Abstract
Somatic stem cells maintain tissue homeostasis by dynamically adjusting proliferation and differentiation in response to stress and metabolic cues. Here we identify Ca2+ signalling as a central regulator of intestinal stem cell (ISC) activity in Drosophila. We show that dietary L-glutamate stimulates ISC division and gut growth. The metabotropic glutamate receptor (mGluR) is required in ISCs for this response, and for an associated modulation of cytosolic Ca2+ oscillations that results in sustained high cytosolic Ca2+ concentrations. High cytosolic Ca2+ concentrations induce ISC proliferation by regulating Calcineurin and CREB-regulated transcriptional co-activator (Crtc). In response to a wide range of dietary and stress stimuli, ISCs reversibly transition between Ca2+ oscillation states that represent poised or activated modes of proliferation, respectively. We propose that the dynamic regulation of intracellular Ca2+ levels allows effective integration of diverse mitogenic signals in ISCs to adapt their proliferative activity to the needs of the tissue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
O’Brien, L. E., Soliman, S. S., Li, X. & Bilder, D. Altered modes of stem cell division drive adaptive intestinal growth. Cell 147, 603–614 (2011)
Buchon, N., Broderick, N. A. & Lemaitre, B. Gut homeostasis in a microbial world: insights from Drosophila melanogaster. Nature Rev. Microbiol. 11, 615–626 (2013)
Biteau, B., Hochmuth, C. E. & Jasper, H. Maintaining tissue homeostasis: dynamic control of somatic stem cell activity. Cell Stem Cell 9, 402–411 (2011)
Lemaitre, B. & Miguel-Aliaga, I. The digestive tract of Drosophila melanogaster. Annu. Rev. Genet. 47, 377–404 (2013)
Ayyaz, A. & Jasper, H. Intestinal inflammation and stem cell homeostasis in aging Drosophila melanogaster. Front. Cell. Infect. Microbiol. 3, 98 (2013)
Skorupa, D. A., Dervisefendic, A., Zwiener, J. & Pletcher, S. D. Dietary composition specifies consumption, obesity, and lifespan in Drosophila melanogaster. Aging Cell 7, 478–490 (2008)
Lee, K. P. et al. Lifespan and reproduction in Drosophila: new insights from nutritional geometry. Proc. Natl Acad. Sci. USA 105, 2498–2503 (2008)
Reeds, P. J., Burrin, D. G., Stoll, B. & Jahoor, F. Intestinal glutamate metabolism. J. Nutr. 130, 978S–982S (2000)
Newsholme, P., Procopio, J., Lima, M. M., Pithon-Curi, T. C. & Curi, R. Glutamine and glutamate–their central role in cell metabolism and function. Cell Biochem. Funct. 21, 1–9 (2003)
Xiao, W. et al. Glutamate prevents intestinal atrophy via luminal nutrient sensing in a mouse model of total parenteral nutrition. FASEB J. 28, 2073–2087 (2014)
Jiang, H. et al. Cytokine/Jak/Stat signaling mediates regeneration and homeostasis in the Drosophila midgut. Cell 137, 1343–1355 (2009)
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci. 24, 251–254 (2001)
Lewerenz, J. et al. The cystine/glutamate antiporter system xc− in health and disease: from molecular mechanisms to novel therapeutic opportunities. Antioxid. Redox Signal. 18, 522–555 (2013)
Freedman, N. J. & Lefkowitz, R. J. Desensitization of G protein-coupled receptors. Recent Prog. Horm. Res. 51, 319–351; discussion 352–313 (1996)
Danbolt, N. C. Glutamate uptake. Prog. Neurobiol. 65, 1–105 (2001)
Burrin, D. G. & Stoll, B. Metabolic fate and function of dietary glutamate in the gut. Am. J. Clin. Nutr. 90, 850S–856S (2009)
Rival, T. et al. Decreasing glutamate buffering capacity triggers oxidative stress and neuropil degeneration in the Drosophila brain. Curr. Biol. 14, 599–605 (2004)
Li, H., Qi, Y. & Jasper, H. Dpp signaling determines regional stem cell identity in the regenerating adult Drosophila gastrointestinal tract. Cell Rep 4, 10–18 (2013)
McBride, S. M. et al. Pharmacological rescue of synaptic plasticity, courtship behavior, and mushroom body defects in a Drosophila model of fragile X syndrome. Neuron 45, 753–764 (2005)
Brini, M. & Carafoli, E. Calcium pumps in health and disease. Physiol. Rev. 89, 1341–1378 (2009)
Dupont, G., Combettes, L., Bird, G. S. & Putney, J. W. Calcium oscillations. Cold Spring Harb. Perspect. Biol. 3, a004226 (2011)
Clapham, D. E. Calcium signaling. Cell 131, 1047–1058 (2007)
Shim, J. et al. Olfactory control of blood progenitor maintenance. Cell 155, 1141–1153 (2013)
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nature Methods 6, 875–881 (2009)
Akerboom, J. et al. Optimization of a GCaMP calcium indicator for neural activity imaging. J. Neurosci. 32, 13819–13840 (2012)
Wong, C. O. et al. A TRPV channel in Drosophila motor neurons regulates presynaptic resting Ca2+ levels, synapse growth, and synaptic transmission. Neuron 84, 764–777 (2014)
Daniels, R. W., Rossano, A. J., Macleod, G. T. & Ganetzky, B. Expression of multiple transgenes from a single construct using viral 2A peptides in Drosophila. PLoS ONE 9, e100637 (2014)
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013)
Roti, G. et al. Complementary genomic screens identify SERCA as a therapeutic target in NOTCH1 mutated cancer. Cancer Cell 23, 390–405 (2013)
Periz, G. & Fortini, M. E. Ca2+-ATPase function is required for intracellular trafficking of the Notch receptor in Drosophila. EMBO J. 18, 5983–5993 (1999)
Nakai, Y. et al. Calcineurin and its regulator Sra/DSCR1 are essential for sleep in Drosophila. J. Neurosci. 31, 12759–12766 (2011)
Takeo, S., Tsuda, M., Akahori, S., Matsuo, T. & Aigaki, T. The calcineurin regulator sra plays an essential role in female meiosis in Drosophila. Curr. Biol. 16, 1435–1440 (2006)
Altarejos, J. Y. & Montminy, M. CREB and the CRTC co-activators: sensors for hormonal and metabolic signals. Nature Rev. Mol. Cell Biol. 12, 141–151 (2011)
Wang, B. et al. The insulin-regulated CREB coactivator TORC promotes stress resistance in Drosophila. Cell Metab. 7, 434–444 (2008)
Hirano, Y. et al. Fasting launches CRTC to facilitate long-term memory formation in Drosophila. Science 339, 443–446 (2013)
Buchon, N., Broderick, N. A., Kuraishi, T. & Lemaitre, B. Drosophila EGFR pathway coordinates stem cell proliferation and gut remodeling following infection. BMC Biol. 8, 152 (2010)
Hochmuth, C. E., Biteau, B., Bohmann, D. & Jasper, H. Redox regulation by Keap1 and Nrf2 controls intestinal stem cell proliferation in Drosophila. Cell Stem Cell 8, 188–199 (2011)
Wu, Q., Zhang, Y., Xu, J. & Shen, P. Regulation of hunger-driven behaviors by neural ribosomal S6 kinase in Drosophila. Proc. Natl Acad. Sci. USA 102, 13289–13294 (2005)
Biteau, B. & Jasper, H. EGF signaling regulates the proliferation of intestinal stem cells in Drosophila. Development 138, 1045–1055 (2011)
Wang, L., Zeng, X., Ryoo, H. D. & Jasper, H. Integration of UPRER and oxidative stress signaling in the control of intestinal stem cell proliferation. PLoS Genet. 10, e1004568 (2014)
Ayyaz, A., Li, H. & Jasper, H. Haemocytes control stem cell activity in the Drosophila intestine. Nature Cell Biol. 17, 736–748 (2015)
Julio-Pieper, M., Flor, P. J., Dinan, T. G. & Cryan, J. F. Exciting times beyond the brain: metabotropic glutamate receptors in peripheral and non-neural tissues. Pharmacol. Rev. 63, 35–58 (2011)
DeBerardinis, R. J. & Cheng, T. Q’s next: the diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene 29, 313–324 (2010)
Guo, L., Karpac, J., Tran, S. L. & Jasper, H. PGRP-SC2 promotes gut immune homeostasis to limit commensal dysbiosis and extend lifespan. Cell 156, 109–122 (2014)
Yan, D. Protection of the glutamate pool concentration in enteric bacteria. Proc. Natl Acad. Sci. USA 104, 9475–9480 (2007)
Biteau, B., Hochmuth, C. E. & Jasper, H. JNK activity in somatic stem cells causes loss of tissue homeostasis in the aging Drosophila gut. Cell Stem Cell 3, 442–455 (2008)
Lin, G., Xu, N. & Xi, R. Paracrine Wingless signalling controls self-renewal of Drosophila intestinal stem cells. Nature 455, 1119–1123 (2008)
Takashima, S. et al. Development of the Drosophila entero-endocrine lineage and its specification by the Notch signaling pathway. Dev. Biol. 353, 161–172 (2011)
Ja, W. W. et al. Prandiology of Drosophila and the CAFE assay. Proc. Natl Acad. Sci. USA 104, 8253–8256 (2007)
Grandison, R. C., Piper, M. D. & Partridge, L. Amino-acid imbalance explains extension of lifespan by dietary restriction in Drosophila. Nature 462, 1061–1064 (2009)
Lee, W. C. & Micchelli, C. A. Development and characterization of a chemically defined food for Drosophila. PLoS ONE 8, e67308 (2013)
Olds, W. H. & Xu, T. Regulation of food intake by mechanosensory ion channels in enteric neurons. Elife 3, 04402 (2014)
Cognigni, P., Bailey, A. P. & Miguel-Aliaga, I. Enteric neurons and systemic signals couple nutritional and reproductive status with intestinal homeostasis. Cell Metab. 13, 92–104 (2011)
Sanyal, S., Jennings, T., Dowse, H. & Ramaswami, M. Conditional mutations in SERCA, the Sarco-endoplasmic reticulum Ca2+-ATPase, alter heart rate and rhythmicity in Drosophila. J. Comp. Physiol. B 176, 253–263 (2006)
Ha, E. M. et al. Regulation of DUOX by the Gαq-phospholipase Cβ-Ca2+ pathway in Drosophila gut immunity. Dev. Cell 16, 386–397 (2009)
Acknowledgements
This work was supported by the National Institute on Aging (R01 AG028127), the National Institute on General Medical Sciences (R01 GM100196), and by a Glenn Foundation for Medical Research postdoctoral fellowship to H.D. The Zeiss 7MP was purchased using National Institutes of Health (NIH) grant S10OD010414. We would like to thank G. Hasan, Y. Hirano, A. Toshiro, M. Parmentier, W. Lee, S. X. Hou, M. Montminy, N. Perrimon, B. Ohlstein A. DiAntonio, H. Hayashi, R. W. Daniels, K. Venkatachalam, the Vienna Drosophila RNAi Center, and the Bloomington Stock Center for flies, and Developmental Studies Hybridoma Bank for antibodies. We also thank Jasper laboratory members for discussion.
Author information
Authors and Affiliations
Contributions
H.D. and H.J. designed and conceived the study. H.D. and A.A.G. performed Ca2+ recording and analysis, H.D. performed all other experiments in the study. H.J. and H.D. analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.A.G. declares financial interest in Image Analyst Software.
Extended data figures and tables
Extended Data Figure 1 Integration of stress and dietary signals by Ca2+ signalling.
a, We propose that Ca2+ signalling integrates and transduces nutritional and stress signals from the environment and from systemic and local paracrine sources to respond appropriately to tissue needs. In homeostatic conditions, ISCs are largely quiescent. Glutamate derived from protein in the food is absorbed by enterocytes through Eaat1, which is expressed throughout the gut, but at relatively higher levels in the anterior midgut. We propose that excessive dietary L-Glu is not efficiently resorbed by enterocytes and activates mGluR in ISCs. The L-Glu signal is integrated with local and systemic stress and growth factor signals to promote ISC proliferation in a CaN/CRTC dependent manner. CaN/CRTC signalling is required and sufficient downstream of InR and Jak/Stat signalling to induce ISC proliferation. We propose that Crtc acts together with other transcription factors regulated by InR and Jak/Stat signalling. b, Ca2+ signalling components in Drosophila (Extended Data Fig.1b).
Extended Data Figure 2 Dietary L-Glu stimulates mGluR-dependent gut growth.
a, Left, mitotic figures in intestines refed with a range of L-Glu (w/vol%) in food with varying yeast concentrations for 4 h. Flies were starved after eclosing for 48 h, and then refed with food containing varying concentrations of yeast as only amino acid source, but supplemented with varying concentrations of L-Glu (between 0.1 and 10% L-Glu in food containing between 0.1% and 5% yeast; note that the L-Glu concentration in standard fly food is about 0.1–0.3%; refs 50, 51). Middle, 10% L-Glu refeeding inhibits ISC proliferation. Mitotic figures were quantified 24 h after flies were refed with yeast-enriched (5%) or yeast-restricted (0.1%) food supplemented with 10% glutamate. Blue food dye was included in the food to monitor food intake. Right, distribution of mitotic ISCs along the gastrointestinal tract after L-Glu refeeding. Number of proliferating ISCs (pH3+) in anterior and posterior midguts (AM and PM, respectively) is quantified. b, Food intake is not affected by changing the L-Glu concentration in the food. CAFÉ assay49 and ingestion of coloured food are shown after 4 h of refeeding. c, Injection of L-Glu fails to promote ISC proliferation. Food-naive flies were starved for 2 days before injection with the indicated solutions. Injected or non-injected flies were examined after recovery for the indicated time points. A typical example of injected fly was shown on the top. Blue food dye was mixed into the injected solutions to monitor its distribution throughout the body. d, Representative images of guts after refeeding. The red lines indicate the length and width measurements used to quantify relative gut size. e, Left, ISC proliferation in animals refed sucrose solution supplemented with indicated amino acids (1% w/vol final). The number of pH3+ ISCs was determined 6 h after refeeding. Right, ISC proliferation in animals refed isocaloric sucrose-only solution (5.2% sucrose, the calorie content of 0.2% sucrose is equivalent to 1% glutamate) or refed excess sucrose (10%). f, Feeding L-Glu for 7 days promotes growth of esgts FLP-out (F/O) clones. Note large clones in L-Glu-fed intestines (box). Genotype of esgts F/O: esg::GAL4, UAS-FLP, tub::GAL80ts; UAS-nls-GFP; tub->FRT-CD2-FRT>GAL4. g, Increased cell density, lineage growth and intestinal diameter in L-Glu-treated guts. Cell density in the posterior midgut was determined in esgts F/O flies exposed to mock or L-Glu-supplemented food for 7 days (esgts F/O allows lineage tracing from ISCs). Genotype: esg::GAL4, UAS-FLP, UAS-GFP, Act > FRT-stop-FRT > GAL4; tub::GAL80ts. Cross sections through the posterior midgut and overview images are shown highlighting the increase in intestinal diameter and in clonal growth after L-Glu supplementation. Clones were analysed 7 days after clone induction at 29 °C. h, L-Glu supplementation promotes growth of ISC lineages. ISC lineages were marked by MARCM (genotype: hs-FLP, UAS-GFP;tub::GAL4,FRT82B,tub::GAL80/FRT82B). Cell numbers in GFP+ ISC clones were counted 7 days after heat-shock induction and refeeding. i, Related to Fig. 1b. Mitotic figures and gut width measured after L-Glu refeeding. mGluR112b is a null allele of mGluR. For pH3 number, n = 12 per condition, for gut width measurement, n = 8 per condition. Data and s.e.m. are shown. P values from Student’s t-test (a (middle), b, c, g, h and i) and one-way ANOVA (a (left and right) and e). The sample size (per condition) is as follows: n = 8 for a (left), n = 7 for a (middle), n = 11 for a (right), n = 12 for b and c, n = 9 for d, n = 11 for e (left), n = 6 for e (right), n = 6 for g, n = 14 for i (left) and n = 9 for i (right). For h, clones (n = 32 for control and n = 42 for glutamate food) from 5 guts were analysed. Data are representative of 3 (a, e and g) and 2 (b, c, h and i) independently performed experiments.
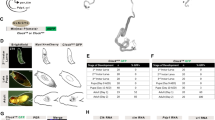
Extended Data Figure 3 ISC regulation by Eaat1.
a, Eaat1 transcript is enriched in the anterior midgut. In situ hybridization with antisense RNA probes against Eaat1 (detected using NBT/BCIP; sense probes shown in bottom panels). Guts overexpressing Eaat1 in enterocytes (Eaat1OE), or depleted of Eaat1 in enterocytes (Eaat1RNAi) used as positive/negative controls. b, Expression pattern of Eaat1::GAL4 in midgut. Expression of Eaat1:: GAL4> UAS-GFP (green) in anterior and posterior midgutis shown in higher magnification in the bottom panels. Cell membranes highlighted by anti-armadillo staining (red). c, Knockdown efficiency of Eaat1RNAi determined by qRT–PCR. Eaat1RNAi (BL43287) was used to knockdown Eaat1 in the gut using NP1::GAL4, tub::GAL80ts (29 °C for 10 days). NP1::GAL4ts (NP1::GAL4; tub::GAL80ts) drives expression in enterocytes throughout the gut when flies are shifted to 29 °C. Ten guts were pooled for RNA extraction and three independent groups were repeated for evaluation. d, Knocking down Eaat1 in enterocytes promotes ISC proliferation. Distribution of mitotic ISCs and percentage of Dl+ cells in intestines of flies in which Eaat1 was knocked down in enterocytes shown in the middle panels. Representative image from confocal microscopy is shown on the right. Arrowheads point to select ISCs identified by anti-Dl staining (white). e, Overexpressing Eaat1 in enterocyte suppresses L-Glu-mediated ISC proliferation. Eaat1 was overexpressed in enterocytes using NP1::GAL4ts (NP1::GAL4; tub::GAL80ts) at 29 °C. Flies were shifted to 29 °C before hatching, then maintained on normal food (0.1–0.3% L-Glu) for 4 days. Flies were then starved (with water) for another 2 days. Mitotic index was determined in intestines of flies refed with 1% glutamate for 6 h as shown in a. f, upd3 transcript in whole guts quantified by qRT–PCR (transcript levels normalized to Act5C). g, Knocking down Eaat1 in enterocytes by NP1::GAL4ts does not increase Jak/Stat signalling activity in the posterior midgut. 2×STAT-GFP is an activity reporter for Jak/Stat signalling (green). Apoptosis in enterocytes is indicated by TUNEL staining (bottom, apoptotic nuclei in red). As a positive control, bleomycin (25 μg ml−1) treatment strongly induces 2×STAT-GFP expression in visceral muscle and epithelium, results in widespread TUNEL-positive nuclei in the intestinal epithelium, and substantially increases upd3 transcripts in the posterior midgut. Data are mean and s.e.m. P values are from ANOVA (d (middle) and f) and Student’s t-test (c, d (left and right) and e). The sample size is n = 3 independent samples for each condition in c, n = 12 guts for each condition (mitotic figures), and n = 14 guts for control and n = 24 guts for Eaat1RNAi (percentage of Dl+ cells) in d; n = 15 for control and n = 21 guts for Eaat1OE in e; n = 3 independent samples for each condition in f. For d and e, a representative experiment is shown (three biological replicates). For a, b and g, representative images from a representative of two independent experiments are shown (a: n = 8 for each condition; b: n = 12; g (left): n = 12 per condition, and g (right): n = 9 per condition).
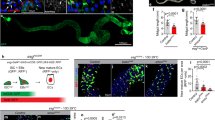
Extended Data Figure 4 Neuronal projections and GCaMP3 fluorescence pattern in the intestinal epithelium.
a, Schematic showing neuronal projections in adult Drosophila gastrointestinal tract (modified from ref. 52). Only three segments are innervated by neurons from the brain: crop, proventriculus (PV) and foregut, hindgut (HG) pylorus, and the rectum (R). b, No glutamatergic neurites are observed in the posterior midgut52,53. All neurites are stained by anti-horseradish-peroxidase (HRP) staining in white. mCD8-GFP driven by GAL4vGlut::GAL4CNS (a glutamatergic neuron driver readily expressed in adults) labels all glutamatergic neurons. Glutamatergic neurons are also detected by VGlut (vesicular glutamate transporter) immunostaining. Higher magnification is shown on the right. c, Summary of neurites innervating the Drosophila gastrointestinal tract (see also ref. 53). While L-Glu in the diet may stimulate glutamatergic neurons innervating the intestine, and a role for these neurons in stimulating ISC activity under certain circumstances cannot be ruled out, the widespread induction of ISC proliferation throughout the gut, as well as the ISC-specific requirement for mGluR suggests that the locally restricted glutamatergic innervation is not directly involved in stimulating ISC proliferation. d, GAL4vGlut::GAL4CNS>mCD8-GFP and anti-vGlut stain the presynaptic neurons of third instar larval neuromuscular junctions. HRP (white) stains all neurites. e, Detection of Ca2+ levels in the Drosophila posterior midgut (fixed tissue). Ubiquitous expression of UAS-GCaMP3 (green) in the posterior midgut using Actin5C::GAL4 reveals high Ca2+ levels in subsets of small, basally located cells (likely to be ISCs and enteroblasts; asterisks), and at the brushed border of enterocytes (arrows). Superficial view in top panels, sagittal view in bottom panels. Right, Actin5C::GAL4 drives expression of mCD8-GFP homogeneously in the posterior midgut. Armadillo (red) marks cell boundaries. In enterocytes, Ca2+ is enriched in the microvilli (marked with arrows), potentially to facilitate Ca2+ dependent absorption and innate immune processes4. f, Ca2+ concentration is higher in ISCs than in enteroblasts. ISCs and enteroblasts are labelled by expression of mCherry (red) under the control of esg::GAL4. Ca2+ in these cells is detected using GCaMP3 (green). Su(H)Gbe-lacZ marks enteroblasts (white in top panels) and Dl marks ISCs (white in bottom panels). g, Quantification of relative Ca2+ levels in ISCs versus enteroblasts measured as fluorescence ratio between GCaMP3 and mCherry. Note the high variability of [Ca2+] in ISCs. Data are mean and s.e.m. P values from Student’s t-test. n = 56 cell pairs from 5 different guts. Four independent experiments were performed. One representative image from 5 flies in a single experiment (two independent duplicates) is shown in b, e and f. One representative image from 6 larvae in a single experiment (two independent duplicates) is shown in d.
Extended Data Figure 5 Ca2+ oscillations in ISCs.
a, Typical ex vivo recording of Ca2+ oscillations in ISCs from young flies. GCaMP3 (green) and mCherry (red) are expressed specifically in ISCs. A total of 56 frames with 15-s intervals are shown (see Methods for details). Frames were exported from ZEN software. Genotype: UAS-GCaMP3; esg::GAL4, UAS-mCherry; Su(H)Gbe::GAL80, tub::GAL80ts. b, Traces of Ca2+ oscillations in ISCs of intestines incubated with EGTA (5 mM) or 2-APB (10 μM) for 5–10 min and then recorded immediately. c, Ca2+ oscillation frequency of ISCs incubated in AHL with or without EGTA, CdCl2 (5 μM) or LaCl3 (5 mM). Values for individual ISCs (collected from 3–4 different guts) are plotted. d, Ca2+ oscillation in ISCs is inhibited by LaCl3. Scheme of oscillation parameters is shown on the left. The local amplitude of oscillation spikes and the baseline Ca2+ level between oscillations were derived from Gaussian fits on detected oscillation spikes. Average oscillation frequencies and average baselines are shown. e, Observed Ca2+ oscillation pattern in ISCs is independent of the genetic reporter system used. Oscillation frequency and average fluorescence ratio are compared between different genetic Calcium reporters: UAS-GCaMP3, UAS-tdTomato-P2A-GCaMP5 and UAS-IVS-GCaMP6s. Genetic reporters are expressed specifically in ISCs using esg::GAL4ts; Su(H)Gbe::GAL80. f, Ca2+ oscillation pattern in ISCs of guts incubated with or without isradipine (10 μg ml−1 incubation in AHL medium). Isradipine inhibits L-type VGCCs to paralyze visceral muscles. Data are mean and s.e.m. P value from ANOVA (c and e) or Student’s t-test (d, f). Individual ISCs pooled from 3–4 guts were plotted in c, e and f. The sample size for c is n = 12, 8, 9 and 14; for e, n = 12, 16, 13, 14, 20 and 24; and for f, n = 22, 14, 25 and 22 (from left to right for each panel). Two (b–d) and three (e, f) independent experiments were performed.
Extended Data Figure 6 Glutamate feeding modulates Ca2+ oscillation pattern through the mGluR/Gαq/PLCβ pathway.
a, Automated quantification of oscillation parameters in recordings from wild-type and mGluRRNAi expressing intestines refed L-Glu containing or control food (compare with Fig. 3a). Average oscillation frequencies and average baselines are shown. b, 10% L-Glu feeding inhibits Ca2+ oscillations. Ca2+ oscillations recorded in animals after 6–8 h refeeding with 10% L-Glu and incubation of the intestine in AHL supplemented with 10% L-Glu. Prolonged incubation (20 min) in 10% L-Glu causes further decrease of average GCaMP3 fluorescence. c, Manual and automated quantification of oscillation parameters on recordings from individual guts of the indicated genotypes. Values for individual cells from 3–4 guts are plotted on the left. Values for individual cells from single guts are shown in the third and fourth panels. Typical traces of recordings for individual cells are shown in the middle and the corresponding oscillation parameters calculated automatically by Gaussian fits in Mathematica 8.0 are shown on the right. Average oscillation frequencies and average baselines are shown. d, Gαq, but not other Gα subunits (GαO, GαS or Gαi) is required for Ca2+ oscillations in ISCs. While knockdown of Gαq, Gβ13F and PLCβ impairs Ca2+ oscillations, oscillation frequency was not affected when GαO, GαS or Gαi were knocked down in ISCs for 8 days at 29 °C (driver genotype: UAS-GCaMP3; esg::GAL4, UAS-mCherry, Su(H)Gbe::GAL80, tub::GAL80ts). Pertussis toxin (UAS-PTX) was used to inhibit Gαq specifically. Typical traces are shown on the right. e, Average oscillation frequencies and average baselines from Gaussian fits are shown. f, Voltage-gated calcium channel (VGCC) components Cavβ or Cac are not required for Ca2+ oscillations in ISCs. Oscillations are also not affected in ISCs expressing molecules expected to affect plasma membrane potential (ΔOrk1 (hyperpolarizing) and UAS-NaChBac (hypo-polarizing)). Driver genotype: UAS-GCaMP3; esg::GAL4, UAS-mCherry; Su(H)Gbe::GAL80, tub::GAL80ts. Data are mean and s.e.m. P values are from ANOVA (c (right), d and f) and Student’s t-test (a, b, c (left) and e). Individual ISCs pooled from 3–4 guts are plotted in b–d and f. The sample size for a and e is n = 3; for b n = 10, 8, 12, 17, 15, 20, 10 and 11; for c, n = 16, 20, 17 and 12; for d, n = 15, 18, 13, 22, 22, 10, 8 and 9; and for f, n = 13, 15, 15, 14 and 16 (from left to right for all panels). For c, values for individual cells from single guts are shown in the third and fourth panels. Data are representative of 2 (d–f) and 3 (a–c) independent experiments.
Extended Data Figure 7 Prolonged increase of cytosolic [Ca2+] promotes ISC proliferation.
a, Typical traces of Ca2+ recordings of indicated genotypes. Transgenes were induced at 29 °C for 4 days and 3–4 guts of each genotype were recorded. Thapsigargin concentration was 2 μM in the medium while recording. Genotypes: UAS-GCaMP3; esg::GAL4, UAS-mCherry; Su(H)Gbe::GAL80, tub::GAL80ts combined with w1118 for control or with the indicated transgenes. b, Knockdown of SERCA in ISCs, but not enteroblasts promotes ISC proliferation. SERCA was knocked down using esg::GAL4 (targeting ISCs and enteroblasts), Su(H)Gbe::GAL4 (enteroblasts), or esg::GAL4; Su(H)Gbe-GAL80 (ISCs) combined with tub::GAL80ts by incubating at 29 °C for 4 days. ISCs and/or enteroblasts are labelled by nuclear GFP (UAS-nls-GFP) in green and mitotic ISCs are stained by pH3 (red). Number of dividing ISCs (pH3+) per gut were quantified and analysed. c, Knocking down SERCA or PMCA promotes growth of ISC-derived clones. Typical MARCM clones expressing SERCARNAi, PMCARNAi and StimRNAi 7 days after induction are shown. ISC clones are marked in green and pH3 staining was used to detect dividing ISCs. Note: SERCARNAi clones contain many more, but smaller cells, resulting in similar clonal area compared to wild type. Quantification is shown on the right. d, Left, mitotic figures quantified in intestines of flies homozygous for SERCAkum170, a temperature-sensitive SERCA loss-of-function allele. Flies were exposed to heat shock (42 °C, 5 min for two consecutive days, this permanently inactivates SERCA54), and proliferation was assessed 7 days after heat shock. Right, size of MARCM clones of ISCs homozygous for SERCAkum170 analysed 7 days after clone induction. e, MARCM clones of ISCs homozygous for the Stim null allele, StimA, analysed 7 days after clone induction. Dashed lines delineate individual clones (GFP, green; DAPI, blue, armadillo, red membrane; prospero, red nuclei; Dl, white). Dl channel is shown separately in greyscale. Quantification of clone sizes (cells per clone) is shown on the right. f, Guts of the indicated genotypes visualized in basal views (six panels) and in cross section (two panels). In basal views, ISCs are identified by Dl immunostaining (red) and GFP (green). In cross sections, visceral muscle is identified by phalloidin staining (red), and ISCs/enteroblasts by GFP (green). Data are mean and s.e.m. P value from ANOVA (c) or Student’s t-test (b, d, e). For mitotic figures in b, n = 11 for each genotype. For mitotic figures in d (left), n = 10 for each condition. For clonal analysis in c, clones (FRT40A, n = 50; SERCARNAi, n = 42; StimRNAi, n = 40; PMCARNAi, n = 56) from 8–10 guts were quantified. For clonal analysis in d (right), clones (FRT42D, n = 80; SERCAkum170, n = 60) from 10 guts were quantified. For clonal analysis in e (FRT19A, n = 120; StimA, n = 102) from 7 guts were assessed. For a, traces from individual ISCs from a representative gut of each indicated genotype are shown, 3–4 guts were recorded for each experiment and 2 independent experiments were performed. For b–f, 1 of 3 independent experiments is presented.
Extended Data Figure 8 Time-course analysis of Notch activity in ISC lineage after manipulation of Ca2+ signalling.
a, ISC lineage. After an asymmetric division, ISCs (expressing Delta) activate Notch in enteroblasts (thus activating Su(H)Gbe-lacZ). Enteroblasts with high Notch activity differentiate into polyploid enterocytes (expressing Pdm1), enteroblasts with low Notch activity differentiate into enteroendocrine cells (expressing prospero). b, ISC proliferation rates (mitotic nuclei per gut) at different time points after perturbation of Notch (N) or Ca2+ signalling. Number of days after shift to 29 °C is listed. c–e, Representative images of guts perturbed as in b, immunostained for Su(H)Gbe-lacZ (bGal, red, reporter for Notch activity) and Dl (white). GFP, green; DAPI, blue. For 7-day time point, higher magnification images (boxed area) of bGal channel are shown in bottom row in greyscale. Note that knockdown of Notch results in rapid loss of Su(H)Gbe-lacZ+ cells and accumulation of Dl+ cells. SERCA knockdown results in loss of Su(H)Gbe-lacZ only after prolonged expression (14 days), even though proliferation is induced more strongly than with NotchRNAi already at 4 days. Su(H)Gbe-lacZ expression is not lost in PMCARNAi or CrtcOE guts, although proliferation is induced as strongly as in NotchRNAi-expressing guts at 4 days. Quantification of ratio of Dl+ cells and Gbe-lacZ+ cells in the gut after at the indicated time points after shift to 29 °C is shown in e. f, Quantification of prospero+ cells of posterior midguts in which cytosolic Ca2+ was increased by knocking down SERCA or PMCA, or by overexpressing Stim and Orai using esgts. Increased numbers of enteroendocrine cells (an indication of impaired Notch signalling) were only observed in animals in which SERCA was knocked down for a prolonged period of time (14 days, 29 °C). Anti-prospero stains enteroendocrine cell nuclei. g, Clonal analysis of ISC differentiation process after manipulation of Ca2+ signalling or Notch activity. ISC MARCM clones of control, NotchRNAi, SERCARNAi or PMCARNAi were analysed at 4 or 7 days after heat-shock (AHS) induction. Clones (marked in GFP) are circled in dashed line. The nuclei of differentiated cells are stained by Pdm1 in red. Although PMCARNAi clones are significantly larger, the differentiation process is largely normal based on Pdm1 staining. Whereas differentiation of SERCARNAi clones at 7 days after heat-shock is significantly perturbed based on Pdm1 staining, as compared with surrounding wild-type enterocytes. As expected, no Pdm1+ cells were observed in NotchRNAi clones. h, Related to g, clone size and percentage of Pdm1+ cells per clone was quantified 4 days after heat-shock induction. i, NotchRNAi induced proliferation can be partially rescued by knocking down Ca2+ signalling components, such as Stim or CanB2. Mitotic index shown on right. j, Inhibition of SERCA stimulates ISC proliferation even when Notch signalling is induced by overexpression of NotchICD (Notch intracellular domain). Data are mean and s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (ANOVA in b, e, f, h and i; Student’s t-test in b (between SERCARNAi and SERCARNAi; STIMRNAi), f (between control and PMCARNAi), and j). For b, mitotic cells are calculated at indicated time points for each genotype: 4 days, control (n = 16), PMCARNAi (8), CrtcOE (9), NotchRNAi (12), SERCARNAi (14), SERCARNAi; StimRNAi (9); 7 days, control (12), PMCARNAi (10), CrtcOE (9), NotchRNAi (11), SERCARNAi (10), SERCARNAi; StimRNAi (8); 14 days, control (6), PMCARNAi (18), CrtcOE (9), NotchRNAi (12), SERCARNAi (8), SERCARNAi; StimRNAi (11). For c–e, cells in posterior midgut from 4–7 guts of each genotype and time point were analysed. For e, fraction of bGal+ or Dl+ cells in 100–200 total cells counted in a field of the posterior midgut for each condition was quantified. Data are mean and s.e.m. for the following number of guts (left to right) n = 4, 4, 5, 4, 4, 4, 4, 7, 4, 6, 4, 4, 4, 5, 4, 4, 4, 5, 4, 4, 5, 4, 6, 5, 4, 4, 5, 4, 5, 5, 6, 4, 5, 7, 5, 6, 4 and 5. For f, prospero+ cells of indicated genotype at 4 days (n = 5 guts) and 14 days (n = 7 guts) were analysed. For clonal analysis in g and h, clones (FRT40A, n = 50; SERCARNAi, n = 45; NotchRNAi, n = 30; and PMCARNAi, n = 43) were analysed. For mitotic analysis in i and j, n = 12 for each genotype. Data in b–f and i are representative of 2 independent experiments; data in g, h and j represent one of triplicate experiments.
Extended Data Figure 9 Cytosolic Ca2+ regulates ISC proliferation through CaN/CRTC pathway.
a, Acute knockdown of SERCA in ISCs does not induce ER stress. Phospho-eIF2α (red, a PERK-mediated phosphorylation and marker of ER stress; see also ref. 40), is only weakly detected in control ISCs. No significant increase of phospho-eIF2α staining is observed in ISCs expressing SERCARNAi, whereas tunicamycin treatment (an inducer of ER stress; 50 μM, 24 h) increases phospho-eIF2α strongly in ISCs (asterisk). Quantification of average fluorescent intensity of phospho-eIF2α shown on right. b, Increasing ER folding capacity by overexpressing spliced Xbp1 (Xbp1s) does not limit ISC proliferation in SERCA loss-of-function conditions. Representative images shown on the left. Mitotic ISCs stained by anti-pH3 staining (red). Mitotic index quantified on the right. c, Increased cytosolic [Ca2+] induces proliferation without perturbing Egfr pathway activity. Indicated Ca2+ signalling components were knocked down or overexpressed using esgts;UAS-nls-GFP. Diphospho-ERK (dpERK) staining as a readout of Egfr pathway activity was quantified after 4-day induction at 29 °C. InR was overexpressed using esgts;UAS-mCD8-GFP as a positive control. d, Proliferation of SERCA-deficient ISCs is suppressed when CaN subunits are silenced simultaneously, but not when CaMKI and CaMKII are knocked down. e, Proliferation of ISCs in which cytosolic [Ca2+] is increased by overexpression of Stmi and Orai, or by knockdown of PMCA, is rescued when CaNB2 is silenced simultaneously. f, Overexpression of constitutive active forms of CaN catalytic subunits (CanA14F, Pp2B14D, CanA1) promotes ISC proliferation. Mitotic figures were quantified after 4 days of transgene expression at 29 °C. g, CaN is required for growth of ISC lineages. Quantification of clone sizes of MARCM clones homozygous for the null allele CanB2KO, or expressing dsRNA against CanB2 (CanB2RNAi), or constitutively active CanA14Fact. h, Loss of Crtc (homozygosity for null allele crtc25-3) rescues increased ISC proliferation when PMCA is knocked down or when Pp2B-14Dact is overexpressed using esg::GAL4, tub::GAL80ts (esgts). i, Overexpression of Crtc promotes ISC proliferation. Mitotic index of intestines overexpressing haemagglutinin (HA)-tagged forms of wild-type Crtc (UAS-CRTC-HA) or constitutively nuclear forms of Crtc (UAS-CRTC-SA-HA) using esg::GAL4ts was analysed after 4 days of incubation at 29 °C. j, Increasing cytosolic Ca2+ promotes ISC proliferation via the Ca2+/calcineurin/Crtc pathway. Guts of indicated genotypes were dissected and stained with anti-pH3 (indicating mitotic ISCs), anti-armadillo (arm, labelling cell boundaries) and anti-prospero (pro, labelling enteroendocrine cells). k, Crtc overexpression is sufficient to promote ISC proliferation when Gαq or InsP3R are silenced. l, CREB and its partner CBP are required for ISC survival, while overexpressing CREB promotes proliferation. Representative images of intestines in which CREB and CBP were genetically perturbed in ISC/enteroblasts using esg::GAL4. Increased numbers of GFP+ ISCs/enteroblasts are observed when CREB is overexpressed, while a significant loss of GFP+ cells is observed when CREB or CBP are knocked down. Guts are stained for armadillo (membrane, white) and prospero (nuclear, white) to identify enteroendocrine cells and enterocytes; DAPI, blue. Genotype: esg::GAL4; UAS-nls-GFP; tub::GAL80ts/UAS-X. m, Knockdown efficiency of RNAi lines determined by qRT–PCR. PMCARNAi (BL31572) and CanB2RNAi (BL27270) were used to knock down respective genes in the gut using NP1::GAL4, tub::GAL80ts (29 °C for 10 days). mGluRRNAi (BL41668) was used to knock down mGluR in the brain using elav::GAL4. Expression levels were normalized using Act5C and to uninduced controls (NP1::GAL4/+; tub::GAL80ts/+ or elav::GAL4/+). Effectiveness of other constructs used has been reported in the literature: UAS-GαqRNAi and UAS-PLCβRNAi were obtained from and verified by Ha et al.55, UAS-CamKIRNAi (BL26726), UAS-CamKIIRNAi (BL29401), UAS-InsP3RRNAi (BL25937), and UAS-RyRRNAi (BL28919) were verified by Shim et al.23, and UAS-SERCARNAi (BL44581) by Roti et al.29. Data are mean and s.e.m. P values from Student’s t-test (b, g and h) and ANOVA (a, c, d–f, i and k). For a and c, fluorescence intensities for phospho-eIF2α (a) or diphospho-ERK (c) in 30–50 ISCs/enteroblasts doublets in single fields of several independent posterior midguts were averaged for each condition. In a, control, n = 5 guts, SERCARNAi, n = 5, tunicamycin, n = 6; in c, control, n = 6, SERCARNAi, n = 4, CRTCOE, n = 9, and PMCARNAi, n = 8. For mitotic analysis, in: b, n = 13 guts for control, n = 10 for the rest; d, n = 17 for each genotype; e, n = 18 for control, n = 12 for the rest; f and k, n = 11 each genotype; and i, n = 12 for each condition. For clonal analysis in g, clones (FRT42D, n = 56 clones; CanB2KO, n = 60; FRT40A, n = 50; CanB2RNAi, n = 70; FRT82B, n = 62; CanA14FACT, n = 58) from 10 guts each were assessed. One representative image from 10 flies in a single experiment (two independent experiments) is shown in j and l. Data in a–i, and k are representative of three independent experiments. Data in m is from n = 3 technical replicates of samples pooled from 10 guts each for PMCA and CanB2, 4 heads each for mGluR. Representative of two independent experiments.
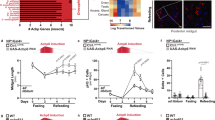
Extended Data Figure 10 Ca2+ oscillation pattern as an indicator of ISC proliferation status.
a, Typical traces of live recordings of indicated genotypes. Genotype for GCaMP3 control: w1118 X UAS-GCaMP3; esg::GAL4, UAS-mCherry; Su(H)Gbe::GAL80, tub::GAL80ts. Genotype for bicistronic control: w1118 X; esg::GAL4, UAS-tdTomato-P2A-GCaMP5;Su(H)Gbe::GAL80, tub::GAL80ts. b, Ca2+ oscillation patterns of ISCs in which proliferation was stimulated or inhibited by genetic or environmental perturbations: knockdown of Notch, overexpression of InR, Unpaired2 or RasV12, infection with Ecc15, or ageing results in high proliferative activity. Overexpression of InRDN and CncC inhibits proliferation of ISCs. c, Acute Ecc15 infection transiently increases cytosolic [Ca2+] while decreasing oscillations in ISCs. d, Acute Ecc15 infection increases cytosolic Ca2+ while decreasing oscillations in ISCs as determined using the bi-cistronic calcium reporter UAS-tdTomato-P2A-GCaMP5G. e, ISCs in which proliferation is impaired exhibit more frequent oscillations than controls. Oscillation frequency of ISCs from indicated genotypes is plotted individually (three guts for each genotype and each dots represents one ISC). f, Expressing InRDN in ISCs is sufficient to inhibit stress- or diet-induced proliferation. Quantification of mitotic figures of indicated genotype is shown. For bleomycin treatment, flies were dry-starved for 4 h before feeding on 25 μg ml−1 (final) bleomycin for 24 h. For refeeding, Flies were maintained on normal food for 4 days at 29 °C, then starved for 2 days, and refed with yeast-supplemented food. Genotype: esg::GAL4, UAS-GFP; Su(H)Gbe::GAL80, tub::GAL80ts/UAS-InRDN. g, ISC proliferation induced by oral infection with Ecc15 or by bleomycin treatment is suppressed by silencing InsP3R or Orai. Mitotic figures were analysed 6 h after oral infection with Ecc15 or 24 h after feeding with bleomycin. h, Increased cytosolic [Ca2+] in ISCs activated by oral infection with Ecc15 or by bleomycin treatment. This increase is suppressed by silencing InsP3R or Orai. i, ISC proliferation induced by bleomycin treatment or Ecc15 infection is suppressed by silencing Crtc or InsP3R or CanB2, or in crtc25-3 homozygous mutants. mGluR, in turn, is not required for bleomycin-induced proliferation. Mitotic figures were analysed 24 h after feeding with bleomycin or 6 h after oral infection with Ecc15. j, Quantification of mitotic figures in animals of the indicated genotypes. ISC proliferation induced by overexpression of InR or Hep can be suppressed by knockdown of InsP3R, Crtc, Stim or CanB2 (see also Fig. 4a). k, mGluR is not required for Ecc15-induced proliferation and changes in cytosolic Ca2+. Mitotic figures and Ca2+ oscillation patterns analysed 6 h after oral infection. l, Increasing cytosolic [Ca2+] promotes ISCs proliferation in Jak/Stat loss-of-function conditions. Jak/Stat pathway (Dome and Hop) is required for ISC proliferation induced by Ecc15 infection. Increasing cytosolic [Ca2+] by knocking down PMCA (left) or SERCA (right) is sufficient to rescue ISC proliferation. m, Knocking down Fos can substantially suppress Crtc overexpression induced ISC proliferation. n, Left, segregation of active and resting ISCs into Ca2+ oscillation modes as calculated by automatic peak detection and Gaussian fits. As shown for ‘manual’ calculations in Fig. 4, Ca2+ oscillation patterns segregate into two modes associated with the proliferative status of the ISCs. ISCs in which the core components of Ca2+ homeostatic machinery are perturbed exhibit both low oscillation frequency and low average signal intensity (bottom left corner). Transition from quiescence to active proliferation after L-Glu feeding is indicated by the blue arrow. Right, ISC activity does not segregate when local oscillation amplitudes are plotted against oscillation frequency, suggesting that the primary driver of ISC proliferation is not the amplitude of individual Ca2+ spikes, but the increase in basal or average cytosolic Ca2+ concentration within ISC. Data are mean and s.e.m. P values from ANOVA (b, h, i, j (left) and l (right)) and Student’s t-test (c, d, f, g, k, m, j (right) and l (left)). For Ca2+ recordings in b–e, h and k, individual ISCs pooled from 3–4 guts were plotted. The sample size for b is n = 10, 14, 14, 12, 16, 18, 9, 12, 11, 20, 22, 38, 29, 12, 19, 8 and 12; for c, n = 7, 8, 9, 7, 13, 18, 10 and 7; for d, n = 14, 16, 15 and 18; for e, n = 5, 4, 5, 9, 4, 6, 8, 6 and 5; for h, n = 12, 11, 12, 11, 12, 14, 12, 11, 17, 12, 14, 11, 19, 9, 10 and 8; for k, n = 11, 12, 11, 11, 13, 10, 11 and 11 (from left to right for all panels). For mitotic analysis in f and g, n = 12, 10, 10, 10, 12, 11, 10, 12, 12, 14, 10, 9, 9, 10, 9, 9, 10 and 13; and in i and m, n = 13, 12, 11, 18, 11, 18, 12, 13, 13, 12, 10, 10, 13, 9, 12, 12, 16, 12, 9, 10, 10, 8, 9, 9, 10, 9, 12, 13, 13, 14, 14, 15, 12, 10, 19, 10, 12, 10, 9, 8, 8, 9 and 10 (from left to right for all panels). Data in a–h, j and m are representative of three independently performed experiments, and those shown in i, k and l are a composite from two separate experiments.
Supplementary information
Supplementary Information
This file PDF contains a Supplementary Discussion with detailed description and interpretation of the effects of specific perturbations shown in Figure 3 and Extended Data Figures 5 and 7-9. It also contains a complete list of genotypes for each Figure. (PDF 340 kb)
Cytosolic Ca2+ oscillates regularly in control young guts raised on conventional food
Guts from flies incubated for 4 days at 29 °C were dissected and recorded in AHL medium using two photon microscopy. The videos was composed using NIH Image J. For this and the following videos, brightness of the RFP and GFP channels were adjusted for visibility. For quantification of oscillation parameters, fluorescence intensities were not adjusted and gut movements were manually or automatically compensated for using Image Analyst MKII (See methods for details). 100 frames with 15s intervals are shown. Video 2 was recorded using the bicistronic Calcium reporter. Genotype Video 1: esg::Gal4, UAS::mCherry, UAS::GCaMP3; Su(H)Gbe::Gal80, tub::Gal80ts. Genotype Video 2: esg::Gal4, UAS::tdTomato-2A-GCaMP5; Su(H)Gbe::Gal80, tub::Gal80ts. (MP4 1857 kb)
Cytosolic Ca2+ oscillates regularly in control young guts raised on conventional food
Guts from flies incubated for 4 days at 29 °C were dissected and recorded in AHL medium using two photon microscopy. Videos were composed using NIH Image J. For this and the following videos, brightness of the RFP and GFP channels were adjusted for visibility. For quantification of oscillation parameters, fluorescence intensities were not adjusted and gut movements were manually or automatically compensated for using Image Analyst MKII (See methods for details). 100 frames with 15s intervals are shown. Video 2 was recorded using the bicistronic Calcium reporter. Genotype Video 1: esg::Gal4, UAS::mCherry, UAS::GCaMP3; Su(H)Gbe::Gal80, tub::Gal80ts. Genotype Video 2: esg::Gal4, UAS::tdTomato-2A-GCaMP5; Su(H)Gbe::Gal80, tub::Gal80Mts (AVI 221 kb)
L-Glutamate feeding reduces Ca2+ oscillation frequency and increases the average GCaMP3 signal intensity.
Freshly hatched files were starved for 2 days at 29 ºC before transferring to yeast restricted food. Video 3 was recorded from flies fed yeast-restricted food with no Glutamate (0% L-Gu + 0.1% Yeast) for 4 days (100 frames, 15s intervals), and Video 4 was recorded from flies maintained on L-Glu supplemented yeast restricted food (1% L-Gu+ 0.1% Yeast) for 4 days (100 frames, 15s intervals). (MOV 555 kb)
L-Glutamate feeding reduces Ca2+ oscillation frequency and increases the average GCaMP3 signal intensity
Freshly hatched files were starved for 2 days at 29 ºC before transferring to yeast restricted food. Video 3 was recorded from flies fed yeast-restricted food with no Glutamate (0% L-Gu + 0.1% Yeast) for 4 days (100 frames, 15s intervals), and Video 4 was recorded from flies maintained on L-Glu supplemented yeast restricted food (1% L-Gu+ 0.1% Yeast) for 4 days (100 frames, 15s intervals). (MP4 1547 kb)
mGluR is required for L-Glutamate induced changes in Ca2+ oscillations.
Flies in which mGluR was specifically knocked down in ISCs were incubated and re-fed as described for Supplementary Videos 3-4. Genotype: esg::Gal4, UAS::mCherry, UAS::GCaMP3; mGluRRNAi, Su(H)Gbe::Gal80, tub::Gal80ts. Video 5 is without glutamate (100 frames, 15s intervals), and video 6 is with 1% glutamate (100 frames, 15s intervals). (MP4 1851 kb)
mGluR is required for L-Glutamate induced changes in Ca2+ oscillations.
Flies in which mGluR was specifically knocked down in ISCs were incubated and re-fed as described for Supplementary Videos 3-4. Genotype: esg::Gal4, UAS::mCherry, UAS::GCaMP3; mGluRRNAi, Su(H)Gbe::Gal80, tub::Gal80ts. Video 5 is without glutamate (100 frames, 15s intervals), and video 6 is with 1% glutamate (100 frames, 15s intervals). (MP4 1863 kb)
Gαq is required for Ca2+ oscillations in ISCs.
Intact guts from flies raised on conventional food were imaged after transgenes were induced for 8 days at 29 °C. Genotype: esg::Gal4, UAS::mCherry, UAS::GCaMP3; GαqRNAi, Su(H)Gbe::Gal80, tub::Gal80ts (67 frames, 15s intervals). (MP4 1303 kb)
Knock-down of SERCA inhibits Ca2+ oscillations in ISCs.
Intact guts from flies raised on conventional food were imaged after transgenes were induced for 4 days at 29 °C. Genotype: esg::Gal4, UAS::mCherry, UAS::GCaMP3; SercaRNAi, Su(H)Gbe::Gal80, tub::Gal80ts (100 frames, 15s intervals). (MP4 1819 kb)
Knock-down of STIM inhibits Ca2+ oscillations in ISCs.
Intact guts from flies raised on conventional food were imaged after transgenes were induced for 4 days at 29 °C. Genotype: esg::Gal4, UAS::mCherry, UAS::GCaMP3; StimRNAi, Su(H)Gbe::Gal80, tub::Gal80ts (100 frames, 15s intervals). (MOV 549 kb)
Over-expression of InRDN increases Ca2+ oscillations in ISCs.
Intact guts from flies raised on conventional food were imaged after transgenes were induced for 4 days at 29 °C. Genotype: esg::Gal4, UAS::mCherry, UAS::GCaMP3; InRDN, Su(H)Gbe::Gal80, tub::Gal80ts (100 frames, 15s intervals). (MP4 1242 kb)
Rights and permissions
About this article
Cite this article
Deng, H., Gerencser, A. & Jasper, H. Signal integration by Ca2+ regulates intestinal stem-cell activity. Nature 528, 212–217 (2015). https://doi.org/10.1038/nature16170
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16170
This article is cited by
-
Calcineurin stimulation by Cnb1p overproduction mitigates protein aggregation and α-synuclein toxicity in a yeast model of synucleinopathy
Cell Communication and Signaling (2023)
-
Gut AstA mediates sleep deprivation-induced energy wasting in Drosophila
Cell Discovery (2023)
-
IFNγ-Stat1 axis drives aging-associated loss of intestinal tissue homeostasis and regeneration
Nature Communications (2023)
-
Cholinergic neurons trigger epithelial Ca2+ currents to heal the gut
Nature (2023)
-
The CRTC-CREB axis functions as a transcriptional sensor to protect against proteotoxic stress in Drosophila
Cell Death & Disease (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.