Abstract
Somaclonal variation arises in plants and animals when differentiated somatic cells are induced into a pluripotent state, but the resulting clones differ from each other and from their parents. In agriculture, somaclonal variation has hindered the micropropagation of elite hybrids and genetically modified crops, but the mechanism responsible remains unknown1. The oil palm fruit ‘mantled’ abnormality is a somaclonal variant arising from tissue culture that drastically reduces yield, and has largely halted efforts to clone elite hybrids for oil production2,3,4. Widely regarded as an epigenetic phenomenon5, ‘mantling’ has defied explanation, but here we identify the MANTLED locus using epigenome-wide association studies of the African oil palm Elaeis guineensis. DNA hypomethylation of a LINE retrotransposon related to rice Karma, in the intron of the homeotic gene DEFICIENS, is common to all mantled clones and is associated with alternative splicing and premature termination. Dense methylation near the Karma splice site (termed the Good Karma epiallele) predicts normal fruit set, whereas hypomethylation (the Bad Karma epiallele) predicts homeotic transformation, parthenocarpy and marked loss of yield. Loss of Karma methylation and of small RNA in tissue culture contributes to the origin of mantled, while restoration in spontaneous revertants accounts for non-Mendelian inheritance. The ability to predict and cull mantling at the plantlet stage will facilitate the introduction of higher performing clones and optimize environmentally sensitive land resources.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Gene Expression Omnibus
Sequence Read Archive
Data deposits
Microarray data have been deposited in the NCBI Gene Expression Omnibus (GEO) and are accessible through GEO Series accession number GSE68410. Small RNA sequence data from the region of interest have been deposited in the NCBI Sequence Read Archive (SRA) database under the accession numbers SAMN03569290–SAMN03569351. Whole-genome bisulfite sequence data have been deposited in the NCBI SRA under accession numbers SAMN03569063–SAMN03569077. The cDNA sequence of the kDEF1 transcript has been deposited in GenBank under accession number KR347486.
References
Stroud, H. et al. Plants regenerated from tissue culture contain stable epigenome changes in rice. Elife 2, e00354 (2013)
Corley, R. H. V. in CRC Handbook of Fruit Set and Development (ed. Monselise, S. P.) 253–259 (CRC Press, 1986)
Zeven, A. C. The ‘mantled’ oil palm (Elaeis guineensis Jacq.). J. W. Afr. Inst. Oil Palm Res. 5, 31–33 (1973)
Mgbeze, G. C. & Iserhienrhien, A. Somaclonal variation associated with oil palm (Elaeis guineensis Jacq.) clonal propagation: a review. Afr. J. Biotechnol. 13, 989–997 (2014)
Jaligot, E., Rival, A., Beule, T., Dussert, S. & Verdeil, J. L. Somaclonal variation in oil palm (Elaeis guineensis Jacq.): the DNA methylation hypothesis. Plant Cell Rep. 19, 684–690 (2000)
Corley, R. H. V. & Law, I. H. in Plantation Management for the 21st Century (ed. Pushparajah, E.) 279–289 (Incorp. Soc. Planters, 1997)
Singh, R. et al. The oil palm SHELL gene controls oil yield and encodes a homologue of SEEDSTICK. Nature 500, 340–344 (2013)
Adam, H. et al. Functional characterization of MADS box genes involved in the determination of oil palm flower structure. J. Exp. Bot. 58, 1245–1259 (2007)
Rao, V. & Donough, C. R. Preliminary evidence of a genetic cause for the floral abnormalities in some oil palm ramets. Elaies 2, 199–207 (1990)
Matthes, M., Singh, R., Cheah, S. C. & Karp, A. Variation in oil palm (Elaeis guineensis Jacq.) tissue culture-derived regenerants revealed by AFLPs with methylation-sensitive enzymes. Theor. Appl. Genet. 102, 971–979 (2001)
Kubis, S. E., Castilho, A. M., Vershinin, A. V. & Heslop-Harrison, J. S. Retroelements, transposons and methylation status in the genome of oil palm (Elaeis guineensis) and the relationship to somaclonal variation. Plant Mol. Biol. 52, 69–79 (2003)
Jaligot, E. et al. DNA methylation and expression of the EgDEF1 gene and neighboring retrotransposons in mantled somaclonal variants of oil palm. PLoS ONE 9, e91896 (2014)
Syed Alwee, S. et al. Characterization of oil palm MADS box genes in relation to the mantled flower abnormality. Plant Cell Tissue Organ Cult. 85, 331–344 (2006)
Singh, R. et al. Oil palm genome sequence reveals divergence of interfertile species in Old and New worlds. Nature 500, 335–339 (2013)
Lippman, Z. et al. Role of transposable elements in heterochromatin and epigenetic control. Nature 430, 471–476 (2004)
Tanurdzic, M. et al. Epigenomic consequences of immortalized plant cell suspension culture. PLoS Biol. 6, e302 (2008)
Komatsu, M., Shimamoto, K. & Kyozuka, J. Two-step regulation and continuous retrotransposition of the rice LINE-type retrotransposon Karma. Plant Cell 15, 1934–1944 (2003)
Cui, X. et al. Control of transposon activity by a histone H3K4 demethylase in rice. Proc. Natl Acad. Sci. USA 110, 1953–1958 (2013)
Martienssen, R., Barkan, A., Taylor, W. C. & Freeling, M. Somatically heritable switches in the DNA modification of Mu transposable elements monitored with a suppressible mutant in maize. Genes Dev. 4, 331–343 (1990)
Cubas, P., Vincent, C. & Coen, E. An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401, 157–161 (1999)
Saze, H. & Kakutani, T. Heritable epigenetic mutation of a transposon-flanked Arabidopsis gene due to lack of the chromatin-remodeling factor DDM1. EMBO J. 26, 3641–3652 (2007)
Perbal, M. C., Haughn, G., Saedler, H. & Schwarz-Sommer, Z. Non-cell-autonomous function of the Antirrhinum floral homeotic proteins DEFICIENS and GLOBOSA is exerted by their polar cell-to-cell trafficking. Development 122, 3433–3441 (1996)
Regulski, M. et al. The maize methylome influences mRNA splice sites and reveals widespread paramutation-like switches guided by small RNA. Genome Res. 23, 1651–1662 (2013)
Baubec, T., Finke, A., Mittelsten Scheid, O. & Pecinka, A. Meristem-specific expression of epigenetic regulators safeguards transposon silencing in Arabidopsis. EMBO Rep. 15, 446–452 (2014)
Taniguchi-Ikeda, M. et al. Pathogenic exon-trapping by SVA retrotransposon and rescue in Fukuyama muscular dystrophy. Nature 478, 127–131 (2011)
Alló, M. et al. Control of alternative splicing through siRNA-mediated transcriptional gene silencing. Nature Struct. Mol. Biol. 16, 717–724 (2009)
Arteaga-Vazquez, M. et al. RNA-mediated trans-communication can establish paramutation at the b1 locus in maize. Proc. Natl Acad. Sci. USA 107, 12986–12991 (2010)
Lamb, R. S. & Irish, V. F. Functional divergence within the APETALA3/PISTILLATA floral homeotic gene lineages. Proc. Natl Acad. Sci. USA 100, 6558–6563 (2003)
Sommer, H. et al. Deficiens, a homeotic gene involved in the control of flower morphogenesis in Antirrhinum majus: the protein shows homology to transcription factors. EMBO J. 9, 605–613 (1990)
Jack, T., Brockman, L. L. & Meyerowitz, E. M. The homeotic gene APETALA3 of Arabidopsis thaliana encodes a MADS box and is expressed in petals and stamens. Cell 68, 683–697 (1992)
Jones, L. H. Propagation of clonal oil palms by tissue culture. Planter 50, 374–381 (1974)
Rabechault, H. & Martin, J.-P. Multiplication vegetative du palmier a huile (Elaeis guineensis Jacq.) a l’aide de cultures de tissues foliares. C. R. Acad. Sci. Paris. Ser. D 283, 1735–1737 (1976)
Durand-Gasselin, T., Le Guen, V., Konan, K. & Duval, Y. Oil palm (Elaeis guineensis Jacq.) plantations in Cote-d’Ivoire obtained through in vitro culture. First results. Oléagineux 45, 10–11 (1990)
Kushairi, A. et al. Current status of oil palm tissue culture in Malaysia. In Proceedings of the Clonal and Quality Replanting Material Workshop. Towards Increasing the Annual National Productivity by One Tonne FFB/ha/year 3–14 (Malaysian Palm Oil Board, 2006)
Rohani, O. et al. in Advances in Oil Palm Research (eds Yusof, B., Jalani, B. S. & Chan, K. W.) 238–283 (Malaysian Palm Oil Board, 2000)
Adam, H. et al. Determination of flower structure in Elaeis guineensis: do palms use the same homeotic genes as other species? Ann. Bot. 100, 1–12 (2007)
Stewart, F. J. & Raleigh, E. A. Dependence of McrBC cleavage on distance between recognition elements. Biol. Chem. 379, 611–616 (1998)
Ordway, J. M. et al. Comprehensive DNA methylation profiling in a human cancer genome identifies novel epigenetic targets. Carcinogenesis 27, 2409–2423 (2006)
Chan, P. L. et al. Evaluation of reference genes for quantitative real-time PCR in oil palm elite planting materials propagated by tissue culture. PLoS ONE 9, e99774 (2014)
Jaligot, E. et al. Epigenetic imbalance and the floral developmental abnormality of the in vitro-regenerated oil palm Elaeis guineensis. Ann. Bot. 108, 1453–1462 (2011)
Acknowledgements
We acknowledge the contributions of staff members of the Breeding and Tissue Culture Unit at MPOB for creating the valuable clonal lines, and for their extensive data collection and sampling efforts. We thank Genomics Unit at MPOB for conducting DNA fingerprinting to verify clonal lines. We thank The McDonnell Genome Institute at Washington University for genomic bisulfite sequencing and transcriptome sequencing support, and MOgene for microarray hybridizations. At Orion Genomics, we thank N. Sander, J. Reed, J. Brune, K. Soe, J. McDonald, C. Brown and B. Dove for technical support, and M.-F. Wu and M. Sachdeva for assistance with the manuscript and additional informatics support. We would also like to thank T. Dalmay for recommendations on sRNA library construction. We appreciate the constant support of the Director-General of MPOB, Datuk Dr. Yuen-May Choo, and the Ministry of Plantation Industries and Commodities, Malaysia.
Author information
Authors and Affiliations
Contributions
M.O.-A. led the work on the MANTLED marker/gene. M.O.-A., R.Si., E.-T.L.L. and R.Sa. conceptualized the research programme. M.O.-A., R.Si., E.-T.L.L., R.N., N.L., S.W.S., J.M.O., R.Sa. and R.A.M. developed the overall strategy, designed experiments and coordinated the project. A.T.H., Z.I. and S.K.R. performed tissue culture on selected ortets and field-planted the ramets. Field data collection and fruit bunch census were conducted at various research stations by F.A.M., N.A.A.B., M.M., N.A., Z.Y. and M.D.A. M.O.-A., C.-C.L., X.A., C.-N.C., W.-C.W., S.S.R.S.A. and Y.-Y.K. identified samples for discovery and validation panels. M.O.-A., C.-C.L. and X.A. identified materials used in the mosaic experiments. M.O.-A., S.-E.O., S.-Y.K., N.S. and N.A. conducted laboratory experiments, histological staging of inflorescences and data analyses. N.J. and S.W.S performed microarray analyses. B.B. and M.A.B. prepared fractions for microarray hybridizations. B.B. designed and analysed qPCR experiments. B.B. and M.B. performed qPCR assays. A.V.B. designed and analysed clone-based bisulfite sequencing experiments, and A.V.B. and M.B. performed bisulfite sequencing assays. M.B. designed and performed qRT–PCR experiments. C.W. and J.M.O. analysed transcriptome data. K.-L.C., N.A., S.W.S., M.H., C.W. and A.V.B. provided bioinformatics support. M.O.-A., R.Si., E.-T.L.L., R.N., N.L., S.W.S., J.M.O., R.Sa. and R.A.M. prepared and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.A.M. is a former consultant of Orion Genomics, LLC.
Extended data figures and tables
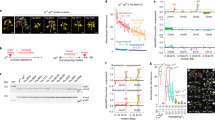
Extended Data Figure 1 Spikelets from clonal palms of different fruit form phenotypes.
a, Spikelets from a normal ramet. b, Spikelet from a fertile mantled ramet. c, Spikelet from a parthenocarpic mantled ramet. d, Spikelet from a revertant ramet displaying both normal (N) and mantled (M) fruits in the same spikelet.
Extended Data Figure 2 Annotation of genome-wide differentially methylated loci.
Sequences of microarray features reporting significant differential DNA methylation between fully normal and fully mantled leaf DNA samples of one or more clonal lineages (P < 0.05, two-sided Student’s t-test, Methods) were mapped to the reference E. guineensis pisifera genome14. Numbers of biological replicates per clonal lineage are provided in Extended Data Fig. 3. Features were assigned to gene and repeat classes according to annotations of genomic elements mapped within 3 kb of the microarray feature sequence, as this is the distance at which McrBC is capable of monitoring DNA methylation density. The repeat class includes all repetitive sequences, including transposons and pisifera-specific repetitive sequences14. Features mapping within 3 kb of both a gene and a repeat were assigned to both classes. The number of features reporting hypermethylation (red) and hypomethylation (green) are plotted.
Extended Data Figure 3 Summary of DNA methylation changes predicted by EWAS within clonal lineages.
Rows indicate independent clonal lineages from four oil palm industry sources (source A–D, as indicated in Fig. 1e). Clone lineages map to sources as follows: 7–9, source A; 1 and 2, source B; 3–6, source C; 10 and 11, source D. The numbers of fully normal and fully mantled palms per lineage represented are indicated to the left. Columns represent each microarray feature mapping to the EgDEF1 (open box at top) and upstream region. The relative positions of Rider, Karma and Koala elements are indicated. The arrow indicates the direction of EgDEF1 transcription. Features reporting significant hypomethylation or hypermethylation in mantled relative to normal clones are indicated as black and grey boxes, respectively. White boxes indicate features reporting no significant DNA methylation difference. Only clonal lineages including more than 1 ramet per phenotype are shown to determine statistical significance within each clonal lineage (n = 41 normal; n = 37 parthenocarpic mantled). Ramets from four additional clonal lineages were included in the source-by-source analysis shown in Fig. 1e.
Extended Data Figure 4 DNA methylation assays and supporting DNA methylation data.
a, Diagram of the EgDEF1 gene including the Karma element within intron 5 (orange box). Black boxes represent exons and the horizontal line represents introns. Scale bar is in base pair units. b, Blue tick marks represent the relative positions of the four microarray features reporting significant hypomethylation of mantled clones in all source lineages. The left-most feature includes the Karma splice acceptor site. Horizontal lines labelled B (BbvI) and R (RsaI) indicate the relative positions of amplicons used for qPCR-based CHG methylation assays. The BbvI amplicon also includes a ScrFI site (S) used in d. The relative position of the bisulfite sequencing amplicon used to determine Karma splice site CHG methylation is shown below the qPCR amplicons. c, Diagrams of the three alternatively spliced EgDEF1 transcripts. Black boxes represent exons included in each transcript. The dotted lines represent intronic sequences spliced out of the mature mRNA transcripts. The red box represents Karma element sequence spliced to EgDEF1 exon 5 in the kDEF1 transcript. The blue box represents EgDEF1 intron 5 sequence included in the tDEF1 transcript that does not use the exon 5 splice donor site. d, In addition to adult leaf samples analysed by BbvI and RsaI qPCR assays (Fig. 2c), 37 samples were found to have a SNP in the BbvI site and were therefore analysed by ScrFI and RsaI qPCR assays (Methods and Extended Data Fig. 4b). Linear discriminant analysis was performed between normal (n = 14) and mantled (n = 22 parthenocarpic mantled; n = 1 fertile mantled) samples. Combining these results with those shown in Fig. 2c, sensitivity and specificity for detection of mantling are each 94%. e, Bisulfite sequencing of controls, FN1 and FN2 (Fig. 2c). mCHG density was calculated for the three CHG sites covered by the unique common microarray feature (Figs 1e and 2d–g). FN1, FN2 and the mantled control were significantly hypomethylated relative to the normal control (*P < 0.0001, two-tailed Fisher’s exact test).
Extended Data Figure 5 Clone based bisulfite sequencing maps of normal and mantled phenotype fruits from epigenetic mosaics.
The heatmap format is as described in Fig. 2d–g. Grey boxes indicate a site in which a SNP on allele a results in a CHG to CHH site conversion. Mosaic clone 1 represents a revertant clone yielding 95% normal fruit. Mosaic clone 2 represents a revertant clone yielding 99% normal fruit. Alleles were analysed independently based on a SNP not affecting a potentially methylated base. Statistical analyses of methylation at the three CHG sites spanning the Karma splice site are shown in Fig. 3e, f.
Extended Data Figure 6 CHG methylation in rachis sectors of an oil palm yielding 7% normal fruit (clone lineage 2 in Fig. 3d).
Rachis of three successive fronds was dissected into 8 equal sectors. DNA methylation in each sector per frond was measured by BbvI and ScrFI assays, as described in Methods. Average DNA methylation density measurements of three technical replicates per frond, per sector, per assay are plotted on a radial graph representing the 8 rachis sections around the palm trunk (ScrFI assay, light blue; BbvI assay dark blue). Sector numbering was ratcheted for frond 2 versus 1, and frond 3 versus 2 based on the R2 best fit of CHG methylation density around the circumference of the palm to correct for out-of-register numbering of rachis sectors between successive fronds (data not shown). Consistent with the fact that this oil palm yields only 7% normal phenotype fruit, most DNA methylation measurements are consistent with the mantled phenotype. However, sectors 8 and 2 display gains of CHG methylation in rachis sectors of all three fronds, and reach or approach normal levels in sectors 8 and 2 of frond 2, thus demonstrating mosaicism directly.
Extended Data Figure 7 Protein sequences and summary of qRT–PCR assay designs.
a, Residues highlighted in red are encoded by Karma sequence splice to exon 5 of EgDEF1. The alternate splicing event disrupts the transcription activation domain of EgDEF1. Twelve variant amino acids are coded by Karma sequencing, followed by a stop codon. b, Diagram of EgDEF1 locus including positions of qRT–PCR primers. cDEF1 transcripts were detected using primer a (spanning the splice junction of exons 1 and 2) and primer c (internal to exon 7). kDEF1 transcripts were detected using primer b (spanning the splice junction of exons 4 and 5) and primer d (internal to Karma ORF2). tDEF1 transcripts were detected using primer a and primer e (spanning the 3′ end of exon 5 and including tDEF1-specific intron 5 sequence). c, All assays were confirmed to give a single band of the correct size by agarose gel electrophoresis. Amplicons were Sanger sequence verified. Note that no band is amplified using the kDEF1 primer pair in samples from normal inflorescence, consistent with lack of expression of kDEF1 in normal inflorescence. d, Sequences of primers diagrammed in b. e, f, Standard curves for qRT–PCR assays. PCR amplicons including each qRT–PCR amplified sequence were serially diluted and quantified in triplicate by qPCR using the indicated primer pairs. Dilutions (x axis) were plotted against the measured cycle threshold (y axis). e, Standard curves for cDEF1 (blue), kDEF1 (red) and tDEF1 (green). Line equations were used to calculate the efficiency of each primer pair. The efficiency of each primer pair was used in calculations for quantification of expression of each associated transcript. f, Standard curves for two endogenous oil palm control genes. The efficiency of each primer pair was used in calculations for quantification of expression of each associated transcript. Expression of each alternative transcript was calculated relative to the control PD00569 control. Control qRT–PCR primers are described previously39.
Extended Data Figure 8 Antisense 24-nucleotide siRNA analysis of inflorescence development.
siRNA expression at inflorescence stages 0 (shoot apical meristem), 2, 3, 4 and 5, was analysed by Illumina siRNA sequencing (Methods). FPKM normalized expression values for each measured 24-nucleotide siRNA are plotted in scale with the genomic elements diagrammed at the top of the figure. Grey bars indicate detected 24-nucleotide siRNAs that are not significantly differentially expressed between normal relative to mantled tissues (P > 0.05, Student’s t-test, two-tailed assuming equal variance). Differentially expressed 24-nucleotide siRNAs are plotted as green or red bars for normal or mantled tissues, respectively. Bars above and below the zero line represent sense and antisense siRNAs, respectively, and are plotted on the same scale in both directions.
Extended Data Figure 9 Relative abundance of 21- and 24-nucleotide sRNA in normal and mantled reclones and stage 0 inflorescence.
a, Distribution of sRNA lengths derived from mantled reclone (blue) and stage 0 inflorescence (red). b, Distribution of sRNA lengths derived from normal reclone (blue) and stage 0 inflorescence (red). Read lengths of sRNA sequencing reads are plotted as the percentage of total reads for each incremental sRNA nucleotide length.
Extended Data Figure 10 CHG methylation in recloned tissue cultures.
Tissue cultures were reconstituted from normal and mantled ramets from two clonal lineages (‘clones of clones’). Methylation at three CHG sites across the Karma DMR was quantified by qPCR assays at two (SC2) and seven (SC7) passages in tissue culture. Cultures derived from normal ramets displayed higher CHG methylation than those derived from mantled ramets. In both normal and mantled reclones, CHG methylation generally decreased with time in culture. At SC2, the time point at which 24-nucleotide siRNAs were measured (Fig. 4c), the culture from normal ramet lineage 1 had lost methylation at the BbvI (the site nearest the Karma splice acceptor site).
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1. (PDF 149 kb)
Rights and permissions
About this article
Cite this article
Ong-Abdullah, M., Ordway, J., Jiang, N. et al. Loss of Karma transposon methylation underlies the mantled somaclonal variant of oil palm. Nature 525, 533–537 (2015). https://doi.org/10.1038/nature15365
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15365
This article is cited by
-
Genome-wide methylome stability and parental effects in the worldwide distributed Lombardy poplar
BMC Biology (2024)
-
Genetic stability analysis of the temporary immersion bioreactors–derived sugarcane seedlings with simple sequence repeat (SSR) markers
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
-
DNA methylome analysis provides insights into gene regulatory mechanism for better performance of rice under fluctuating environmental conditions: epigenomics of adaptive plasticity
Planta (2024)
-
DNA methylation: an emerging paradigm of gene regulation under drought stress in plants
Molecular Biology Reports (2024)
-
Somatic embryogenesis in oil palm from immature leaves with emphasis on leaf position, sequential callus re-collection, use of temporary immersion system, and assessment of genetic and epigenetic fidelity of the resulting clones
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.