Abstract
Large cancer cell line collections broadly capture the genomic diversity of human cancers and provide valuable insight into anti-cancer drug response. Here we show substantial agreement and biological consilience between drug sensitivity measurements and their associated genomic predictors from two publicly available large-scale pharmacogenomics resources: The Cancer Cell Line Encyclopedia and the Genomics of Drug Sensitivity in Cancer databases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Sharma, S. V., Haber, D. A. & Settleman, J. Cell line-based platforms to evaluate the therapeutic efficacy of candidate anticancer agents. Nature Rev. Cancer 10, 241–253 (2010)
Neve, R. M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006)
Caponigro, G. & Sellers, W. R. Advances in the preclinical testing of cancer therapeutic hypotheses. Nature Rev. Drug Discov . 10, 179–187 (2011)
Garraway, L. A. et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 436, 117–122 (2005)
Solit, D. B. et al. BRAF mutation predicts sensitivity to MEK inhibition. Nature 439, 358–362 (2006)
Sos, M. L. et al. Predicting drug susceptibility of non-small cell lung cancers based on genetic lesions. J. Clin. Invest. 119, 1727–1740 (2009)
Haibe-Kains, B. et al. Inconsistency in large pharmacogenomic studies. Nature 504, 389–393 (2013)
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012)
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012)
Acknowledgements
We thank T. Golub, E. Lander, S. Schreiber, P. Clemons and J. Engelman for helpful discussions. This work was supported by research grants from the Novartis Institutes for BioMedical Research (CCLE; L.A.G., M.G., and G.V.K.) and by grants from the Wellcome Trust (086357 and 102696; D.A.H., M.R.S., U.M., M.J.G., A.A., C.H.B.) and the National Institutes of Health (1U54HG006097-01, A.A. and C.H.B.). L.A.G. was supported in part by grants from Novartis and the Dr. Miriam and Sheldon Adelson Medical Research Foundation. G.V.K. was supported in part by the Slim Foundation. F.I. was supported in part by the EMBL-EBI and Wellcome Trust Sanger Institute Post-Doctoral (ESPOD) programme, and U.M. was funded by a Cancer Research UK Clinician Scientist Fellowship (A16629).
Author information
Authors and Affiliations
Consortia
Contributions
N.S., L.A.G., A.A., D.A.H., C.H.B., S.R., J.L., J.B., G.C., R.S., W.R.S., F.S., M.P.M., F.I., M.M., J.S.-R., M.R.S., U.M. and M.J.G. conceived the studies; N.S., M.G., G.V.K., A.A., I.P.-M., J.L., M.L., D.S., A.K., K.V., E.J.E., M.P.M., F.I. and M.M. performed analyses; N.S., M.G., A.A., M.L., M.R., F.I. and M.M. wrote/tested the R code, and N.S., M.G., A.A., L.A.G., C.H.B., J.L., M.P.M. and J.S.-R. wrote the paper. The Cancer Cell Line Encyclopedia investigators are N.S., M.G., G.V.K., L.A.G., J.L., M.L., D.S., A.K., K.V., E.J.E., M.R., J.B., G.C., R.S., W.R.S., F.S. and M.M.; the Genomics of Drug Sensitivity in Cancer investigators are A.A., I.P.-M., D.A.H., S.R., C.H.B., F.I., M.R.S., U.M. and M.J.G.
Corresponding authors
Ethics declarations
Competing interests
N.S. is an employee and shareholder of Blueprint Medicines. L.A.G. is a consultant for Foundation Medicine, Novartis, Boehringer Ingelheim, an equity holder in Foundation Medicine, and a member of the Scientific Advisory Board at Warp Drive. L.A.G. receives sponsored research support from Novartis. J.L., M.L., A.K., K.V., J.B., G.C., R.S., W.R.S, F.S., M.P.M. are employees and shareholders of Novartis. U.M. is a founder and consultant for 14M Genomics Ltd. M.R.S. is a founder and shareholder of 14M Genomics Ltd.
Extended data figures and tables
Extended Data Figure 1 Comparison of pharmacological data from the CCLE and GDSC studies.
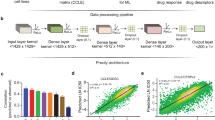
a, b, Scatter plots (blue dots) represent the drug sensitivity measured as the area under the dose–response curve (a) and IC50 (b) in overlapping cell lines between CCLE and GDSC studies. For this analysis, IC50 values for insensitive compounds were set to the highest concentration tested in both data sets. The number of overlapping cell lines n for each drug is indicated, as well as the Pearson correlation coefficient R and P value. In this representation, lower values denote insensitive cell lines. The full distribution of sensitivity values for each drug and study is depicted as ‘violin plots’ (green, CCLE; purple, GDSC) and accounts for all tested cell lines, as opposed to the overlapping set; the grey dot represents the median, thick black line represents the first to third quartile range, and shape of the plot represents the kernel density of the distribution.
Extended Data Figure 2 Power analysis of Spearman and Pearson correlation tests.
a, Example of a clear signal that appears in only 2% (20 out of 1,000) data points using synthetic data. The Spearman statistic completely fails to detect such a signal which is typical for selective cancer therapeutics. b, c, Expected Spearman and Pearson correlation coefficients between the two data sets assuming different percentages of drug-sensitive cell lines (α = 2%, 5%, 10% and 50%) and different number of overlapping cell lines. The error bars depict ± one standard deviation. d, e, Estimated statistical power for Spearman and Pearson correlation tests using a P value cutoff of 0.05 for rejecting the null hypothesis. This analysis was done using synthetic data as described in the Methods.
Extended Data Figure 3 Waterfall analysis for categorization of cell lines.
a, Schematic of the waterfall analysis methodology and example of outcome for PLX4720. b, Consistency in cell line sensitivity categorization for all drugs. The waterfall method using all data available was used to determine thresholds between ‘sensitive’ and ‘resistant’ cell lines (blue). Alternatively a 1 μM threshold was used (green). Asterisks indicate significance of Cohen’s Kappa coefficients (P < 0.05).
Extended Data Figure 4 Overlap in ANOVA genomic correlates of drug sensitivity.
a–d, Volcano plots showing ANOVA outcomes using drug responses from CCLE (left, a, c) or GDSC (right, b, d) data set from overlapping set of cell lines, and mutational status of 71 cancer genes from the GDSC. a, b, Analyses using AUC values. c, d, Analyses using IC50 values. Points represent drug–gene interactions (with sizes proportional to the number of screened mutant cell lines). Positions on x axis indicate effect size magnitudes: negative values (green circle) indicate mutations associated with increase in sensitivity, positive values (red circle) mutations associated with increased resistance. Positions on y axis indicate association significances (corrected P values) and the horizontal dashed line indicates a significance threshold (FDR 20%). Corresponding drug name, target(s) and cancer gene are reported for a subset of therapeutically relevant interactions.
Extended Data Figure 5 Consistency of drug sensitivity/tissue-of-origin associations between the CCLE and GDSC data sets.
Each point is a tested association between drug response and a given cell line’s tissue of origin. Positions of the points on the two axes correspond to ‘signed log q-values’ of the corresponding tests for the two data sets, respectively. Point labels indicate drug names and targets (in italics) and tested tissue (among round brackets). The sign indicates the effect of the marker (neg = increased sensitivity and pos = increased resistance) and the magnitude indicates the log P value of the corresponding t-test, after correcting for multiple hypothesis testing. Fisher’s exact test P values for independence of columns and rows of the contingency table determined by sign and significance of the associations are also reported (over all the tests and for significant associations only, respectively).
Extended Data Figure 6 Comparison of genomic features selected by elastic net between the CCLE and GDSC data sets.
a, Consistency in predictors of response identified by elastic net regression across 21,013 genome features (copy number variations, messenger RNA expression and sequence variants). Statistical significance of the number of genomic features identified in common (χ2 test) using the GDSC and CCLE drug sensitivity data sets. Only drugs where features were found in both studies are represented. b, Corresponding contingency tables. Out of the 4,957 drug–gene associations with non-zero elastic net weight coefficients, only one divergent result was found (weight coefficient with opposite signs), corresponding to a feature with the lowest possible frequency (non-zero coefficient in 1 out of 100 bootstrap trials in the elastic net analysis).
Extended Data Figure 7 Comparison of genomic feature-drug associations in the CCLE and GDSC data sets.
a, b, Ridge regression coefficients for all the drugs with successful elastic net regression in the indicated data set are plotted using either overlapping (a) or all available (b) cell lines. To select cell line features, elastic net was performed using the indicated data set. Then, ridge regression was performed on each data set using the selected features. For plotting, the weights associated with the features were multiplied by the standard deviation of the features as in Garnett et al.9, and then standardized per drug. Colour scale indicates the number of times a feature is selected in 100 independent runs of the elastic net. Green and red colouring indicate features associated with sensitivity or resistance, respectively.
Extended Data Figure 8 Agreement in genomic predictors of drug response identified by elastic net regression in the GDSC and CCLE studies.
Elastic net selection of genomic features was performed on the indicated data set and their effects were computed using a non-selective regression (ridge). Total number of features selected by elastic net is reported above the bars. Number of cell lines used in the regression is in parentheses on the x axis. Consistency is reported as the proportion of features with the overall same direction of effect (association with sensitivity or resistance): proportion of features with same sign, using either the cosine correlation that takes into account the sign associated with the features or the Pearson’s correlation that does not.
Extended Data Figure 9 Gene expression correlates of drug response identified previously have better agreement when using more stringent FDR cut-offs.
Data from Haibe-Kains et al.7. a, Scatter plots of the IC50 based gene-drug association statistic (column “stat” in Haibe-Kains et al.7; Supplementary Data 2 and 3 and Extended Data Fig. 6) with FDR between 0 and 0.01 (purple), 0.01 and 0.05 (cyan), 0.05 and 0.2 (green). In each panel the two black lines intersect at the origin and define the agreement quadrants (top right and bottom left quadrants). b, Proportion of genes in the agreement quadrants (same sign between the two studies). c, Additional measures of agreement between the two studies: Agreement measures increase with more stringent FDR cut-off, suggesting that false discovery drives agreement down. Uncentred measures (cosine correlation, uncentred covariance, agreement quadrant proportion) yield better agreement between the studies (see Supplementary Discussion).
Extended Data Figure 10 Example of significant change in observed correlation by addition of a few sensitive cell lines.
For lapatinib sensitivity data, there are 86 overlapping cell lines between the CCLE and GDSC data sets. a, Left panel is an excerpt from Haibe-Kains et al.7 figure 2 comparing the sensitivity data of lapatinib for the two data sets. b, Right panel shows the two sensitive cell lines (BT-474 and NCI-H1648) that were omitted in the analysis of Haibe-Kains et al.7. The inclusion of these two cell lines drastically changes the observed Pearson correlation (from 0.25 to 0.53). This is consistent with the simulation results (Extended Data Fig. 2c) that show high variability in the observed Pearson correlation for low sample numbers.
Supplementary information
Supplementary Data 1
This file contains cell line collections and drug responses. (XLSX 870 kb)
Supplementary Data 2
This file contains Waterfall analysis. (XLSX 58 kb)
Supplementary Data 3
This file contains ANOVA results for gene-drug associations. (XLSX 86 kb)
Supplementary Data 4
This file contains t-test results for tissue-drug associations. (XLSX 39 kb)
Supplementary Data 5
This file contains Elastic Net results. (XLSX 1054 kb)
Supplementary Data 6
This file contains Elastic Net and Ridge regression results. (XLSX 2950 kb)
Supplementary Data 7
This file contains Drug/Genotype associations missed in one dataset. (XLSX 13 kb)
Supplementary Text
This file contains a Supplementary Discussion and additional references. (PDF 154 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
The Cancer Cell Line Encyclopedia Consortium., The Genomics of Drug Sensitivity in Cancer Consortium. Pharmacogenomic agreement between two cancer cell line data sets. Nature 528, 84–87 (2015). https://doi.org/10.1038/nature15736
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15736
This article is cited by
-
Prevalence, causes and impact of TP53-loss phenocopying events in human tumors
BMC Biology (2023)
-
Salvage of ribose from uridine or RNA supports glycolysis in nutrient-limited conditions
Nature Metabolism (2023)
-
Whole-genome sequencing reveals an association between small genomic deletions and an increased risk of developing Parkinson’s disease
Experimental & Molecular Medicine (2023)
-
Systematic analysis of transcriptome signature for improving outcomes in lung adenocarcinoma
Journal of Cancer Research and Clinical Oncology (2023)
-
StringFix: an annotation-guided transcriptome assembler improves the recovery of amino acid sequences from RNA-Seq reads
Genes & Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.