Abstract
Roots and leaves of healthy plants host taxonomically structured bacterial assemblies, and members of these communities contribute to plant growth and health. We established Arabidopsis leaf- and root-derived microbiota culture collections representing the majority of bacterial species that are reproducibly detectable by culture-independent community sequencing. We found an extensive taxonomic overlap between the leaf and root microbiota. Genome drafts of 400 isolates revealed a large overlap of genome-encoded functional capabilities between leaf- and root-derived bacteria with few significant differences at the level of individual functional categories. Using defined bacterial communities and a gnotobiotic Arabidopsis plant system we show that the isolates form assemblies resembling natural microbiota on their cognate host organs, but are also capable of ectopic leaf or root colonization. While this raises the possibility of reciprocal relocation between root and leaf microbiota members, genome information and recolonization experiments also provide evidence for microbiota specialization to their respective niche.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
BioProject
European Nucleotide Archive
Data deposits
Sequencing reads (454 16S rRNA, MiSeq 16S rRNA and WGS HiSeq reads) have been deposited in the European Nucleotide Archive (ENA) under accession numbers PRJEB11545, PRJEB11583 and PRJEB11584, and genome assemblies and annotations corresponding to the leaf, root and soil culture collections have been deposited in the BioProject database under accession numbers PRJNA297956, PRJNA297942 and PRJNA298127. Isolates have been deposited at the Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures (https://www.dsmz.de/).
References
Rosenberg, E. & Xilber-Rosenberg, I. The Hologenome Concept: Human, Animal and Plant Microbiota (Springer, 2013)
Spor, A., Koren, O. & Ley, R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nature Rev. Microbiol. 9, 279–290 (2011)
Berendsen, R. L., Pieterse, C. M. & Bakker, P. A. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486 (2012)
Subramanian, S. et al. Cultivating healthy growth and nutrition through the gut microbiota. Cell 161, 36–48 (2015)
Delmotte, N. et al. Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc. Natl Acad. Sci. USA 106, 16428–16433 (2009)
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012)
Lundberg, D. S. et al. Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90 (2012)
Vorholt, J. A. Microbial life in the phyllosphere. Nature Rev. Microbiol. 10, 828–840 (2012)
Bodenhausen, N., Horton, M. W. & Bergelson, J. Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PLoS One 8, e56329 (2013)
Guttman, D. S., McHardy, A. C. & Schulze-Lefert, P. Microbial genome-enabled insights into plant-microorganism interactions. Nature Rev. Genet. 15, 797–813 (2014)
Horton, M. W. et al. Genome-wide association study of Arabidopsis thaliana leaf microbial community. Nat. Commun. 5, 5320 (2014)
Schlaeppi, K., Dombrowski, N., Oter, R. G., Ver Loren van Themaat, E. & Schulze-Lefert, P. Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc. Natl Acad. Sci. USA 111, 585–592 (2014)
Edwards, J. et al. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl Acad. Sci. USA 112, E911–E920 (2015)
Hacquard, S. et al. Microbiota and host nutrition across plant and animal kingdoms. Cell Host Microbe 17, 603–616 (2015)
Bulgarelli, D. et al. Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17, 392–403 (2015)
Lebeis, S. L. et al. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349, 860–864 (2015)
Maignien, L., DeForce, E. A., Chafee, M. E., Eren, A. M. & Simmons, S. L. Ecological succession and stochastic variation in the assembly of Arabidopsis thaliana phyllosphere communities. MBio 5, e00682–e13 (2014)
Zarraonaindia, I. et al. The soil microbiome influences grapevine-associated microbiota. MBio 6, e02527–14 (2015)
Lebeis, S. L., Rott, M., Dangl, J. L. & Schulze-Lefert, P. Culturing a plant microbiome community at the cross-Rhodes. New Phytol. 196, 341–344 (2012)
Goodman, A. L. et al. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl Acad. Sci. USA 108, 6252–6257 (2011)
Faure, D., Vereecke, D. & Leveau, J. J. Molecular communication in the rhizosphere. Plant Soil 321, 279–303 (2009)
Bais, H. P., Weir, T. L., Perry, L. G., Gilroy, S. & Vivanco, J. M. The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 57, 233–266 (2006)
Ramachandran, V. K., East, A. K., Karunakaran, R., Downie, J. A. & Poole, P. S. Adaptation of Rhizobium leguminosarum to pea, alfalfa and sugar beet rhizospheres investigated by comparative transcriptomics. Genome Biol. 12, R106 (2011)
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nature Methods 10, 996–998 (2013)
Chi, F. et al. Ascending migration of endophytic rhizobia, from roots to leaves, inside rice plants and assessment of benefits to rice growth physiology. Appl. Environ. Microbiol. 71, 7271–7278 (2005)
Chelius, M. K. & Triplett, E. W. The diversity of Archaea and Bacteria in association with the roots of Zea mays L. Microb. Ecol. 41, 252–263 (2001)
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature Methods 7, 335–336 (2010)
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006)
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26, 266–267 (2010)
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014)
Tritt, A., Eisen, J. A., Facciotti, M. T. & Darling, A. E. An integrated pipeline for de novo assembly of microbial genomes. PLoS One 7, e42304 (2012)
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010)
Delcher, A. L., Harmon, D., Kasif, S., White, O. & Salzberg, S. L. Improved microbial gene identification with GLIMMER. Nucleic Acids Res. 27, 4636–4641 (1999)
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014)
Overbeek, R. et al. The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res. 33, 5691–5702 (2005)
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000)
Kanehisa, M. et al. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res. 42, D199–D205 (2014)
Eddy, S. R. Accelerated profile HMM searches. PLOS Comput. Biol. 7, e1002195 (2011)
Wu, M. & Eisen, J. A. A simple, fast, and accurate method of phylogenomic inference. Genome Biol. 9, R151 (2008)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539–539 (2011)
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010)
Dröge, J., Gregor, I. & McHardy, A. C. Taxator-tk: precise taxonomic assignment of metagenomes by fast approximation of evolutionary neighborhoods. Bioinformatics 31, 817–824 (2015)
Whitman, W. B., Coleman, D. C. & Wiebe, W. J. Prokaryotes: the unseen majority. Proc. Natl Acad. Sci. USA 95, 6578–6583 (1998)
Bodenhausen, N., Bortfeld-Miller, M., Ackermann, M. & Vorholt, J. A. A synthetic community approach reveals plant genotypes affecting the phyllosphere microbiota. PLoS Genet. 10, e1004283 (2014)
Acknowledgements
We thank D. Lundberg, S. Lebeis, S. Herrera-Paredes, S. Biswas and J. Dangl for sharing the calcined clay utilization protocol before publication; M. Kisielow of the ETH Zurich Flow Cytometry Core Facility for help with bacterial cell sorting as well as M. Baltisberger, D. Jolic and D. Weigel for their help in finding natural Arabidopsis populations; E. Kemen and M. Agler for sharing the Illumina Mi-Seq protocol for profiling of defined communities before publication and A. Sczyrba for his advice with the genome assembly. This work was supported by funds to P.S.-L. from the Max Planck Society, a European Research Council advanced grant (ROOTMICROBIOTA), the ‘Cluster of Excellence on Plant Sciences’ program funded by the Deutsche Forschungsgemeinschaft, the German Center for Infection Research (DZIF), by funds to J.A.V. from ETH Zurich (ETH Research Grant ETH-41 14-2), a grant from the Swiss National Research Foundation (310030B_152835), and a European Research Council advanced grant (PhyMo).
Author information
Authors and Affiliations
Contributions
J.A.V. and P.S.-L. initiated, coordinated and supervised the project. Y.B., M.R., N.D. and S.S. isolated root and soil bacteria strains. Y.B. collected root material and performed culture-independent community profiling. D.B.M., E.P. and M.R.-E. collected environmental leaf material, D.B.M. and E.P. isolated leaf strains and performed culture-independent community profiling. G.S. and R.G.-O. analysed culture-independent 16S rRNA amplicon sequencing data. Y.B., D.B.M. isolated DNA and prepared samples for genome sequencing. R.G.-O., P.C.M, B.H. and A.C.M. organized the genome sequencing data. R.G.-O. assembled and annotated draft genomes and performed comparative genome analyses. Y.B. and D.B.M. performed recolonization experiments; G.S. and R.G.-O. analysed the recolonization data. Y.B., D.B.M., R.G.-O., J.A.V. and P.S.-L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
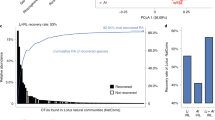
Extended Data Figure 1 Culture-dependent coverage of A. thaliana root- and leaf-associated OTUs identified in several cultivation-independent studies.
a–d, The inner circle depicts taxonomic assignments of top 100 root-associated OTUs (filled dots) for the indicated phyla and families that were identified in the current (a), ref. 6 (b) and ref. 12 (c) studies with Cologne-soil-grown plants, and current leaf (d) study at locations around Tübingen and Zurich. Black squares of the outer ring highlight OTUs sharing ≥ 97% 16S rRNA gene similarity to Arabidopsis root or leaf bacterial culture collection.
Extended Data Figure 2 16S rRNA gene community profiling of phyllosphere samples from different locations.
a–d, The indicated Beta-diversity indices were calculated from leaf samples (n = 60) collected from natural A. thaliana populations growing in the areas around Tübingen and Zurich. The indicated colour code refers to sampling locations, sampling sites, sampling season, and combined or individual leaves of respective plants.
Extended Data Figure 3 At-RSPHERE, At-LSPHERE and soil bacterial culture collections.
a, At-RSPHERE (n = 206 isolates), a culture collection of the A. thaliana root microbiota. b, At-LSPHERE (n = 224 isolates), a culture collection of the A. thaliana leaf microbiota. c, Bacteria isolated from Cologne soil (n = 33 isolates). Numbers inside white circles indicate the number of bacterial isolates sharing ≥ 97% sequence identity, but isolated from independent roots, leaves and soil batches.
Extended Data Figure 4 Taxonomy overlap between A. thaliana root- and leaf-associated bacterial community from plants grown in natural soils.
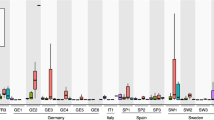
a, b, Rank abundance plots of top 20 genera (a) and OTUs (b) in root bacterial communities (n = 8) from Cologne with corresponding genera detected in leaf bacterial communities (n = 60) from Zurich and Tübingen. c, d, Rank abundance plots of top 20 genera (c) and OTUs (d) in leaf bacterial communities from Zurich and Tübingen with corresponding genera detected in root bacterial communities from Cologne. Boxplot whiskers extend to the most extreme data point which is no more than 1.5 times the interquartile range from the upper or lower quartiles.
Extended Data Figure 5 Phylogenetic distribution of ‘carbohydrate metabolism’ genes across sequenced isolates.
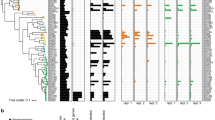
a, Phylogeny of sequenced leaf (n = 206), root (n = 194) and soil (n = 32) isolates based on the concatenated alignment of the 31 conserved AMPHORA phylogenetic marker genes. The origin of each genome (leaf, root or soil) is shown by different shapes and their taxonomic affiliation (phylum level) is depicted using various colours. Shaded areas correspond to the different clusters of genomes and are annotated with their consensus taxonomy (family level). b, Relative abundance of protein coding genes classified as belonging to the KEGG general term ‘carbohydrate metabolism’, measured as percentage of annotated proteins per genome.
Extended Data Figure 6 Phylogenetic distribution of ‘xenobiotic biodegradation and metabolism’ genes across sequenced isolates.
a, Phylogeny of sequenced leaf (n = 206), root (n = 194) and soil (n = 32) isolates based on the concatenated alignment of the 31 conserved AMPHORA phylogenetic marker genes. The origin of each genome (leaf, root or soil) is shown by different shapes and their taxonomic affiliation (phylum level; class level for Proteobacteria) is depicted using various colours. Shaded areas correspond to the different clusters of genomes and are annotated with their consensus taxonomy (family level). b, Relative abundance of protein coding genes classified as belonging to the KEGG general term ‘xenobiotics biodegradation and metabolism’, measured as percentage of annotated proteins per genome.
Extended Data Figure 7 V. vinifera metagenome comparison.
a, b, Functional enrichment analysis of V. vinifera root and soil shotgun metagenomes (a; n = 47) compared to A. thaliana culture collection root and soil genomes (b; n = 432). Functional category abundances correspond to the percentage of annotated genes in each genome or metagenome sample. Boxplot whiskers extend to the most extreme data point which is no more than 1.5 times the interquartile range from the upper or lower quartiles.
Extended Data Figure 8 Cluster analysis of Bray–Curtis distances between groups of samples in the SynCom colonization of germ-free A. thaliana experiments.
a, Comparison of pairwise distances within input samples and between input and output samples of the RS in clay experiments. b, Comparison of pairwise distances between samples within the same cluster and between different clusters of the RS in clay experiments. c, Comparison of pairwise distances between input samples and between input and output samples of the L spray experiments. d, Comparison of pairwise distances within samples within the same cluster and between different clusters of the L spray experiments. e, Comparison of pairwise distances between samples within the same cluster and between different clusters of the leaf output across experiments. f, Comparison of pairwise distances between leaf output samples in the RSL in clay experiments and leaf output samples in the L in clay and RS in clay experiments. g, Comparison of pairwise distances between root output samples in the RSL in clay experiments and root output samples in the L in clay and RS in clay experiments. All comparisons marked with asterisks were subjected to a Student’s t-test (P < 0.001 in each case). L in clay was tested with 6 independently prepared SynComs (n = 6); RSL in clay experiment was tested with 3 independently prepared SynComs, each used for 3 independent inoculations (n = 9). All other experiments were tested with 6 independently prepared SynComs and each preparation was used for 3 independent inoculations (n = 18). L, leaf-derived strains; RS, root- and soil-derived strains.
Extended Data Figure 9 Similarity of rank abundances of SynCom outputs with corresponding root- and leaf-associated OTUs of plants grown in natural environments.
a–c, Rank abundance plots of SynCom root outputs (n = 69) with corresponding root-associated OTUs in natural communities (n = 8) from plants grown in the present study in Cologne soil at the taxonomic ranks of phylum (a), order (b) and family (c). d–f, Rank abundance plots of SynCom leaf outputs (n = 69) with corresponding leaf-associated OTUs in natural communities (n = 60) from plants grown in the present study around Tuebingen or Zurich at the taxonomic ranks of phylum (d), order (e) and family (f). Boxplot whiskers extend to the most extreme data point which is no more than 1.5 times the interquartile range from the upper or lower quartiles.
Extended Data Figure 10 Fractional contribution of At-LSPHERE and At-RPSHERE-specific OTUs and SynCom competition supports host organ-specific community assemblies.
a, Fractional contribution of At-LSPHERE and At-RPSHERE specific OTUs in the input, leaf and the root output communities in the ‘RSL in clay’ experiment (n = 9). b, c, PCoA of Bray–Curtis distances of root (b; n = 21) and leaf (c; n = 21) outputs of the ‘R in clay’, ‘RS in clay’, and ‘R spray’ SynCom experiments. R, root-derived isolates; S, soil-derived isolates; L, leaf-derived isolates. RSL in clay experiment was tested with 3 independently prepared SynComs, each used for 3 independent inoculations. All other experiments were tested with 3 independently prepared SynComs and each preparation was used for 3 independent inoculations. Boxplot whiskers extend to the most extreme data point which is no more than 1.5 times the interquartile range from the upper or lower quartiles.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-9. (PDF 9119 kb)
Supplementary Data
This zipped folder contains Supplementary Data files 1-7 and a Supplementary Data guide. (ZIP 17144 kb)
Rights and permissions
About this article
Cite this article
Bai, Y., Müller, D., Srinivas, G. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015). https://doi.org/10.1038/nature16192
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16192
This article is cited by
-
Uncovering the mechanisms underlying pear leaf apoplast protein-mediated resistance against Colletotrichum fructicola through transcriptome and proteome profiling
Phytopathology Research (2024)
-
Selective regulation of endophytic bacteria and gene expression in soybean by water-soluble humic materials
Environmental Microbiome (2024)
-
Dynamic root microbiome sustains soybean productivity under unbalanced fertilization
Nature Communications (2024)
-
Commensal lifestyle regulated by a negative feedback loop between Arabidopsis ROS and the bacterial T2SS
Nature Communications (2024)
-
Chromosomal barcodes for simultaneous tracking of near-isogenic bacterial strains in plant microbiota
Nature Microbiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.