Abstract
Males and females share many traits that have a common genetic basis; however, selection on these traits often differs between the sexes, leading to sexual conflict1,2. Under such sexual antagonism, theory predicts the evolution of genetic architectures that resolve this sexual conflict2,3,4,5. Yet, despite intense theoretical and empirical interest, the specific loci underlying sexually antagonistic phenotypes have rarely been identified, limiting our understanding of how sexual conflict impacts genome evolution3,6 and the maintenance of genetic diversity6,7. Here we identify a large effect locus controlling age at maturity in Atlantic salmon (Salmo salar), an important fitness trait in which selection favours earlier maturation in males than females8, and show it is a clear example of sex-dependent dominance that reduces intralocus sexual conflict and maintains adaptive variation in wild populations. Using high-density single nucleotide polymorphism data across 57 wild populations and whole genome re-sequencing, we find that the vestigial-like family member 3 gene (VGLL3) exhibits sex-dependent dominance in salmon, promoting earlier and later maturation in males and females, respectively. VGLL3, an adiposity regulator associated with size and age at maturity in humans, explained 39% of phenotypic variation, an unexpectedly large proportion for what is usually considered a highly polygenic trait. Such large effects are predicted under balancing selection from either sexually antagonistic or spatially varying selection9,10. Our results provide the first empirical example of dominance reversal allowing greater optimization of phenotypes within each sex, contributing to the resolution of sexual conflict in a major and widespread evolutionary trade-off between age and size at maturity. They also provide key empirical evidence for how variation in reproductive strategies can be maintained over large geographical scales. We anticipate these findings will have a substantial impact on population management in a range of harvested species where trends towards earlier maturation have been observed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
Data deposits
Details of the SNPs used in the study have been deposited in dbSNP (http://www.ncbi.nlm.nih.gov/SNP/) under accession numbers ss1867919552–ss1868858426, and re-sequencing data have been deposited in EMBL Nucleotide Sequence Database (European Nucleotide Archive) under accession number PRJEB10744. SNP genotype and phenotype data and detailed DNA sequence information of the main candidate gene regions are available in Dryad (http://dx.doi.org/10.5061/dryad.23h4q).
References
Bonduriansky, R. & Chenoweth, S. F. Intralocus sexual conflict. Trends Ecol. Evol. 24, 280–288 (2009)
Lande, R. Sexual dimorphism, sexual selection, and adaptation in polygenic characters. Evolution 34, 292–305 (1980)
Rice, W. Sex chromosomes and the evolution of sexual dimorphism. Evolution 38, 1416–1424 (1984)
Fisher, R. A. The Genetical Theory of Natural Selection 139–142 (Oxford Univ. Press, 1930)
van Doorn, G. S. Intralocus sexual conflict. Ann. NY Acad. Sci . 1168, 52–71 (2009)
Fry, J. D. The genomic location of sexually antagonistic variation: some cautionary comments. Evolution 64, 1510–1516 (2010)
Turelli, M. & Barton, N. H. Polygenic variation maintained by balancing selection: pleiotropy, sex-dependent allelic effects and G × E interactions. Genetics 166, 1053–1079 (2004)
Schaffer, W. M. in Evolution Illuminated: Salmon and Their Relatives (eds Stearns, S. & Hendry, A. ) 20–51 (Oxford Univ. Press, 2004)
Connallon, T. & Clark, A. G. Balancing selection in species with separate sexes: insights from Fisher’s geometric model. Genetics 197, 991–1006 (2014)
Savolainen, O., Lascoux, M. & Merilä, J. Ecological genomics of local adaptation. Nature Rev. Genet. 14, 807–820 (2013)
Dobzhansky, T. A review of some fundamental concepts and problems of population genetics. Cold Spring Harb. Symp. Quant. Biol. 20, 1–15 (1955)
Mokkonen, M. et al. Negative frequency-dependent selection of sexually antagonistic alleles in Myodes glareolus. Science 334, 972–974 (2011)
Sellis, D., Callahan, B. J., Petrov, D. A. & Messer, P. W. Heterozygote advantage as a natural consequence of adaptation in diploids. Proc. Natl Acad. Sci. USA 108, 20666–20671 (2011)
Connallon, T. & Clark, A. G. Evolutionary inevitability of sexual antagonism. Proc. R. Soc. Lond. B 281, 20132123 (2014)
Kidwell, J. F., Clegg, M. T., Stewart, F. M. & Prout, T. Regions of stable equilibria for models of differential selection in the two sexes under random mating. Genetics 85, 171–183 (1977)
Pennell, T. M. & Morrow, E. H. Two sexes, one genome: the evolutionary dynamics of intralocus sexual conflict. Ecol. Evol . 3, 1819–1834 (2013)
Stearns, S. C. Life history evolution: successes, limitations, and prospects. Naturwissenschaften 87, 476–486 (2000)
Fleming, I. A. & Einum, S. in Atlantic Salmon Ecology 33–65 (Wiley-Blackwell, 2011)
Hutchings, J. A. & Jones, M. E. B. Life history variation and growth rate thresholds for maturity in Atlantic salmon, Salmo salar. Can. J. Fish. Aquat. Sci . 55 (Suppl. 1), 22–47 (1998)
Halperin, D. S., Pan, C., Lusis, A. J. & Tontonoz, P. Vestigial-like 3 is an inhibitor of adipocyte differentiation. J. Lipid Res. 54, 473–481 (2013)
Perry, J. R. B. et al.; Australian Ovarian Cancer Study; GENICA Network; kConFab; LifeLines Cohort Study; InterAct Consortium; Early Growth Genetics (EGG) Consortium. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature 514, 92–97 (2014)
Cousminer, D. L. et al.; ReproGen Consortium; Early Growth Genetics (EGG) Consortium. Genome-wide association and longitudinal analyses reveal genetic loci linking pubertal height growth, pubertal timing and childhood adiposity. Hum. Mol. Genet. 22, 2735–2747 (2013)
Taranger, G. L. et al. Control of puberty in farmed fish. Gen. Comp. Endocrinol. 165, 483–515 (2010)
Thorpe, J., Mangel, M., Metcalfe, N. & Huntingford, F. Modelling the proximate basis of salmonid life-history variation, with application to Atlantic salmon, Salmo salar L. Evol. Ecol. 12, 581–599 (1998)
Reinton, N. et al. Localization of a novel human A-kinase-anchoring protein, hAKAP220, during spermatogenesis. Dev. Biol. 223, 194–204 (2000)
Lee, B. et al. Direct transcriptional regulation of Six6 is controlled by SoxB1 binding to a remote forebrain enhancer. Dev. Biol. 366, 393–403 (2012)
Flatt, T. & Heyland, A. (eds) Mechanisms of Life History Evolution: The Genetics and Physiology of Life History Traits and Trade-Offs (Oxford Univ. Press, 2011)
Woram, R. A. et al. Comparative genome analysis of the primary sex-determining locus in salmonid fishes. Genome Res. 13, 272–280 (2003)
Chaput, G. Overview of the status of Atlantic salmon (Salmo salar) in the North Atlantic and trends in marine mortality. ICES J. Mar. Sci. 69, 1538–1548 (2012)
Allendorf, F. W. & Hard, J. J. Human-induced evolution caused by unnatural selection through harvest of wild animals. Proc. Natl Acad. Sci. USA 106 (Suppl. 1), 9987–9994 (2009)
Bourret, V. et al. SNP-array reveals genome-wide patterns of geographical and potential adaptive divergence across the natural range of Atlantic salmon (Salmo salar). Mol. Ecol. 22, 532–551 (2013)
Karlsson, S., Diserud, O. H., Moen, T. & Hindar, K. A standardized method for quantifying unidirectional genetic introgression. Ecol. Evol . 4, 3256–3263 (2014)
Aykanat, T. et al. Low but significant genetic differentiation underlies biologically meaningful phenotypic divergence in a large Atlantic salmon population. Mol. Ecol. 24, 5158–5174 (2015)
Johnston, S. E. et al. Genome-wide SNP analysis reveals a genetic basis for sea-age variation in a wild population of Atlantic salmon (Salmo salar). Mol. Ecol. 23, 3452–3468 (2014)
Yano, A. et al. The sexually dimorphic on the Y-chromosome gene (sdY) is a conserved male-specific Y-chromosome sequence in many salmonids. Evol. Appl . 6, 486–496 (2013)
Friedland, K. D. & Haas, R. E. Marine post-smolt growth and age at maturity of Atlantic salmon. J. Fish Biol. 48, 1–15 (1996)
Fisher, J. P. & Pearcy, W. G. Spacing of scale circuli versus growth-rate in young Coho salmon. Fish Bull. 88, 637–643 (1990)
ICES. Report of the Workshop on Age Determination of Salmon (WKADS). Report CM 2011/ACOM:44 (ICES, 2011)
Einum, S., Thorstad, E. B. & Næsje, T. F. Growth rate correlations across life-stages in female Atlantic salmon. J. Fish Biol. 60, 780–784 (2002)
Jonsson, N. & Jonsson, B. Sea growth, smolt age and age at sexual maturation in Atlantic salmon. J. Fish Biol. 71, 245–252 (2007)
Gjedrem, T., Gjøen, H. M. & Gjerde, B. Genetic origin of Norwegian farmed Atlantic salmon. Aquaculture 98, 41–50 (1991)
GenABEL Project Developers. GenABEL: genome-wide SNP association analysis. R package version 1.8-0 (2013)
R Core Team. R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2014)
Christensen, R. H. B. ordinal - regression models for ordinal data. R package version 2015.1-21 (2015)
Aschard, H., Vilhjálmsson, B. J., Joshi, A. D., Price, A. L. & Kraft, P. Adjusting for heritable covariates can bias effect estimates in genome-wide association studies. Am. J. Hum. Genet. 96, 329–339 (2015)
Bonhomme, M. et al. Detecting selection in population trees: the Lewontin and Krakauer test extended. Genetics 186, 241–262 (2010)
Lotterhos, K. E. & Whitlock, M. C. Evaluation of demographic history and neutral parameterization on the performance of FST outlier tests. Mol. Ecol. 23, 2178–2192 (2014)
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, v067.i01 (2015)
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010)
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at http://arXiv.org/abs/1207.3907 (2012)
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012)
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nature Methods 7, 248–249 (2010)
Browning, B. L. & Browning, S. R. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 194, 459–471 (2013)
Sabeti, P. C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002)
Gautier, M. & Vitalis, R. rehh: an R package to detect footprints of selection in genome-wide SNP data from haplotype structure. Bioinformatics 28, 1176–1177 (2012)
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006)
Fijarczyk, A. & Babik, W. Detecting balancing selection in genomes: limits and prospects. Mol. Ecol. 24, 3529–3545 (2015)
Messer, P. W. & Petrov, D. A. Population genomics of rapid adaptation by soft selective sweeps. Trends Ecol. Evol. 28, 659–669 (2013)
Ferrer-Admetlla, A., Liang, M., Korneliussen, T. & Nielsen, R. On detecting incomplete soft or hard selective sweeps using haplotype structure. Mol. Biol. Evol. 31, 1275–1291 (2014)
Garud, N. R., Messer, P. W., Buzbas, E. O. & Petrov, D. A. Recent selective sweeps in North American Drosophila melanogaster show signatures of soft sweeps. PLoS Genet. 11, e1005004 (2015)
Acknowledgements
We thank L. Andersson, T. F. Hansen and H. Granroth-Wilding for commenting on earlier drafts of the manuscript. We also acknowledge the numerous fishers who contributed scales and phenotypic information. We thank J. Haantie, J. G. Jensås, J. Kuusela, I. Torvi and G. Østborg for scale measurements, T. Andersstuen, T. Balstad, L. Birkeland Eriksen, S. Karoliussen, J. Kuismin, M. Lindqvist, T. Pajula, K. Salminen, K. Sõstar, M. Spets and K. Vagonyte-Hallan for laboratory assistance, M. Ellmen, O. Guttorm, T. Kanniainen, A. Koskinen, T. Pöyhönen and S. Uusi-Heikkilä for sampling assistance, and T. Mulugeta for informatics support. Bioinformatic analyses used resources at the Finnish Centre for Scientific Computing, the Abel Cluster, owned by the University of Oslo and the Norwegian Metacenter for High Performance Computing, and operated by the Department for Research Computing at the University of Oslo IT Department and the Orion Computing Cluster at CIGENE. This study was funded by the Finnish Academy (grants 137710, 141231, 272836, 284941), the Research Council of Norway (QuantEscape, grant 216105 and RCN-project 221734/O30) and by AquaGen (SNP array development).
Author information
Authors and Affiliations
Contributions
C.R.P., S.L., N.J.B., T.A. and K.H. conceived the study. C.R.P., S.L., N.J.B., T.A., K.H., C.J., S.K. and S.E.J. designed the experiments. T.M. led the development of the 220K SNP array, and M.K. and T.N. generated and conducted bioinformatics on the molecular data. K.H., P.F., A.J.J., T.F.N., H.S., K.U., J.E., P.O., A.R. and E.N. coordinated the collection of phenotypic data. T.A., N.J.B., M.B., G.H.B., S.K. and C.J. analysed the data. N.J.B., T.A. and C.R.P. wrote the manuscript. All authors read and commented on the manuscript. C.R.P. and S.L. contributed equally as senior authors.
Corresponding authors
Extended data figures and tables
Extended Data Figure 1 Map of study populations.
Bars indicate the proportion of individuals maturing after 1 (light blue), 2 (medium blue) or ≥3 years (dark blue) at sea; 1–54, NOR data set; 55–56, TAN; 57, BAL (Extended Data Table 1). Data for lake and river coordinates were obtained from European Environmental Agency (under a Creative Commons Attribution 4 License) and the Norwegian Water Resources and Energy Directorate.
Extended Data Figure 2 GWAS analyses for the TAN (n = 463), NOR (n = 941) and combined (n = 1,404) data sets.
a, Manhattan and quantile–quantile plots of the GWAS for age at maturity in Atlantic salmon before (left) and after (right) correction for population structure. The first three rows are models including phenotypic covariates (that is, the FULL model), and the next three rows are models without phenotypic covariates (that is, the BASIC model). The y axis shows the association statistic (−log10(P values)) for each SNP ordered by chromosome and position (x axis). The genome-wide statistical significance adjusted for multiple comparisons and genomic inflation is indicated by a horizontal dashed line. The VGLL3TOP (the SNP with the highest association with age at maturity) and VGLL3TAG (the SNP strongest linkage disequilibrium with the missense mutations in the VGLL3 gene) SNPs are shown with red arrows. QQ plots showing the deviation of P values (red line) from the null expectation (black line) are in the insets. b, Proportion of SNPs showing no evidence of significant population structure (Hnull: Akaike information criterion<−2) as a function of the number of principal components included in the model, for TAN (squares), NOR (circles) and the combined data set (TAN + NOR; triangles). The numbers of principal components used in population corrected models are marked with red. c, Relationship between population average age at maturity and allele frequency at the VGLL3TOP SNP and (d) SIX6TOP SNP. e, Relationship between the VGLL3TOP SNP and the SIX6TOP SNP allele frequencies.
Extended Data Figure 3 GWAS analyses for the BAL data set.
Manhattan plots and quantile–quantile plots of the GWAS for age at maturity in the BAL data set (n = 114), (a) before and (b) after correction for population structure. The y axis shows the association statistic (−log10(P values)) for each SNP ordered by chromosome and position (x axis). The genome-wide statistical significance adjusted for multiple comparisons and genomic inflation is indicated by a horizontal dashed line. The VGLL3TOP and VGLL3TAG SNPs are shown with red arrows. The QQ plot shows the deviation of P values (red line) from the null expectation (black line). c, Distribution of association statistics for the VGLL3TOP SNP in 100,000 bootstrapped replicates with resampling, using the TAN + NOR data set combined (n = 1,404). An equivalent sampling design to the BAL data set (n = 114 and the same age at maturity structure; see Supplementary Table 1) was used in the resampling. The red arrow indicates the P value of the VGLL3TOP SNP in the BAL data set.
Extended Data Figure 4 Gene model diagrams detailing regions around the VGLL3TOP and SIX6TOP loci.
a, Gene models and genomic positions of the two genes in the genome region on chromosome 25 significantly associated with age at maturity. Missense SNPs identified by re-sequencing within the genes are indicated in green. Amino acids indicated above and below the gene model were associated with the late (L) and early (E) maturation alleles, respectively. Longer tick marks show custom 220K Affymetrix axiom array SNPs, and shorter tick marks indicate re-sequencing variants. Notable SNPs are colour coded with red (VGLL3TOP), blue (VGLL3iHS) and green (the SNP tagging missense mutations in VGLL3 and the AKAP11 missense SNP). Note that missense variants on VGLL3 were identified by whole genome sequencing. The array SNP in tightest linkage disequilibrium with the VGLL3 missense variants identified by re-sequencing is 306 and 2,356 base pairs upstream (R2 = 1 and 0.71, respectively). b, Gene model and linkage disequilibrium plots of an ~0.5 Mb region on chromosome 9 where a significant GWAS signal was observed before correction for population structure. The association plot shown is before correction for population structure, using the combined data set (TAN + NOR). The SIX6TOP locus is shown in red. Shorter tick marks in the SNP axis indicate re-sequencing variants. FST estimates for SNPs in the region are also shown (lower graph). Closed circles indicate SNPs significantly diverged from null (neutral) expectations (FLK FST outlier test, 99.5% quantile of the null distribution, (56 populations, total n = 1,404). c, Conserved elements (PhastCons) of the 200 kb region around the SIX6 gene showing the predicted forebrain distal regulatory element (red tick mark) that is located close to the SIX6TOP SNP. One re-sequenced variant in strong linkage disequilibrium with the SIX6TOP SNP was located in this region.
Extended Data Figure 5 Details of modelling the genetic architecture of age at maturity.
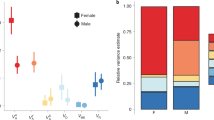
a, Threshold logistic models explaining variation in age at maturity in relation to the VGLL3TOP SNP in the TAN (n = 220 females, 243 males), NOR (n = 473 females, 468 males) and the combined (n = 693 females, 711 males) data sets for females (left panels) and males (right panels). Shaded grey areas around the logistic curves indicate one standard error of the threshold coefficients, and shaded red and blue areas indicate one standard error around genotype coefficients for females and males, respectively. The y axis depicts the probability of delaying maturation from one maturity age class to the next. LL genotypes were centred to zero (intercept) and had no standard error because of the rank deficiency of the model (that is, threshold degrees of freedom is prioritized in the model). Threshold coefficients are sex independent, which was the optimal model explaining the data (see Extended Data Table 2 and Supplementary information 3). Small insets to the right of each logistic curve depict the odds of delaying maturation for the LL genotype in relation to the EE genotype (median, 50% parametric sampling quantile) and the degree of partial dominance (median, 50% parametric sampling quantile) on the unobserved liability scale (that is, the x axis in the logistic curves). The dominance estimates (δ) given above each panel are scaled to [−1,1] range (δ = (2βEL + (βLL − βEE))/(|βLL − βEE|)), where negative and positive values indicate an EE-like, and LL-like, expression of the phenotype (that is, delayed maturation), respectively. P values in the upper insets show the significance of the model deviating from additivity (Padd, 10,000 parametric permutations). The difference in dominance between females and males is highly significant for all data sets (P = 0.0082 for TAN, and P < 0.001 for NOR and the combined data sets.). P values for all odds of delaying maturation are significant (P < 0.001, 100,000 parametric permutations). b, Predicted mean and 50% sampling quantiles (10,000 parametric permutations) of age at maturity using the logit transformation model. The y axis is log scaled. Padd values in the insets shows significance of the model deviating from additivity (10,000 parametric permutations).
Extended Data Figure 6 Haplotype length analysis summary.
a, Manhattan plot of each SNP in the study showing the P values of the correlation between population iHS values (46 populations, 32 haplotypes per population) and the average age at maturity. Ten SNPs flanking the VGLL3TOP and SIX6TOP SNPs are marked with red circles and triangles, respectively. b, c, Same as a but showing a 5 Mb magnified view of the (b) VGLL3 and (c) SIX6 regions. d, Histogram showing the statistic distribution of the association between iHS and average age at maturity for all SNPs analysed in the study. Ten SNPs around the VGLL3TOP and SIX6TOP SNPs are marked with blue and red arrows, respectively, where longer arrow tails show the VGLL3TOP and SIX6TOP SNPs. e, f, iHS concordance (Pearson’s r) in the TAN data set between the reduced (n = 16) and full data sets for (e) a sub-population (55) with lower average age at maturity (n = 137) and (f) a sub-population (56) with higher average age at maturity (n = 326). Each point shows a single SNP. The lower panel shows the concordance (Pearson’s r) of the TAN full data sets to all populations (n = 46) included in the iHS analysis. The self-concordance, as in the upper panel, is indicated with red. g, Relationship between population iHS score and VGLL3TOP allele frequency. iHS = 0 (no haplotype length difference) is marked with a horizontal grey line. Positive iHS values indicate longer haplotype blocks, and therefore stronger selection, around the E allele in a population relative to the L allele, and vice versa for negative iHS values.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References and Supplementary Tables 1-5. (PDF 968 kb)
Rights and permissions
About this article
Cite this article
Barson, N., Aykanat, T., Hindar, K. et al. Sex-dependent dominance at a single locus maintains variation in age at maturity in salmon. Nature 528, 405–408 (2015). https://doi.org/10.1038/nature16062
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16062
This article is cited by
-
The roles of sexual selection and sexual conflict in shaping patterns of genome and transcriptome variation
Nature Ecology & Evolution (2023)
-
Tests for associations between sexual dimorphism and patterns of quantitative genetic variation in the water strider, Aquarius remigis
Heredity (2023)
-
Preadapted to adapt: underpinnings of adaptive plasticity revealed by the downy brome genome
Communications Biology (2023)
-
Gene-poor Y-chromosomes substantially impact male trait heritabilities and may help shape sexually dimorphic evolution
Heredity (2023)
-
Investigation of VGLL3 and sub-target genes in the aetiology of paediatric acute appendicitis: a prospective case–control study
Pediatric Surgery International (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.