Abstract
The human kidney contains up to 2 million epithelial nephrons responsible for blood filtration. Regenerating the kidney requires the induction of the more than 20 distinct cell types required for excretion and the regulation of pH, and electrolyte and fluid balance. We have previously described the simultaneous induction of progenitors for both collecting duct and nephrons via the directed differentiation of human pluripotent stem cells1. Paradoxically, although both are of intermediate mesoderm in origin, collecting duct and nephrons have distinct temporospatial origins. Here we identify the developmental mechanism regulating the preferential induction of collecting duct versus kidney mesenchyme progenitors. Using this knowledge, we have generated kidney organoids that contain nephrons associated with a collecting duct network surrounded by renal interstitium and endothelial cells. Within these organoids, individual nephrons segment into distal and proximal tubules, early loops of Henle, and glomeruli containing podocytes elaborating foot processes and undergoing vascularization. When transcription profiles of kidney organoids were compared to human fetal tissues, they showed highest congruence with first trimester human kidney. Furthermore, the proximal tubules endocytose dextran and differentially apoptose in response to cisplatin, a nephrotoxicant. Such kidney organoids represent powerful models of the human organ for future applications, including nephrotoxicity screening, disease modelling and as a source of cells for therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
The RNA-seq data have been deposited in the Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE70101.
References
Takasato, M. et al. Directing human embryonic stem cell differentiation towards a renal lineage generates a self-organizing kidney. Nature Cell Biol. 16, 118–126 (2014)
James, R. G. & Schultheiss, T. M. Patterning of the avian intermediate mesoderm by lateral plate and axial tissues. Dev. Biol. 253, 109–124 (2003)
Mae, S. et al. Monitoring and robust induction of nephrogenic intermediate mesoderm from human pluripotent stem cells. Nat. Commun. 4, 1367 (2013)
Xia, Y. et al. Directed differentiation of human pluripotent cells to ureteric bud kidney progenitor-like cells. Nature Cell Biol. 15, 1507–1515 (2013)
Taguchi, A. et al. Redefining the in vivo origin of metanephric nephron progenitors enables generation of complex kidney structures from pluripotent stem cells. Cell Stem Cell 14, 53–67 (2014)
Lam, A. Q. et al. Rapid and efficient differentiation of human pluripotent stem cells into intermediate mesoderm that forms tubules expressing kidney proximal tubular markers. J. Am. Soc. Nephrol. 25, 1211–1225 (2014)
Kang, M. & Han, Y. M. Differentiation of human pluripotent stem cells into nephron progenitor cells in a serum and feeder free system. PLoS One 9, e94888 (2014)
Xu, J. et al. Eya1 interacts with Six2 and Myc to regulate expansion of the nephron progenitor pool during nephrogenesis. Dev. Cell 31, 434–447 (2014)
Duester, G. Retinoic acid synthesis and signaling during early organogenesis. Cell 134, 921–931 (2008)
Sakai, Y. et al. The retinoic acid-inactivating enzyme CYP26 is essential for establishing an uneven distribution of retinoic acid along the anterio-posterior axis within the mouse embryo. Genes Dev. 15, 213–225 (2001)
Abu-Abed, S. et al. The retinoic acid-metabolizing enzyme, CYP26A1, is essential for normal hindbrain patterning, vertebral identity, and development of posterior structures. Genes Dev. 15, 226–240 (2001)
Sweetman, D., Wagstaff, L., Cooper, O., Weijer, C. & Münsterberg, A. The migration of paraxial and lateral plate mesoderm cells emerging from the late primitive streak is controlled by different Wnt signals. BMC Dev. Biol. 8, 63 (2008)
Takasato, M. & Little, M. H. The origin of the mammalian kidney: implications for recreating the kidney in vitro. Development 142, 1937–1947 (2015)
Park, J. S. et al. Six2 and Wnt regulate self-renewal and commitment of nephron progenitors through shared gene regulatory networks. Dev. Cell 23, 637–651 (2012)
Roost, M. S. et al. KeyGenes, a tool to probe tissue differentiation using a human fetal transcriptional atlas. Stem Cell Reports 4, 1112–1124 (2015)
Mugford, J. W., Sipilä, P., McMahon, J. A. & McMahon, A. P. Osr1 expression demarcates a multi-potent population of intermediate mesoderm that undergoes progressive restriction to an Osr1-dependent nephron progenitor compartment within the mammalian kidney. Dev. Biol. 324, 88–98 (2008)
Sims-Lucas, S. et al. Endothelial progenitors exist within the kidney and lung mesenchyme. PLoS One 8, e65993 (2013)
Brunskill, E. W., Georgas, K., Rumballe, B., Little, M. H. & Potter, S. S. Defining the molecular character of the developing and adult kidney podocyte. PLoS One 6, e24640 (2011)
Kobayashi, A. et al. Identification of a multipotent self-renewing stromal progenitor population during mammalian kidney organogenesis. Stem Cell Reports 3, 650–662 (2014)
Floege, J. et al. Localization of PDGF alpha-receptor in the developing and mature human kidney. Kidney Int. 51, 1140–1150 (1997)
Loughna, S., Yuan, H. T. & Woolf, A. S. Effects of oxygen on vascular patterning in Tie1/LacZ metanephric kidneys in vitro. Biochem. Biophys. Res. Commun. 247, 361–366 (1998)
Brunskill, E. W. et al. Atlas of gene expression in the developing kidney at microanatomic resolution. Dev. Cell 15, 781–791 (2008)
Thiagarajan, R. D. et al. Identification of anchor genes during kidney development defines ontological relationships, molecular subcompartments and regulatory pathways. PLoS One 6, e17286 (2011)
Cheng, X. & Klaassen, C. D. Tissue distribution, ontogeny, and hormonal regulation of xenobiotic transporters in mouse kidneys. Drug Metab. Dispos. 37, 2178–2185 (2009)
Mese, H., Sasaki, A., Nakayama, S., Alcalde, R. E. & Matsumura, T. The role of caspase family protease, caspase-3 on cisplatin-induced apoptosis in cisplatin-resistant A431 cell line. Cancer Chemother. Pharmacol. 46, 241–245 (2000)
Cummings, B. S. & Schnellmann, R. G. Cisplatin-induced renal cell apoptosis: caspase 3-dependent and -independent pathways. J. Pharmacol. Exp. Ther. 302, 8–17 (2002)
Short, K. M. et al. Global quantification of tissue dynamics in the developing mouse kidney. Dev. Cell 29, 188–202 (2014)
Briggs, J. A. et al. Integration-free induced pluripotent stem cells model genetic and neural developmental features of down syndrome etiology. Stem Cells 31, 467–478 (2013)
Takasato, M., Er, X. P., Chiu, S. H. & Little, H. M. Generation of kidney organoids from human pluripotent stem cells. Protoc. Exch. http://dx.doi.org/10.1038/protex.2015.087 (2015)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Acknowledgements
This research was supported by National Health and Medical Research Council (NHMRC) of Australia (APP1041277, APP1037320), Australian Research Council (ARC) (SRI110001002, CE140100036), Bontius Stiching and Organovo Inc. M.H.L. and R.G.P. are NHMRC Senior Principal Research Fellows. B.M. is a Rosamond Siemon Postgraduate Scholar. We thank A. Christ and T. Bruxner at the IMB Sequencing Facility for providing NGS service. We also acknowledge the IMB ACRF Imaging Facility and the Australian Microscopy & Microanalysis Research Facility at the Center for Microscopy and Microanalysis at The University of Queensland.
Author information
Authors and Affiliations
Contributions
M.T. and M.H.L. planned the project, designed experiments, analysed and interpreted data and wrote the manuscript. M.T. performed experiments. P.X.E. maintained hES/iPS cells. P.X.E. and H.S.C. performed experiments under the supervision of M.T and M.H.L.; B.M. generated organoids for TEM. G.J.B. analysed bioinformatic data. C.F. performed TEM. R.G.P. captured and interpreted TEM images. E.J.W. provided the iPS cell line and advised on general iPS cell quality control. M.S.R. and S.M.C.d.S.L. developed NGS analytical tools and analysed data for RNA-seq profiling.
Corresponding authors
Ethics declarations
Competing interests
M.T. and M.H.L. are named inventors on a patent relating to this methodology. Some research funding was provided by Organovo Inc.
Extended data figures and tables
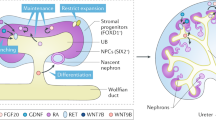
Extended Data Figure 1 Antero-posterior intermediate mesoderm specification is regulated by the timing of FGF9 exposure and the presence of RA signalling.
a, Immunofluorescence at day 18 of monolayer differentiation from cultures exposed to different timing of FGF9 addition (after 2, 3, 4 and 5 days of CHIR99021). The ureteric epithelium is represented by GATA3+PAX2+ECAD+ cells. The metanephric mesenchyme and its derivatives are marked by PAX2+GATA3−ECAD− (metanephric mesenchyme) and PAX2+GATA3−ECAD+ (nephrons), respectively. Scale bars, 100 μm. b, Immunofluorescence at day 7 and 18 of monolayer differentiation using 5 days of CHIR99021 followed by RA or AGN193109 (AGN) on top of FGF9. RA reduced the specification of posterior intermediate mesoderm, as indicated by the reduction of HOXD11 at day 7 (top panel). This resulted in less metanephric mesenchyme but some ureteric epithelium by RA at day 18 (bottom panel). Scale bars, 100 μm.
Extended Data Figure 2 Induction of both kidney progenitors at the same time.
a, b, Immunofluorescence at day 18 of the monolayer differentiation using the 4 days CHIR99021 before FGF9 protocol. The metanephric mesenchyme is marked by SIX2+SIX1+HOXD11+ cells (a). GATA3+PAX2+ECAD+KRT8+ cells representing the ureteric epithelium were also induced (b). Scale bars, 50 μm.
Extended Data Figure 3 Regulation of nephrogenesis in the kidney organoid.
a, Stimulating organoids with 5 μM CHIR99021 for 1 h immediately after aggregation promoted nephrogenesis (CHIR pulse), whereas only limited numbers of nephrogenesis events happened without CHIR99021 (no pulse). Scale bars, 1 mm. b, Without the addition of FGF9 after this CHIR99021 pulse, organoids did not initiate nephrogenesis (−FGF9). Scale bars, 200 μm.
Extended Data Figure 4 The timing of FGF9 exposure affects the ratio of collecting duct to nephron in the kidney organoid.
a, b, Immunofluorescence of kidney organoids at day 18 after-aggregation after exposure to different timings of initial FGF9 exposure (2, 3, 4 and 5 days of CHIR99021 pre-FGF9), demonstrating the regulation of collecting duct/nephron ratio by varying this timing. GATA3+ECAD+ cells represent the collecting duct (a), whereas WT1+NPHS1+ cells mark podocytes of the glomerulus (b). Scale bars, 200 μm.
Extended Data Figure 5 Changes of gene expression during development of the kidney organoid.
a–c, Graphs showing expression changes of selected marker genes at 4 time points (day 0, 3, 11 and 18) of the kidney organoid culture. y axis represents the count of detection for each gene in an RNA sequencing analysis. Markers of the nephron progenitor (cap mesenchyme) and collecting duct progenitor (ureteric tip) were peaked by day 3 then dropped (a). Markers of early nephron increased by day 3, while those of mature nephron components (Proximal and distal tubule and Podocytes) started after day 3. Illustrations show expression regions (blue coloured) of each selected gene in the developing kidney (b). Markers of endothelial and renal interstitial cells were also increased by day 11 (c).
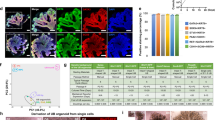
Extended Data Figure 6 Transcriptional similarity of the kidney organoid to human fetal organs.
Dendrogram showing the hierarchical clustering of day 0, 3, 11 and 18 differentiation experiments and 21 human fetal organs from first and second trimester (Gene Expression Omnibus accession number GSE66302)15. Sample name is composed of individual ID followed by an organ name and gestation week. For instance, ‘DJ1 kidney_9’ represents a kidney at ninth week gestation from individual ID: DJ1. Day 0 and 3 kidney organoids cluster with gonad, in agreement with the common origin of both gonad and kidney from the intermediate mesoderm. Day 11 and 18 kidney organoids show strongest similarity to trimester 1 human kidney. The classifier genes used for this analysis are detailed in Supplementary Table 3.
Extended Data Figure 7 Evidence of endothelial cells in the kidney organoid.
a, Immunofluorescence of day 11 kidney organoids showing the presence of CD31+KDR+ endothelial cells surrounding NPHS1+ glomeruli. Scale bar, 100 μm. b, Two representative images demonstrating the expression of another endothelium marker SOX17 in CD31+ endothelial cells. Scale bars, 100 μm. c, Immunofluorescence of day 18 kidney organoids displaying endothelia with lumen formation, as indicated by asterisks. This image also shows the endothelial invasion into a glomerulus. Scale bar, 100 μm.
Extended Data Figure 8 Characterization of non-epithelial structures in the kidney organoid.
All images were taken from day 18 kidney organoids. a, PDGFRA+ pericytic cells attaching on KDR+ vessels. Scale bar, 50 μm. b, Some glomeruli contained PDGFRA+ cells likely to represent early mesangial cells19. Scale bar, 50 μm. c, Laminin staining (LAM) demonstrates the presence of basement membrane in glomerulus structures (white arrowheads). Scale bar, 100 μm. d, TEM images of avascular glomeruli showing early podocytes surrounding a basement membrane (yellow arrowheads) and exhibiting foot processes on the basement membrane. e, Immunofluorescence showing FOXD1 expression in podocytes (WT1+FOXD1+)18 and a subpopulation of MEIS1+ interstitium (white arrowheads). This is suggestive of the presence of both cortical stroma (FOXD1+MEIS1+) and medullary stroma (FOXD1−MEIS1+). Scale bar, 100 μm.
Extended Data Figure 9 Functional assay of proximal tubule maturation within kidney organoids.
a, Fluorescent microscopy showing the dextran uptake in both the kidney organoids and E14 mouse embryonic kidneys organ culture after 24 h presence of dextran–Alexa488 (10 μg ml−1) in the culture medium (24 h dextran–Alexa488). 1 h incubation was insufficient for either organoids or mouse kidney explants to uptake dextran from the culture media (1 h dextran–Alexa488). No background signals were detected in a control without dextran (no dextran). Dashed line circles the organoids and kidneys. Scale bars, 1 mm. b, Endocytosis mediator cubilin (CUBN) was present on apical surface of the proximal tubules in kidney organoids (left panel). The same staining without detergent during the process showed the complete absence of CUBN staining on apical surface (right panel), demonstrating that the tubules within the organoids are intact. This explains the requirement for a 24 h incubation with dextran before evidence of apical uptake. Dashed line circles LTL+ proximal tubules. Scale bars, 50 μm. c, Low power immunofluorescence microscopy of day 18 kidney organoids after being treated by cisplatin for 24 h. No apoptosis was observed in proximal tubules in the absence of cisplatin (0 μM, left panel). LTL+ECAD+ proximal tubular cell-specific apoptosis was observed only in response to either 5 μM (not shown) or 20 μM cisplatin (arrowheads in middle panel). Global cell death was observed after culture in 100 μM cisplatin (right panel). Scale bars, 100 μm.
Supplementary information
Supplementary Table 1
This table contains a list of primer sequences used for qPCR. (XLSX 41 kb)
Supplementary Table 2
This table shows RNA-seq profiling of organoid differentiation timecourse (day 0, 3, 11, 18). Data presented is normalized read counts including average counts for each timepoint. (XLSX 5655 kb)
Supplementary Table 3
This file contains a table of 85 Classifer genes15 used for the unbiased comparative analysis of RNAseq profiling data generated from trimester 1 (1T) and trimester 2 (2T) human fetal tissues compared to kidney organoids day 0, 3, 11 and 18 after commencement of 3D culture (Figure 2g, Extended Data Figure 6). (XLSX 13 kb)
Z-stacks within developing kidney organoids at day 11 of culture
Confocal microscopic Z-stack images from the bottom to the top of kidney organoids after 11 days in 3D culture. GATA3+ECAD+ CD, yellow with green nucleus; ECAD+ DT, yellow; LTL+ PT, red; NPHS1+ glomerulus, green membrane; DAPI, blue. Fields of view: 354.25 μm x 354.25 μm. (MOV 12982 kb)
Z-stacks within developing kidney organoids at day 11 of culture
Confocal microscopic Z-stack images from the bottom to the top of kidney organoids after 11 days in 3D culture. GATA3+ECAD+ CD, yellow with green nucleus; ECAD+ DT, yellow; LTL+ PT, red; NPHS1+ glomerulus, green membrane; DAPI, blue. Fields of view: 354.25 μm x 354.25 μm. (MOV 14698 kb)
Z-stacks of vascularizing glomeruli within a kidney organoid
Both videos represent confocal microscopic Z-stack images within a day 18 organoid scanning through vascularizing glomeruli. NPHS1+ podocytes of the glomerulus, green; CD31+ endothelium, pink; DAPI, blue. Fields of view: 212.55 μm x 212.55 μm. (MP4 3231 kb)
Z-stacks of vascularizing glomeruli within a kidney organoid
Both videos represent confocal microscopic Z-stack images within a day 18 organoid scanning through vascularizing glomeruli. NPHS1+ podocytes of the glomerulus, green; CD31+ endothelium, pink; DAPI, blue. Fields of view: 212.55 μm x 212.55 μm. (MP4 2669 kb)
Rights and permissions
About this article
Cite this article
Takasato, M., Er, P., Chiu, H. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568 (2015). https://doi.org/10.1038/nature15695
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15695
This article is cited by
-
Insights on Three Dimensional Organoid Studies for Stem Cell Therapy in Regenerative Medicine
Stem Cell Reviews and Reports (2024)
-
A disease-specific iPS cell resource for studying rare and intractable diseases
Inflammation and Regeneration (2023)
-
Review on kidney diseases: types, treatment and potential of stem cell therapy
Renal Replacement Therapy (2023)
-
HIF-1α promotes kidney organoid vascularization and applications in disease modeling
Stem Cell Research & Therapy (2023)
-
The interplay of cells, polymers, and vascularization in three-dimensional lung models and their applications in COVID-19 research and therapy
Stem Cell Research & Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.