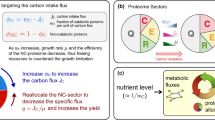
Abstract
Overflow metabolism refers to the seemingly wasteful strategy in which cells use fermentation instead of the more efficient respiration to generate energy, despite the availability of oxygen. Known as the Warburg effect in the context of cancer growth, this phenomenon occurs ubiquitously for fast-growing cells, including bacteria, fungi and mammalian cells, but its origin has remained unclear despite decades of research. Here we study metabolic overflow in Escherichia coli, and show that it is a global physiological response used to cope with changing proteomic demands of energy biogenesis and biomass synthesis under different growth conditions. A simple model of proteomic resource allocation can quantitatively account for all of the observed behaviours, and accurately predict responses to new perturbations. The key hypothesis of the model, that the proteome cost of energy biogenesis by respiration exceeds that by fermentation, is quantitatively confirmed by direct measurement of protein abundances via quantitative mass spectrometry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neidhardt, F. C., Ingraham, J. L. & Schaechter, M. Physiology of the Bacterial Cell: A Molecular Approach Ch. 5 (Sinauer Associates Inc, 1990)
Wolfe, A. J. The acetate switch. Microbiol. Mol. Biol. Rev. 69, 12–50 (2005)
De Deken, R. H. The Crabtree effect: a regulatory system in yeast. J. Gen. Microbiol. 44, 149–156 (1966)
De Mey, M., De Maeseneire, S., Soetaert, W. & Vandamme, E. Minimizing acetate formation in E. coli fermentations. J. Ind. Microbiol. Biotechnol. 34, 689–700 (2007)
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009)
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011)
Weinberg, R. A. The Biology of Cancer 2nd edn, Ch. 2.6 (Garland Science, 2013)
Majewski, R. A. & Domach, M. M. Simple constrained-optimization view of acetate overflow in E. coli. Biotechnol. Bioeng. 35, 732–738 (1990)
Varma, A. & Palsson, B. O. Stoichiometric flux balance models quantitatively predict growth and metabolic by-product secretion in wild-type Escherichia coli W3110. Appl. Environ. Microbiol. 60, 3724–3731 (1994)
Pfeiffer, T., Schuster, S. & Bonhoeffer, S. Cooperation and competition in the evolution of ATP-producing pathways. Science 292, 504–507 (2001)
Pfeiffer, T. & Bonhoeffer, S. Evolutionary consequences of tradeoffs between yield and rate of ATP production. Z. Phys. Chem. 216, 51–63 (2002)
Vazquez, A. et al. Impact of the solvent capacity constraint on E. coli metabolism. BMC Syst. Biol. 2, 7 (2008)
Vemuri, G. N., Altman, E., Sangurdekar, D. P., Khodursky, A. B. & Eiteman, M. A. Overflow metabolism in Escherichia coli during steady-state growth: transcriptional regulation and effect of the redox ratio. Appl. Environ. Microbiol. 72, 3653–3661 (2006)
Molenaar, D., van Berlo, R., de Ridder, D. & Teusink, B. Shifts in growth strategies reflect tradeoffs in cellular economics. Mol. Syst. Biol. 5, 323 (2009)
Valgepea, K. et al. Systems biology approach reveals that overflow metabolism of acetate in Escherichia coli is triggered by carbon catabolite repression of acetyl-CoA synthetase. BMC Syst. Biol. 4, 166 (2010)
Renilla, S. et al. Acetate scavenging activity in Escherichia coli: interplay of acetyl-CoA synthetase and the PEP-glyoxylate cycle in chemostat cultures. Appl. Microbiol. Biotechnol. 93, 2109–2124 (2012)
Zhuang, K., Vemuri, G. N. & Mahadevan, R. Economics of membrane occupancy and respiro-fermentation. Mol. Syst. Biol. 7, 500 (2011)
Huberts, D. H., Niebel, B. & Heinemann, M. A flux-sensing mechanism could regulate the switch between respiration and fermentation. FEMS Yeast Res. 12, 118–128 (2012)
O’Brien, E. J., Lerman, J. A., Chang, R. L., Hyduke, D. R. & Palsson, B. O. Genome-scale models of metabolism and gene expression extend and refine growth phenotype prediction. Mol. Syst. Biol. 9, 693 (2013)
Peebo, K. et al. Proteome reallocation in Escherichia coli with increasing specific growth rate. Mol. Biosyst. 11, 1184–1193 (2015)
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z. & Hwa, T. Interdependence of cell growth and gene expression: origins and consequences. Science 330, 1099–1102 (2010)
Meyer, H.-P., Leist, C. & Fiechter, A. Acetate formation in continuous culture of Escherichia coli K12 D1 on defined and complex media. J. Biotechnol. 1, 355–358 (1984)
Nanchen, A., Schicker, A. & Sauer, U. Nonlinear dependency of intracellular fluxes on growth rate in miniaturized continuous cultures of Escherichia coli. Appl. Environ. Microbiol. 72, 1164–1172 (2006)
el-Mansi, E. M. & Holms, W. H. Control of carbon flux to acetate excretion during growth of Escherichia coli in batch and continuous cultures. J. Gen. Microbiol. 135, 2875–2883 (1989)
Applebee, M. K., Joyce, A. R., Conrad, T. M., Pettigrew, D. W. & Palsson, B. O. Functional and metabolic effects of adaptive glycerol kinase (GLPK) mutants in Escherichia coli. J. Biol. Chem. 286, 23150–23159 (2011)
Scott, M. & Hwa, T. Bacterial growth laws and their applications. Curr. Opin. Biotechnol. 22, 559–565 (2011)
You, C. et al. Coordination of bacterial proteome with metabolism by cyclic AMP signalling. Nature 500, 301–306 (2013)
Hui, S. et al. Quantitative proteomic analysis reveals a simple strategy of global resource allocation in bacteria. Mol. Syst. Biol. 11, 784 (2015)
Maaloe, O. in Biological Regulation and Development Vol. 1 (ed. Goldberger, R. F. ) 487–542 (Plenum, 1979)
Nelson, D. L., Lehninger, A. L. & Cox, M. M. Lehninger Principles of Biochemistry Chs 14, 16 (Macmillan, 2008)
Sandén, A. M. et al. Limiting factors in Escherichia coli fed-batch production of recombinant proteins. Biotechnol. Bioeng. 81, 158–166 (2003)
Neijssel, O. M., Teixeira de Mattos, M. J. & Tempest, D. W. in Escherichia coli and Salmonella: Cellular and Molecular Biology (eds Neidhardt, F. C. et al. ) 1683–1693 (ASM Press, 1996)
Brooker, R. J. An analysis of lactose permease “sugar specificity” mutations which also affect the coupling between proton and lactose transport. I. Val177 and Val177/Asn319 permeases facilitate proton uniport and sugar uniport. J. Biol. Chem. 266, 4131–4138 (1991)
Li, G. W., Burkhardt, D., Gross, C. & Weissman, J. S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157, 624–635 (2014)
Flamholz, A., Noor, E., Bar-Even, A., Liebermeister, W. & Milo, R. Glycolytic strategy as a tradeoff between energy yield and protein cost. Proc. Natl Acad. Sci. USA 110, 10039–10044 (2013)
Woldringh, C. L., Binnerts, J. S. & Mans, A. Variation in Escherichia coli buoyant density measured in Percoll gradients. J. Bacteriol. 148, 58–63 (1981)
Bollenbach, T., Quan, S., Chait, R. & Kishony, R. Nonoptimal microbial response to antibiotics underlies suppressive drug interactions. Cell 139, 707–718 (2009)
El-Mansi, M. Flux to acetate and lactate excretions in industrial fermentations: physiological and biochemical implications. J. Ind. Microbiol. Biotechnol. 31, 295–300 (2004)
Farmer, W. R. & Liao, J. C. Improving lycopene production in Escherichia coli by engineering metabolic control. Nature Biotechnol. 18, 533–537 (2000)
Aristidou, A. A., San, K. Y. & Bennett, G. N. Metabolic engineering of Escherichia coli to enhance recombinant protein production through acetate reduction. Biotechnol. Prog. 11, 475–478 (1995)
Veit, A., Polen, T. & Wendisch, V. F. Global gene expression analysis of glucose overflow metabolism in Escherichia coli and reduction of aerobic acetate formation. Appl. Microbiol. Biotechnol. 74, 406–421 (2007)
Galluzzi, L., Kepp, O., Vander Heiden, M. G. & Kroemer, G. Metabolic targets for cancer therapy. Nature Rev. Drug Discov. 12, 829–846 (2013)
Kuhlman, T., Zhang, Z., Saier, M. H., Jr & Hwa, T. Combinatorial transcriptional control of the lactose operon of Escherichia coli. Proc. Natl Acad. Sci. USA 104, 6043–6048 (2007)
Levine, E., Zhang, Z., Kuhlman, T. & Hwa, T. Quantitative characteristics of gene regulation by small RNA. PLoS Biol. 5, e229 (2007)
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Sys. Biol. 2, 2006.0008 (2006)
Cayley, S., Record, M. T., Jr & Lewis, B. A. Accumulation of 3-(N-morpholino)propanesulfonate by osmotically stressed Escherichia coli K-12. J. Bacteriol. 171, 3597–3602 (1989)
Neidhardt, F. C., Bloch, P. L. & Smith, D. F. Culture medium for enterobacteria. J. Bacteriol. 119, 736–747 (1974)
Sperling, E., Bunner, A. E., Sykes, M. T. & Williamson, J. R. Quantitative analysis of isotope distributions in proteomic mass spectrometry using least-squares Fourier transform convolution. Anal. Chem. 80, 4906–4917 (2008)
Rockwood, A. L. & Van Orden, S. L. Ultrahigh-speed calculation of isotope distributions. Anal. Chem. 68, 2027–2030 (1996)
Lyons, E., Freeling, M., Kustu, S. & Inwood, W. Using genomic sequencing for classical genetics in E. coli K12. PLoS ONE 6, e16717 (2011)
Soupene, E. et al. Physiological studies of Escherichia coli strain MG1655: growth defects and apparent cross-regulation of gene expression. J. Bacteriol. 185, 5611–5626 (2003)
Brown, S. D. & Jun, S. Complete genome sequence of Escherichia coli NCM3722. Genome Announc. 3, e00879–15 (2015)
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997)
Klumpp, S., Zhang, Z. & Hwa, T. Growth rate-dependent global effects on gene expression in bacteria. Cell 139, 1366–1375 (2009)
Keseler, I. M. et al. EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res. 41, D605–D612 (2013)
Unden, G., Steinmetz, P. A. & Degreif-Dünnwald, P. The Aerobic and anaerobic respiratory chain of Escherichia coli and Salmonella enterica: enzymes and energetics. EcoSal Plus http://dx.doi.org/10.1128/ecosalplus.ESP-0005-2013 (2014)
Holms, H. Flux analysis and control of the central metabolic pathways in Escherichia coli. FEMS Microbiol. Rev. 19, 85–116 (1996)
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genet. 25, 25–29 (2000)
Macnab, R. in Escherichia coli and Salmonella: Cellular and Molecular Biology (eds Neidhardt, F. C. et al. ) 123–145 (ASM Press, 1996)
McLaughlin, S. The mechanism of action of DNP on phospholipid bilayer membranes. J. Membr. Biol. 9, 361–372 (1972)
Rhoads, D. B., Waters, F. B. & Epstein, W. Cation transport in Escherichia coli. VIII. Potassium transport mutants. J. Gen. Physiol. 67, 325–341 (1976)
Kochanowski, K. et al. Functioning of a metabolic flux sensor in Escherichia coli. Proc. Natl Acad. Sci. USA 110, 1130–1135 (2013)
Chubukov, V., Gerosa, L., Kochanowski, K. & Sauer, U. Coordination of microbial metabolism. Nature Rev. Microbiol. 12, 327–340 (2014)
Acknowledgements
We are grateful to F. J. Bruggeman, E. O’Brien, U. Sauer, M. H. Saier and members of the Hwa and Sauer laboratories for valuable comments, and J. L. Figueroa for artistic contributions to the model illustration in Box 1. This work was supported by the NIH (grant R01-GM109069) and the Simons Foundation (grant 330378). T.H. additionally acknowledges the support of M. Rössler, the Walter Haefner Foundation and the ETH Foundation. M.B. acknowledges support from SystemsX TPdF. Y.S. acknowledges support from Hong Kong Baptist University (grants FRG2/11-12/159 and SKLP-14-15-P012).
Author information
Authors and Affiliations
Contributions
M.B., S.H., J.R.W. and T.H. designed the study. M.B., S.H., H.O., Z.Z. and Y.S. performed experiments. M.B., S.H. and T.H. analysed the data and developed the model. M.B., S.H., J.R.W. and T.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Acetate excretion data.
a–f, The acetate excretion data is shown for E. coli cells grown in chemostat (a–e) and for cells growing on medium with non-glycolytic carbon sources (f). a, Glucose-limited chemostat data based on figure 1 from ref. 22. b, Glucose-limited and pyruvate–limited chemostat data from table 7 of ref. 57. c, Glucose-limited chemostat data based on figure 3 of ref. 23. Only data with dilution rates less than the apparent washout dilution rate are plotted here. d, Glucose-limited chemostat data from table 1 of ref. 15. e, Glucose-limited chemostat data based on figure 1 of ref. 16. f, E. coli K-12 NCM3722 was grown in minimal medium with one of five non-glycolytic carbon sources, including two gluconeogenic substrates (pyruvate and lactate), one substrate of the pentose phosphate pathway (gluconate), and two intermediates of the TCA pathway (succinate and fumarate). Deviation from the acetate line (the red line, as defined in Fig. 1 and equation (1) of the main text) is seen most notably for pyruvate, which excretes a very large amount of acetate, and to a lesser degree, also lactate and gluconate, which enter glycolysis as pyruvate. In the framework of our model, these deviations result from different proteome efficiencies of fermentation and respiration on these carbon sources. Note: acetate excretion measurement was also attempted for growth on LB. However, growth on LB is not characterized by a single exponential steady-state growth phase, as various constituents of the medium are depleted during the course of batch culture growth. Assuming exponential growth for A600 nm data below 0.3 and alternatively from 0.3 for 0.5 gave doubling time of 18 min and 28 min, respectively. The corresponding acetate excretion rates were 14.3 and 3.6 mM A600 nm−1 h−1. These data should be regarded as semi-quantitative owing to the non-steady nature of growth on LB.
Extended Data Figure 2 The (oxidative) fermentation and respiration pathways.
a, Schematic illustration of the fermentation pathway, using glucose as an exemplary carbon source. The pathway is shown as the coloured part. One molecule of glucose is catabolized into two molecules of acetate and two molecules of CO2 (not shown in the diagram), with four molecules of ATP generated via substrate phosphorylation and also four molecules of NADH produced. In the aerobic environment, NADH molecules can be converted into ATP molecules. The total number of ATP molecules produced per glucose molecule is therefore 4 + 4x, in which the conversion factor x indicates the number of ATP molecules converted from one NADH molecule (that is, ATP:NADH = x:1). In the illustration, we assume that two molecules of ATP are converted from one NADH molecule; that is, x = 2. b, Schematic illustration of the respiration pathway, using glucose as an exemplary carbon source. The pathway is shown as the coloured part. One molecule of glucose is catabolized into 6 molecules of CO2 (not shown in the diagram), producing 4 molecules of ATP, 6 molecules of NADH, 2 molecules of NADPH, and 2 molecules of FADH2. Using ATP:NADH = x:1, ATP:NADPH = x:1 and ATP:FADH2 = x:2, we have the total number of ATP molecules produced as 4 + 9x for the respiration pathway. Here in the illustration, we assumed x = 2. Note that the ratio of total ATP produced from respiration over total ATP produced from fermentation depends on the conversion factor x; that is, (4 + 9x)/(4 + 4x). The value of this ratio ranges from 1 (for x = 0) to 9/4 (for x → ∞).
Extended Data Figure 3 Growth-rate dependence of acetate production and CO2 evolution in bioreactor: data and comparisons to the model.
a, The rate of CO2 evolution was determined in a bioreactor setup for wild-type and titratable LacY cells (NCM3722 and NQ381, respectively) grown in lactose minimum medium with various degrees of lactose uptake titration, and the result was used to deduce the CO2 flux produced by respiration (blue circles). Also plotted (red squares) is the acetate excretion rate measured in the bioreactor. See Supplementary Note 2 for details of this experiment and corresponding analysis. The inducer levels, growth rate, measurements of glucose, acetate and CO2, and the deduced CO2 levels via respiration are shown in the table on the right. b, Deduced energy production fluxes from fermentation and respiration pathways, based on the measurements presented in a. Fluxes are in units of mM A600 nm−1 h−1. c, Comparison of model and experimental data. Using the set of parameters summarized in Extended Data Table 2, the model solution (equations (S14) and (S17)) satisfactorily describes the experimental data obtained for acetate excretion (Jac) and respiratory CO2 production  in the bioreactor for carbon limitation. These results depend on the assumed ratios of ATP-carbon conversion. As described in Supplementary Note D1, the ratios we used in this work are ATP:NADH = 2:1, ATP:NADPH = 2:1 and ATP:FADH2 = 1.15:1. Note that these conversion ratios have never been precisely measured and could be substantially overestimated15. However, the central results presented in this work are robust with respect to the choice of these conversion ratios. As an illustration, we show in d that the model results generated with a very different set of conversion ratios (ATP:NADH = 0.5:1, ATP:NADPH = 0.5:1 and ATP:FADH2 = 0.5:1) even provide a slightly better description of the data. (For these conversion ratios, the energy production of the cell matches the theoretical energy demand for biomass production.) The full model calibration requires the rate of CO2 evolution, which can only be measured in a bioreactor setup. We note a small discrepancy between acetate fluxes and growth rates obtained for cultures grown in bioreactor as compared to batch cultures, possibly caused by differences in aeration.
in the bioreactor for carbon limitation. These results depend on the assumed ratios of ATP-carbon conversion. As described in Supplementary Note D1, the ratios we used in this work are ATP:NADH = 2:1, ATP:NADPH = 2:1 and ATP:FADH2 = 1.15:1. Note that these conversion ratios have never been precisely measured and could be substantially overestimated15. However, the central results presented in this work are robust with respect to the choice of these conversion ratios. As an illustration, we show in d that the model results generated with a very different set of conversion ratios (ATP:NADH = 0.5:1, ATP:NADPH = 0.5:1 and ATP:FADH2 = 0.5:1) even provide a slightly better description of the data. (For these conversion ratios, the energy production of the cell matches the theoretical energy demand for biomass production.) The full model calibration requires the rate of CO2 evolution, which can only be measured in a bioreactor setup. We note a small discrepancy between acetate fluxes and growth rates obtained for cultures grown in bioreactor as compared to batch cultures, possibly caused by differences in aeration.
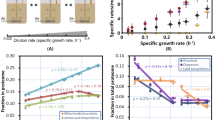
Extended Data Figure 4 The effect of useless protein expression on acetate excretion.
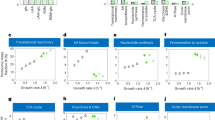
a, Acetate excretion rate by strain NQ1389 is plotted against the absolute abundance of the expressed LacZ proteins, reported as a fraction of total protein (ϕZ), for each of the four carbon sources described in Fig. 2a. The solid lines, depicting the linear decrease in acetate excretion, are model predictions (equation (S23) in Supplementary Information), with the lone parameter ϕmax ≈ 47% (the x-intercept of the line) determined from the least-mean-squares fit of the data in 3D plot of Fig. 2b, by a plane anchored to the acetate line. b, Alternatively, ϕmax can be determined from linear fits of growth rate versus ϕZ for the four carbon sources shown. This results in ϕmax = 42% ± 5%. The solid lines in this panel are linear fits using ϕmax = 42%. (We note that over a broad growth rate range, ϕmax actually exhibits a growth-rate dependence (Dai, X. et al.,manuscript in preparation). Nevertheless, over the narrow growth-rate range relevant for acetate excretion, this dependence is negligible. Hence, for the purposes of our paper, we consider ϕmax to be constant.) c, A different view of the 3D plot in Fig. 2b. d, Glucose uptake rate as a function of growth rate under LacZ overexpression. The circles are the data and the dashed line is the best-fit to the data passing through the origin. e, The relative protein levels of several representative genes, taken from amino acid synthesis (red), central metabolism (blue), protein synthesis genes (green), and nucleotide synthesis (black). As described in ref. 28, the vast majority of genes exhibited an expression pattern that is linearly proportional to the growth rate when growth is changed by increasing LacZ expression. f, Growth-rate dependence of motility proteins under carbon limitation. The proteome fraction data (green symbols) is from the carbon limitation series in ref. 28, in which growth rate was limited by titrating the lactose uptake for the strain NQ381. The motility proteins are proteins that are associated with the Gene Ontology (GO) term 0006810 (with GO name ‘locomotion’) as defined by the Gene Ontology Consortium58. See ref. 28 for detailed description of the experimental procedure and data processing. Note that the fraction of motility proteins increases the most in the growth range where acetate is excreted. Also note that the energy consumption by chemotaxis comprises a very minor fraction of the total energy budget, estimated to be in the order of 0.1% (ref. 59). Disabling the motility function therefore does not affect the cell’s energy requirement.
Extended Data Figure 5 Acetate excretion due to energy dissipation by DNP.
DNP is a chemical known to dissipate membrane potential60 and thus imposes energy stress on the cell. Acetate excretion rates for the titratable glucose uptake strain (NQ1243) grown in medium with glucose and different concentrations of DNP were measured for different degrees of glucose uptake. The results are qualitatively similar to the data for the leaky LacY mutant NQ1313 (Fig. 3d), except for a systematic difference in the slopes of the resulting acetate lines (thin lines of different colours). The origin of this deviation is presumably a more complex action of DNP with additional effects on the cell as compared to the leaky LacY mutant. Indeed, it is known for instance that in addition to leakage of protons, DNP also causes leakage of osmolites through the membrane61.
Extended Data Figure 6 The relative expression levels of glycolysis proteins under proteome perturbation (by LacZ overexpression), and energy dissipation (by expressing leaky LacY).
Orange data points and linear fits result from the overexpression experiment (that is, NQ1389 grown in glucose minimal medium with different induction levels of LacZ expression), and blue data points and fits are the leaky LacY series (that is, the wild-type LacY strain NQ1312 and the leaky LacY strain NQ1313 grown in glucose minimal medium with 200 μM 3-methylbenzyl alcohol (3MBA)). The y axis denotes relative protein levels, which were obtained by mass spectrometry with the same reference for the two series (see Methods). The x axis is the growth rate (in units of h−1). The different trends of protein expression for the two series show the distinct nature of the two perturbations, demonstrating that these seemingly similar predictions for the acetate line (parallel shift to slow growth as shown in Fig. 3a, d) for the two perturbations, have distinct origins and exhibit distinctly different patterns of gene expression in accordance with model predictions derived in section C3 of Supplementary Note 1 (see equations (S26) and (S36)). From the perspective of gene regulation, it is not obvious what causes the increased expression levels of glycolysis genes under energy dissipation; the transcription factor Cra in combination with the key central carbon intermediate fructose-1,6-bisphosphate (FBP) is recognized as the major regulator of glycolysis62. FBP relieves the repression of glycolytic enzymes expression by Cra63. The observed increase in the abundance of glycolytic enzymes under energy dissipation could be caused by a build-up of FBP, as energy stress limits protein polymerization. However, in this case, it is not clear what signalling pathway gives rise to the opposite responses of glycolytic enzymes to LacZ overexpression. Inset, corresponding glucose uptake and acetate excretion rates for the two perturbations presented in the main figure. Glucose uptake and acetate excretion rates decreased proportional to growth rate for LacZ overexpression (as expected from the model equations (S25), (S26) and (S29)). On the other hand, there was a marked increase in acetate excretion with energy dissipation for a roughly constant glucose uptake rate, as correctly anticipated by the model (equation (S36)). Note that the protein abundance of ACS (main panel) shows that this increase in the acetate excretion rate was not caused by a drop in ACS. Instead, the observed increase of acetate excretion, together with the parallel increase in the expression level of glycolysis and TCA enzymes, points to the coordination of glycolytic and TCA fluxes in response to energy demand.
Extended Data Figure 7 The relative expression level of TCA proteins under proteome perturbations (by protein overexpression) and energy dissipation (by using leaky LacY).
Orange data points and linear fits represent the overexpression experiment (that is, NQ1389 with different induction levels of LacZ expression), and blue data points and fits are the leaky LacY series (using strains NQ1312 and NQ1313 with wild-type and leaky LacY expression, respectively). The y axis denotes relative protein levels, which were obtained by mass spectrometry with the same reference for the two series (see Methods). The x axis is growth rate (in units of h−1). The different trends of protein expression for the two series show the distinct nature of the two perturbations, demonstrating that these seemingly similar predictions on the acetate line (parallel shift to slow growth as shown in Fig. 3a, c) for the two perturbations, have distinct origins and exhibit distinctly different patterns of gene expression in accordance with model predictions derived in section C3 of Supplementary Note 1 (see equations (S27) and (S37)). From the perspective of gene regulation, the transcription factor CRP is thought to be the major regulator of TCA enzyme expression in aerobic conditions63. CRP-cAMP activity, which increases under carbon limitation, is known to activate the expression of most TCA enzymes. Together with the known role of CRP in upregulating the enzyme ACS which takes up acetate15,16, CRP is considered to be a major candidate for regulating energy metabolism and acetate excretion. However, our findings that the expression of TCA enzymes increased under energy dissipation while acetate excretion also increased (Extended Data Fig. 6, inset) cannot be accounted for by known mechanisms of CRP regulation, and instead suggest an important role of additional regulators in the coordination of energy biogenesis pathways.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-4, including Supplementary Figures 1-15, Supplementary Tables 1-6 and Supplementary References. (PDF 2951 kb)
Rights and permissions
About this article
Cite this article
Basan, M., Hui, S., Okano, H. et al. Overflow metabolism in Escherichia coli results from efficient proteome allocation. Nature 528, 99–104 (2015). https://doi.org/10.1038/nature15765
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15765
This article is cited by
-
A parallel glycolysis provides a selective advantage through rapid growth acceleration
Nature Chemical Biology (2024)
-
A universal metabolite repair enzyme removes a strong inhibitor of the TCA cycle
Nature Communications (2024)
-
Volatilomes of human infection
Analytical and Bioanalytical Chemistry (2024)
-
Growth-coupled anaerobic production of isobutanol from glucose in minimal medium with Escherichia coli
Biotechnology for Biofuels and Bioproducts (2023)
-
Inter-bacterial mutualism promoted by public goods in a system characterized by deterministic temperature variation
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.