late Holocene plant and animal assemblages differs fundamentally from that of assemblages over the past 300 million years that predate the large-scale impacts of humans. Our results suggest that the rules governing the assembly of communities have recently been changed by human activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
06 January 2016
ED Table 1 was corrected (specifically the data owner for Wisconsin plant data).
References
Botkin, D. B. et al. Forecasting the effects of global warming on biodiversity. Bioscience 57, 227–236 (2007)
Gotelli, N. J. & Ulrich, W. The empirical Bayes approach as a tool to identify non-random species associations. Oecologia 162, 463–477 (2010)
Gotelli, N. J. & McCabe, D. J. Species co-occurrence: a meta-analysis of J.M. Diamond’s assembly rules model. Ecology 83, 2091–2096 (2002)
Marlon, J. R. et al. Global biomass burning: a synthesis and review of Holocene paleofire records and their controls. Quat. Sci. Rev. 65, 5–25 (2013)
Ruddiman, W. F. & Ellis, E. C. Effect of per-capita land use changes on Holocene forest clearance and CO2 emissions. Quat. Sci. Rev. 28, 3011–3015 (2009)
Zeder, M. A. & Smith, B. D. A conversation on agricultural origins: talking past each other in a crowded room. Curr. Anthropol. 50, 681–691 (2009)
Smith, B. D. & Yarnell, R. A. Initial formation of an indigenous crop complex in eastern North America at 3800 B.P. Proc. Natl Acad. Sci. USA 106, 6561–6566 (2009)
Connor, E. F. & McCoy, E. D. The statistics and biology of the species-area relationship. Am. Nat. 113, 791–833 (1979)
Connor, E. F., Collins, M. D. & Simberloff, D. The checkered history of checkerboard distributions. Ecology 94, 2403–2414 (2013)
Ulrich, W. & Gotelli, N. J. Null model analysis of species associations using abundance data. Ecology 91, 3384–3397 (2010)
Wagner, P. J., Kosnik, M. A. & Lidgard, S. Abundance distributions imply elevated complexity of post-Paleozoic marine ecosystems. Science 314, 1289–1292 (2006)
Wilf, P. & Labandeira, C. C. Response of plant-insect associations to Paleocene-Eocene warming. Science 284, 2153–2156 (1999)
Lyons, S. K. A quantitative model for assessing community dynamics of Pleistocene mammals. Am. Nat. 165, E168–E185 (2005)
Jackson, S. T. & Williams, J. W. Modern analogs in Quaternary paleoecology: here today, gone yesterday, gone tomorrow? Annu. Rev. Earth Planet. Sci. 32, 495–537 (2004)
Ulrich, W. Pairs – a FORTRAN program for studying pair wise species associations in ecological matrices. http://www.keib.umk.pl/wp-content/uploads/2013/04/PairsManual.pdf (2008)
Zachos, J. C., Dickens, G. R. & Zeebe, R. E. An early Cenozoic perspective on greenhouse warming and carbon-cycle dynamics. Nature 451, 279–283 (2008)
Augustin, L. et al.; EPICA community members. Eight glacial cycles from an Antarctic ice core. Nature 429, 623–628 (2004)
Bambach, R. K. Phanerozoic biodiversity mass extinctions. Annu. Rev. Earth Planet. Sci. 34, 127–155 (2006)
Bowen, G. J. et al. Two massive, rapid releases of carbon during the onset of the Palaeocene–Eocene thermal maximum. Nature Geosci. 8, 44–47 (2015)
Blois, J. L. et al. A framework for evaluating the influence of climate, dispersal limitation, and biotic interactions using fossil pollen associations across the late Quaternary. Ecography 37, 1095–1108 (2014)
Delcourt, P. A. & Delcourt, H. R. Prehistoric Native Americans and Ecological Change: Human Ecosystems in Eastern North American since the Pleistocene. (Cambridge Univ. Press, 2004)
Barnosky, A. D. Colloquium paper: megafauna biomass tradeoff as a driver of Quaternary and future extinctions. Proc. Natl Acad. Sci. USA 105 (suppl. 1), 11543–11548 (2008)
Haddad, N. M. et al. Habitat fragmentation and its lasting impact on Earth’s ecosystems. Sci. Adv. 1, e1500052 (2015)
Williams, M. Deforesting the Earth: From Prehistory To Global Crisis (Univ. Chicago Press, 2003)
Grayson, D. K. The archaeological record of human impacts on animal populations. J. World Prehist. 15, 1–68 (2001)
van Kleunen, M. et al. Global exchange and accumulation of non-native plants. Nature 525, 100–103 (2015)
Capinha, C., Essl, F., Seebens, H., Moser, D. & Pereira, H. M. The dispersal of alien species redefines biogeography in the Anthropocene. Science 348, 1248–1251 (2015)
Dornelas, M. et al. Assemblage time series reveal biodiversity change but not systematic loss. Science 344, 296–299 (2014)
Tóth, A. B., Lyons, S. K. & Behrensmeyer, A. K. A century of change in Kenya’s mammal communities: increased richness and decreased uniqueness in six protected areas. PLoS ONE 9, e93092 (2014)
Thompson, J. R., Carpenter, D. N., Cogbill, C. V. & Foster, D. R. Four centuries of change in northeastern United States forests. PLoS ONE 8, e72540 (2013)
Ulrich, W. Pairs – a FORTRAN program for studying pair-wise species associations in ecological matrices. http://www.keib.umk.pl/pairs/?lang=en, (2010)
Stone, L. & Roberts, A. The checkerboard score and species distributions. Oecologia 85, 74–79 (1990)
Gotelli, N. J. Null model analysis of species co-occurrence patterns. Ecology 81, 2606–2621 (2000)
Ulrich, W. & Gotelli, N. J. Null model analysis of species nestedness patterns. Ecology 88, 1824–1831 (2007)
Gotelli, N. J. & Ulrich, W. Statistical challenges in null model analysis. Oikos 121, 171–180 (2012)
Efron, B. Bayesians, frequentists, and scientists. J. Am. Stat. Assoc. 100, 1–5 (2005)
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009)
Kidwell, S. M. & Holland, S. M. The quality of the fossil record: implications for evolutionary analyses. Annu. Rev. Ecol. Syst. 33, 561–588 (2002)
Behrensmeyer, A. K., Kidwell, S. M. & Gastaldo, R. A. Taphonomy and paleobiology. Paleobiology 26, 103–147 (2000)
Acknowledgements
We thank G. Dietl for comments that improved the manuscript. Support for this research was provided by a National Museum of Natural History Program grant to the Evolution of Terrestrial Ecosystems Program (ETE) and NSF-DEB 1257625. This is ETE publication 338.
Author information
Authors and Affiliations
Contributions
All authors provided input into the final manuscript. S.K.L. helped design the study, contributed and analysed data, and wrote the paper. K.L.A., J.L.B., J.T.F., and W.U. were involved in the study design, and contributed and analysed data. A.T. was involved in the study design, contributed and analysed data, and created Supplementary Fig. 1. A.K.B., A.B., W.A.D., J.T.E., N.J., C.L., C.V.L., S.W., and B.R. were involved in the study design and contributed data. H.A., J.A., and D.W. contributed data. M.D. and J.H.M. were involved in the study design, analysed data, created figures and wrote text for the supplementary information. A.D. was involved in the study design and analysed data. G.R.G., B.M., D.P., S.P.-M., R.T., R.P., and A.V. were involved in the study design. N.J.G. designed the study and helped write the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
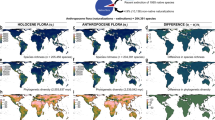
Extended Data Figure 1 Map showing distribution of fossil data sets.
Polygons enclose the localities for each fossil data set included in our analyses. Mammals are in blue, plants are in green. Dark colours represent data sets that are older. This map was created using ArcGIS software by Esri and can be found at http://www.arcgis.com/home/item.html?id=c61ad8ab017d49e1a82f580ee1298931. ArcGIS and ArcMap are the intellectual property of Esri and are used herein under license. Copyright © Esri. All rights reserved. For more information about Esri software, please visit http://www.esri.com.
Extended Data Figure 2 Breakpoint analysis of the composite data.
The analysis was performed on all data including the islands (see Fig. 3 main text), showing the mean estimate (red point; 103.778 years) and 95% confidence interval (error bar at base of plot; 101.606, 105.951 years) of the initiation of reduced percentage of aggregated species pairs, as well as the mean and confidence intervals around the change in slope of the two resulting linear models. The breakpoint analysis was run using all the data resolved to the best possible dates to allow for the greatest amount of power in detecting where the switch occurred. However, the results were similar when island data were excluded.
Extended Data Figure 3 Weighted Loess curve with and without modern data.
Loess curve weighted by number of sites with shaded 95% confidence intervals illustrates the reduction in the proportion of aggregated species pairs towards the present. Data are analysed with (black line and shading) and without (red line and shading) the modern data. Colours indicate continent: North America (green), Eurasia (purple), Australia (dark grey), South America (dark blue), Africa (orange). Point shapes indicate type of data: pollen (square), mammals (triangle), macroplants (circle). Data on terrestrial communities from ref. 2 are diamonds. Only mainland assemblages were included in the calculation for the weighted Loess curve and the density plots here and in Fig. 1.
Extended Data Figure 4 Results of PAIRS analyses of two Pleistocene–Holocene fossil data sets.
a, Mammal data for three periods: late Pleistocene (40,000–10,000 years ago), Holocene (10,000–500 years ago), and modern (present, literature survey data). Note the switch from >50% aggregated pairs to <50% aggregated pairs occurs in the Pleistocene versus Holocene data sets. b, Results for large and small mammals separately. There is a significant difference (P < 0.001) between the Holocene and the Pleistocene for all mammals (blue bars) and for large mammals (purple bars) only (P = 0.015). However, the direction of the shift was different. For all mammals, there were fewer positive associations in the Holocene, whereas, for large mammals only, there were more positive associations in the Holocene. c, North American pollen data from the past 21,000 years (modified from ref. 20). Data are from cores resolved into 1,000-year time slices. The size of the circle is related to the number of sites in the data set. The point at 0 represents a period from the present to 1,000 years ago, but is sampled from the top of the pollen cores using the same methodology as the older time slices. Note the trend of decreasing percentage of aggregations towards the present, especially in times with the largest numbers of sites (after 14,000 years). A weighted regression that takes into account the number of sites in each time slice is significant (dashed green line; P = 0.04, adjusted R2 = 0.15). The final time slice at 0 records a shift from a dominance of aggregated pairs to a dominance of segregated pairs. The sampling methods and data structure are the same for all time slices. Grey dashed line is at 50% in each panel.
Extended Data Figure 5 Relationship between the proportion of aggregated pairs and scale.
The proportion of significant pairs that are aggregated is not the result of temporal or spatial scale of data. Arcsine transformation of the proportion of significant pairs that are aggregated plotted as a function of spatial (a, b) or temporal (c, d) grain (b, d) or extent (a, c). Linear regressions are non-significant and adjusted R2 values are extremely low.
Extended Data Figure 6 Climate variability measured during the temporal extents of the fossil data sets.
Proportion of significant pairs that are aggregated shows no relationship with climate variability within a time interval. a, b, Climate variability was quantified as the standard deviation of all climate proxy data for that time interval (a), or the standard deviation of the first differences in climate across the interval (b). c, d, Climate variability (standard deviation of first differences) and age of data sets show no relationship (c), suggesting no trend in climate variability sampled by the fossil data sets across the Phanerozoic. There is a significant, but weak, positive relationship (d, dashed line) between climate variability and decreasing age of the data sets when the linear model is weighted by sample size of climate proxy data enveloped by the temporal window of the fossil data sets (P = 0.007, adjusted R2 = 0.0998).
Extended Data Figure 7 Relationship between proportion of aggregated pairs and fixed-width time intervals.
High-amplitude Pleistocene climate variability oscillating between glacial and interglacial cycles may have imposed its own novel pressures on floral and faunal communities. Furthermore, ecological impacts may lag behind climate episodes themselves, complicating efforts to quantify climatic links to changes in the proportion of aggregated species pairs over time. Thus, limiting our measure of climate variability to the temporal span of the data sets themselves may potentially not account for important (and possibly ecologically significant) climatic variability from the previous millennia. To incorporate this possibility, we re-analysed the relationship between the proportion of aggregated species pairs and climate variability of each data set, but included climate across the preceding 100,000 years, 10,000 years (not shown), and 1,000 years (not shown). As in the more restrictive analysis (Fig. 3a), there is consistently no relationship between climate variability and the proportion of aggregated species pairs.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data comprising: Dataset criteria, Modern data; Matrix Fill; Mammal Datasets; Plant Datasets. (PDF 13451 kb)
Rights and permissions
About this article
Cite this article
Kathleen Lyons, S., Amatangelo, K., Behrensmeyer, A. et al. Holocene shifts in the assembly of plant and animal communities implicate human impacts. Nature 529, 80–83 (2016). https://doi.org/10.1038/nature16447
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16447
This article is cited by
-
Disruption of trait-environment relationships in African megafauna occurred in the middle Pleistocene
Nature Communications (2023)
-
Climate-driven habitat shifts of high-ranked prey species structure Late Upper Paleolithic hunting
Scientific Reports (2023)
-
Informed selection of corridors through network and graph analyses to enhance dispersal potential through an agricultural matrix
Landscape Ecology (2023)
-
What evidence exists for temporal variability in Arctic terrestrial and freshwater biodiversity throughout the Holocene? A systematic map protocol
Environmental Evidence (2022)
-
The Dingo Barrier Fence: Presenting the case to decommission the world’s longest environmental barrier in the United Nations Decade on Ecosystem Restoration 2021–2030
Biologia Futura (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.