Abstract
Enhancers, critical determinants of cellular identity, are commonly recognized by correlative chromatin marks and gain-of-function potential, although only loss-of-function studies can demonstrate their requirement in the native genomic context. Previously, we identified an erythroid enhancer of human BCL11A, subject to common genetic variation associated with the fetal haemoglobin level, the mouse orthologue of which is necessary for erythroid BCL11A expression. Here we develop pooled clustered regularly interspaced palindromic repeat (CRISPR)-Cas9 guide RNA libraries to perform in situ saturating mutagenesis of the human and mouse enhancers. This approach reveals critical minimal features and discrete vulnerabilities of these enhancers. Despite conserved function of the composite enhancers, their architecture diverges. The crucial human sequences appear to be primate-specific. Through editing of primary human progenitors and mouse transgenesis, we validate the BCL11A erythroid enhancer as a target for fetal haemoglobin reinduction. The detailed enhancer map will inform therapeutic genome editing, and the screening approach described here is generally applicable to functional interrogation of non-coding genomic elements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981)
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 488, 75–82 (2012)
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014)
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009)
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010)
Xu, J. et al. Combinatorial assembly of developmental stage-specific enhancers controls gene expression programs during human erythropoiesis. Dev. Cell 23, 796–811 (2012)
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011)
Parker, S. C. J. et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants. Proc. Natl Acad. Sci. USA 110, 17921–17926 (2013)
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013)
Paul, D. S. et al. Maps of open chromatin guide the functional follow-up of genome-wide association signals: Application to hematological traits. PLoS Genet. 7, e1002139 (2011)
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012)
Hardison, R. C. Variable evolutionary signatures at the heart of enhancers. Nature Genet. 42, 734–735 (2010)
Vierstra, J. et al. Mouse regulatory DNA landscapes reveal global principles of cis-regulatory evolution. Science 346, 1007–1012 (2014)
Villar, D. et al. Enhancer evolution across 20 mammalian species. Cell 160, 554–566 (2015)
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006)
Melnikov, A. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nature Biotechnol. 30, 271–277 (2012)
Patwardhan, R. P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nature Biotechnol. 30, 265–270 (2012)
Sexton, T. & Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell 160, 1049–1059 (2015)
Bender, M., Bulger, M., Close, J. & Groudine, M. β-globin gene switching and DNase I sensitivity of the endogenous β-globin locus in mice do not require the locus control region. Mol. Cell 5, 387–393 (2000)
Johnson, K. D. et al. Cis-element mutated in GATA2-dependent immunodeficiency governs hematopoiesis and vascular integrity. J. Clin. Invest. 122, 3692–3704 (2012)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013)
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014)
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014)
Koike-Yusa, H., Li, Y., Tan, E.-P., Velasco-Herrera, M. D. C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nature Biotechnol. 32, 267–273 (2014)
Zhou, Y. et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014)
Bauer, D. E. et al. An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science 342, 253–257 (2013)
Gröschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in Leukemia. Cell 157, 369–381 (2014)
Mansour, M. R. et al. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346, 1373–1377 (2014)
Sankaran, V. G. et al. Human fetal hemoglobin expression is regulated by the developmental stage-specific repressor BCL11A. Science 322, 1839–1842 (2008)
Sankaran, V. G. et al. Developmental and species-divergent globin switching are driven by BCL11A. Nature 460, 1093–1097 (2009)
Xu, J. et al. Correction of sickle cell disease in adult mice by interference with fetal hemoglobin silencing. Science 334, 993–996 (2011)
Kurita, R. et al. Establishment of immortalized human erythroid progenitor cell lines able to produce enucleated red blood cells. PLoS ONE 8, e59890 (2013)
Hardison, R. C. & Blobel, G. A. GWAS to therapy by genome edits? Science 342, 206–207 (2013)
Canver, M. C. et al. Characterization of genomic deletion efficiency mediated by clusted regularly interspaced palindromic repeats (CRISPR)/Cas9 nuclease system in mammalian cells. J. Biol. Chem. 289, 21312–21324 (2014)
Mandal, P. K. et al. Efficient ablation of genes in human hematopoietic stem and effector cells using CRISPR/Cas9. Cell Stem Cell 15, 643–652 (2014)
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013)
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nature Biotechnol. 31, 827–832 (2013)
Liu, P. et al. Bcl11a is essential for normal lymphoid development. Nature Immunol. 4, 525–532 (2003)
John, A. et al. Bcl11a is required for neuronal morphogenesis and sensory circuit formation in dorsal spinal cord development. Development 139, 1831–1841 (2012)
Yu, Y. et al. Bcl11a is essential for lymphoid development and negatively regulates p53. J. Exp. Med. 209, 2467–2483 (2012)
Porcu, B. S. et al. The human β globin locus introduced by YAC transfer exhibits a specific and reproducible pattern of developmental regulation in transgenic mice. Blood 90, 4602–4609 (1997)
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015)
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered and improved PAM specificities. Nature 523, 481–485 (2015)
Esvelt, K. M. et al. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nature Methods 10, 1116–1121 (2013)
Findlay, G. M., Boyle, E. A., Hause, R. J., Klein, J. C. & Shendure, J. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014)
Bauer, D. E., Kamran, S. C. & Orkin, S. H. Reawakening fetal hemoglobin: Prospects for new therapies for the beta-globin disorders. Blood 120, 2945–2953 (2012)
Basak, A. et al. Persistence of fetal hemoglobin and altered neurodevelopment due to BCL11A deletions. JCI 125, 2363–2368 (2015)
Funnell, P. W. et al. 2p15-p16.1 microdeletions encompassing and proximal to BCL11A are associated with elevated HbF in addition to neurologic impairment. Blood 126, 89–93 (2015)
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nature Methods 11, 783–784 (2014)
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015)
Giarratana, M. et al. Proof of principle for transfusion of in vitro generated red blood cells. Blood 118, 5071–5079 (2011)
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014)
Kowalczyk, M. S. et al. Intragenic enhancers act as alternative promoters. Mol. Cell 45, 447–458 (2012)
Dogan, N. et al. Occupancy by key transcription factors is a more accurate predictor of enhancer activity than histone modifications or chromatin accessibility. Epigenetics Chromatin 8, 16 (2015)
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011)
Mathelier, A. et al. JASPAR 2014: An extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, 142–147 (2014)
Weber, K., Bartsch, U., Stocking, C. & Fehse, B. A multicolor panel of novel lentiviral ‘gene ontology’ (LeGO) vectors for functional gene analysis. Mol. Ther. 16, 698–706 (2008)
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nature Biotechnol. 32, 1262–1267 (2014)
Acknowledgements
We thank J. Hughes and D. Higgs for assistance with analysis of ChIP-seq; R. Mathieu and the Boston Children’s Hospital Hematology/Oncology-HSCI Flow Cytometry Research Facility for cell sorting; Z. Herbert and F. Abderazzaq at the Dana-Farber Cancer Institute Molecular Biology Core Facility and Center for Cancer Computational Biology, respectively, for sequencing; J. Doench for providing TALENs; C. Peng for advice with MEL reporter cell generation; F. Godinho and M. Nguyen for technical help with ESCs and transgenic mice; A. Dass, C. Lin and S. Kamran for general technical assistance; C. Brendel and D. Williams for input regarding lentiviral transduction of HSPCs; J. Desimini for graphical assistance; and J. Xu and G. Lettre for insightful discussions. M.C.C. is supported by F30DK103359-01A1. E.C.S. is supported by a Jane Coffin Childs Memorial Fund for Medical Research Fellowship. L.P. is supported by NHGRI Career Development Award K99HG008399. N.E.S. is supported by a Simons Center for the Social Brain Postdoctoral Fellowship and NIH NHGRI award K99-HG008171. O.S. is supported by a fellowship from the Klarman Family Foundation. S.L. is supported by a Leukemia & Lymphoma Society Fellow Award. T.M. is supported by NIH R01 A1084905. G.-C.Y. is supported by NIH R01HL119099 and R01HG005085. F.Z. is supported by the NIMH (5DP1-MH100706) and NIDDK (5R01-DK097768), a Waterman award from the National Science Foundation, the Keck, McKnight, Damon Runyon, Searle Scholars, Merkin, Vallee, and Simons Foundations, and Bob Metcalfe. S.H.O. is supported by P01HL032262 and P30DK049216 (Center of Excellence in Molecular Hematology). D.E.B. is supported by an NIDDK Career Development Award K08DK093705, Doris Duke Charitable Foundation Innovations in Clinical Research Award (2013137), and Charles H. Hood Foundation Child Health Research Award. Computational tools and instructions for designing CRISPR-Cas9 sgRNA libraries for conducting non-coding screening can be found at the Zhang laboratory website http://www.genome-engineering.org.
Author information
Authors and Affiliations
Contributions
D.E.B. conceived this study. N.E.S., O.S. and F.Z. conceived the pooled non-coding screening strategy using CRISPR-Cas9. M.C.C., N.E.S., O.S., F.Z., S.H.O. and D.E.B. designed and executed the pooled CRISPR screening strategy. E.C.S., F.S., Y.F., S.L., S.H.O. and D.E.B. designed, produced and analysed the transgenic mice. R.K. and Y.N. provided the HUDEP-2 cell line. M.C.C., F.S., T.M., S.H.O. and D.E.B. adapted the HUDEP-2 cell line as a model of globin gene regulation. M.C.C., F.S., D.D.C., P.G.S., D.S.V. and D.E.B. performed all experiments in cell lines. M.C.C., L.P., N.E.S., S.P.G., G.-C.Y., F.Z., S.H.O. and D.E.B. analysed the data. L.P., S.P.G. and G.-C.Y. developed the HMM. M.C.C., S.H.O., and D.E.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
D.E.B. and S.H.O. are inventors on a patent related to this work. N.E.S., O.S. and F.Z. are inventors on a patent application related to the screening technology. F.Z. is a founder of Editas Medicine and scientific advisor for Editas Medicine and Horizon Discovery. S.H.O. is on the Scientific Advisory Board of Editas Medicine.
Additional information
All reagents described in this manuscript have been deposited with Addgene (http://www.addgene.org).
Extended data figures and tables
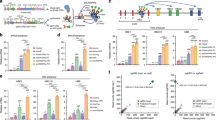
Extended Data Figure 1 Human BCL11A locus.
a, Schematic of the human BCL11A locus (hg19, transcription from right to left) with erythroid chromatin marks and trait-associated haplotype denoted, and composite enhancer as previously defined28. b, Ranked enhancers in primary human adult erythroid precursors by H3K27ac signal intensity, with super-enhancers shaded, and super-enhancer-associated genes indicated.
Extended Data Figure 2 Tiled pooled in situ CRISPR-Cas9 BCL11A enhancer screen.
a, Distribution of NGG and NAG PAM sgRNAs mapped to genomic cleavage position. The vertical lines represent cleavage sites for sgRNAs mapped to plus and minus strands. b, Gap distance between adjacent genomic cleavage position for NAG PAM sgRNAs. c, Library composition by target sequence and PAM restriction. d, Representation of both NGG and NAG sgRNA (1,338 sgRNAs in total) within the plasmid pool by deep sequencing. The median was 718 normalized reads and the 10th and 90th percentiles (indicated by the vertical dotted lines) ranged from 337 to 1,205 normalized reads. e, HbF distribution in HUDEP-2 cells transduced with Cas9 and individual sgRNAs, either non-targeting or targeting BCL11A exon 2. f, HbF enrichment scores of NGG sgRNAs in six biological replicates. g, Sort of library-transduced cells into HbF-high and HbF-low pools. h, Control sgRNA enrichment. Boxes demonstrate 25th, median, and 75th percentiles and whiskers minimum and maximum values. ****P < 0.0001, NS, non-significant. i, NGG sgRNA representation in plasmid pool and cells at conclusion of experiment (left), and in HbF-high and HbF-low pools (right), with dotted lines at x = y and x = 8y. j, Quantile–quantile plots of NGG sgRNA enrichment scores. k, Cellular dropout scores of NGG sgRNAs relative to genomic cleavage position and repetitive elements. Non-targeting sgRNAs pseudo-mapped with 5-bp spacing.
Extended Data Figure 3 Validation of enhancer screen.
a, HbF+ fraction in HUDEP-2 cells transduced in arrayed format with 24 sgRNAs from all 5 mapping categories with enrichment scores ranging from the highest to the lowest in the screen. b, Correlation between HbF enrichment score from pooled sgRNA screen and HbF+ fraction by arrayed validation of individual sgRNAs in HUDEP-2 cells. c, Erythroid differentiation of primary human erythroid precursors evaluated by CD71 and CD235a surface markers, enucleation frequency (CD235a+ Hoescht33342−), and morphology by May–Grünwald Giemsa staining.
Extended Data Figure 4 Functional assessment of enhancer sequences.
a, Topology of the HMM used to infer the three functional enhancer states (active, repressive and neutral). The emission probabilities of HbF enrichment scores from each state were modelled as Gaussian distributions (the values of μ and σ2 are shown). The transition probabilities (arrows) are displayed. b, Frequency distribution of indels from HUDEP-2 cells exposed to Cas9 and individual sgRNAs, sorted into HbF-high and HbF-low pools, and subjected to deep sequencing of the target site. Indels calculated on a per nucleotide basis throughout an amplicon surrounding the sgRNA-1617 and -1621 cleavage sites (dotted lines). An indel enrichment ratio was calculated by dividing normalized indel frequencies in the HbF-high pool by those in the HbF-low pool.
Extended Data Figure 5 Functional cores of the BCL11A enhancer.
a–c, 200 bp at the functional cores of DHSs h+55, h+58 and h+62 defined by HMM states (active, red; repressive, green). HbF enrichment scores are shown by grey lines and circles. HbF indel enrichment per nucleotide based on amplicon genomic sequencing of sorted cells exposed to either sgRNA-1617 (top) or -1621 (bottom) is shown. Common SNPs (MAF > 1%) are shown with dotted lines with HbF-low allele in blue and HbF-high allele in red; no common SNPs are present at the h+58 region. JASPAR motifs (P < 10−4) are depicted in black except for those with allele-specific significance depicted by allelic colour. Selected motifs annotated by transcription factor on the basis of known erythroid-specific function or genomic position. Motif LOGOs at key positions with motif scores P < 10−3 as described in text. Dotted boxes show regions of highest HbF enrichment score at each core with underlying predicted motifs. Orthologous sequences are listed from representative primates and nonprimates of distributed phylogeny. PhyloP (scale from −4.5 to 4.88) and PhastCons (from 0 to 1) estimates of evolutionary conservation among 100 vertebrates are shown. An arrow indicates a 144 bp insertion in the mouse genome relative to the human reference adjacent to the orthologous GATA1 motif at h+58.
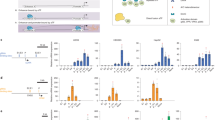
Extended Data Figure 6 Tiled pooled in situ CRISPR-Cas9 Bcl11a enhancer screen.
a, Schematic of the mouse Bcl11a locus (mm9, transcription from left to right) with erythroid chromatin marks (top, dark blue H3K27ac from ref. 55; middle, light blue H3K27ac from ref. 56; and bottom, black DNase I from ref. 28) and regions of primary sequence homology to the human DHSs displayed. y axes for H3K27ac tracks are both scaled to maximum 3.5 reads per million. Composite enhancer as previously defined28. b, Ranked enhancers in mouse erythroid precursors by H3K27ac signal intensity55,56, with super-enhancers shaded. Super-enhancer associated genes are indicated by Venn diagram. c, Strategy to knock-in by homology-directed repair the fluorescent protein mCherry into the mouse embryonic globin Hbb-y locus (encoding the εy embryonic globin chain). d, Distribution of NGG and NAG PAM sgRNAs mapped to genomic cleavage position with vertical lines representing cleavage sites for sgRNAs mapped to plus and minus strands. e, Distance to adjacent genomic cleavage position for NGG (left) and NAG (right) PAM sgRNAs. f, Representation of the 1,271 NGG and NAG sgRNAs within the plasmid pool by deep sequencing. The median was 735 normalized reads and the 10th and 90th percentiles (indicated by the vertical dotted lines) ranged from 393 to 1,240 normalized reads. g, Library composition by target sequence and PAM restriction. h, mCherry expression upon exposure to Cas9 and an individual NGG sgRNA targeting Bcl11a exon 2 in MEL εy:mCherry reporter cells. i, εy:mCherry sort of library transduced cells. j, Control sgRNA enrichment. Boxes demonstrate 25th, median and 75th percentiles and whiskers minimum and maximum values. ****P < 0.0001. k, Enrichment scores of NGG sgRNAs between four biological replicates.
Extended Data Figure 7 Bcl11a enhancer screen analyses.
a, NGG sgRNA representation in plasmid pool and cells at conclusion of experiment (left), and in εy:mCherry-high and εy:mCherry-low pools (right), with dotted lines at x = y and x = 8y. b, Quantile–quantile plots of NGG sgRNA εy enrichment scores. c, Cellular dropout scores of NGG sgRNAs relative to genomic cleavage position and repetitive elements. Non-targeting sgRNAs are pseudo-mapped with 5 bp spacing. d, Correlation between cellular dropout and εy enrichment scores.
Extended Data Figure 8 Functional sequences at the Bcl11a erythroid enhancer.
a–c, HMM segmentation of active functional states at m+55, m+58 and m+62 orthologues. HbF enrichment scores are shown as grey lines and circles with the blue line representing smoothed enrichment score. DNase I sequencing from mouse fetal liver erythroid precursors28. PhyloP (scale from −3.3 to 2.1) and PhastCons (from 0 to 1) estimates of evolutionary conservation among 30 vertebrates are shown. d, Top: Bcl11a expression determined by RT–qPCR displayed as a heat map in 108 hemizygous m+62 orthologue deletion clones ordered by genomic position of deletion midpoint. Each bar demonstrates the genomic position of the deletion breakpoints and the associated colour demonstrates the level of Bcl11a expression. Bottom: Bcl11a expression determined by RT–qPCR in 108 hemizygous m+62 orthologue deletion clones. Per nucleotide mean effect size was calculated as the mean fold change in Bcl11a expression from all clones in which that nucleotide was deleted. Grey shading represents one s.d. The Bcl11a expression data are shown with same x axis as in panel c immediately above.
Extended Data Figure 9 Evaluation of the m+62 functional core.
200 bp at the functional core of the m+62 orthologue defined by HMM state. Enrichment scores are shown as grey lines and circles with the blue line representing smoothed enrichment score. JASPAR motifs (P < 10−4) are depicted with selected motifs annotated by transcription factor name on the basis of known erythroid-specific function or genomic position. Orthologous human sequences are listed. PhyloP (scale from −3.3 to 2.1) and PhastCons (from 0 to 1) estimates of evolutionary conservation among 30 vertebrates are shown. Individual numbered hemizygous deletion clones with indicated breakpoints were evaluated by BCL11A immunoblot (C, control). Clones 9 and 10 encompass the entire m+62 orthologue.
Extended Data Figure 10 Requirement of Bcl11a erythroid enhancer during murine ontogeny.
a, Progeny of heterozygous Bcl11a m+62 orthologue deletion intercrosses as compared to expected Mendelian ratio. b, Fraction of fetal liver comprised of B-cell progenitors at E16.5 from various genotypes. c, Peripheral blood analysis from 4-week-old mice to examine the frequency of various circulating haematopoietic lineages in Bcl11a m+62 orthologue deletion wild-type, heterozygous, and homozygous mice. d, Bcl11a expression in β-YAC/+62 deletion mice (each symbol represents the mean expression from technical replicates from an individual mouse). *P < 0.05; error bars represent s.e.m.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary References and Supplementary Tables 1-5. (PDF 359 kb)
Source data
Rights and permissions
About this article
Cite this article
Canver, M., Smith, E., Sher, F. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015). https://doi.org/10.1038/nature15521
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15521
This article is cited by
-
Detection of CRISPR/Cas9-Mediated Fetal Hemoglobin Reactivation in Erythroblasts Derived from Cord Blood-Hematopoietic Stem Cells
Molecular Biotechnology (2024)
-
Base Editors-Mediated Gene Therapy in Hematopoietic Stem Cells for Hematologic Diseases
Stem Cell Reviews and Reports (2024)
-
METTL16 promotes liver cancer stem cell self-renewal via controlling ribosome biogenesis and mRNA translation
Journal of Hematology & Oncology (2024)
-
Multicenter integrated analysis of noncoding CRISPRi screens
Nature Methods (2024)
-
Viral vectors and extracellular vesicles: innate delivery systems utilized in CRISPR/Cas-mediated cancer therapy
Cancer Gene Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.