Abstract
Epigenetic modifiers have fundamental roles in defining unique cellular identity through the establishment and maintenance of lineage-specific chromatin and methylation status1. Several DNA modifications such as 5-hydroxymethylcytosine (5hmC) are catalysed by the ten eleven translocation (Tet) methylcytosine dioxygenase family members2, and the roles of Tet proteins in regulating chromatin architecture and gene transcription independently of DNA methylation have been gradually uncovered3. However, the regulation of immunity and inflammation by Tet proteins independent of their role in modulating DNA methylation remains largely unknown. Here we show that Tet2 selectively mediates active repression of interleukin-6 (IL-6) transcription during inflammation resolution in innate myeloid cells, including dendritic cells and macrophages. Loss of Tet2 resulted in the upregulation of several inflammatory mediators, including IL-6, at late phase during the response to lipopolysaccharide challenge. Tet2-deficient mice were more susceptible to endotoxin shock and dextran-sulfate-sodium-induced colitis, displaying a more severe inflammatory phenotype and increased IL-6 production compared to wild-type mice. IκBζ, an IL-6-specific transcription factor, mediated specific targeting of Tet2 to the Il6 promoter, further indicating opposite regulatory roles of IκBζ at initial and resolution phases of inflammation. For the repression mechanism, independent of DNA methylation and hydroxymethylation, Tet2 recruited Hdac2 and repressed transcription of Il6 via histone deacetylation. We provide mechanistic evidence for the gene-specific transcription repression activity of Tet2 via histone deacetylation and for the prevention of constant transcription activation at the chromatin level for resolving inflammation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chen, T. & Dent, S. Y. Chromatin modifiers and remodellers: regulators of cellular differentiation. Nature Rev. Genet. 15, 93–106 (2014)
Kohli, R. M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013)
Pastor, W. A., Aravind, L. & Rao, A. TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nature Rev. Mol. Cell Biol. 14, 341–356 (2013)
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009)
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010)
Szabo, S. J. et al. Distinct effects of T-bet in TH1 lineage commitment and IFN-γ production in CD4 and CD8 T cells. Science 295, 338–342 (2002)
Tlaskalová-Hogenová, H. et al. Involvement of innate immunity in the development of inflammatory and autoimmune diseases. Ann. NY Acad. Sci. 1051, 787–798 (2005)
Neurath, M. F. Cytokines in inflammatory bowel disease. Nature Rev. Immunol. 14, 329–342 (2014)
Ichiyama, K. et al. The methylcytosine dioxygenase tet2 promotes DNA demethylation and activation of cytokine gene expression in T cells. Immunity 42, 613–626 (2015)
Yamamoto, M. et al. Regulation of Toll/IL-1-receptor-mediated gene expression by the inducible nuclear protein IκBζ. Nature 430, 218–222 (2004)
Kayama, H. et al. Class-specific regulation of pro-inflammatory genes by MyD88 pathways and IκBζ. J. Biol. Chem. 283, 12468–12477 (2008)
Hildebrand, D. G. et al. IκBζ is a transcriptional key regulator of CCL2/MCP-1. J. Immunol. 190, 4812–4820 (2013)
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011)
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013)
Deplus, R. et al. TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J. 32, 645–655 (2013)
Foster, S. L., Hargreaves, D. C. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007)
Qiao, Y. et al. Synergistic activation of inflammatory cytokine genes by interferon-gamma-induced chromatin remodeling and toll-like receptor signaling. Immunity 39, 454–469 (2013)
Nicodeme, E. et al. Suppression of inflammation by a synthetic histone mimic. Nature 468, 1119–1123 (2010)
Inoue, S., Mai, A., Dyer, M. J. & Cohen, G. M. Inhibition of histone deacetylase class I but not class II is critical for the sensitization of leukemic cells to tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis. Cancer Res. 66, 6785–6792 (2006)
Chen, X. et al. Requirement for the histone deacetylase Hdac3 for the inflammatory gene expression program in macrophages. Proc. Natl Acad. Sci. USA 109, E2865–E2874 (2012)
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20, 11–24 (2011)
Rampal, R. et al. DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep 9, 1841–1855 (2014)
Liu, J. et al. Rhbdd3 controls autoimmunity by suppressing the production of IL-6 by dendritic cells via K27-linked ubiquitination of the regulator NEMO. Nature Immunol. 15, 612–622 (2014)
Hunter, C. A. & Jones, S. A. IL-6 as a keystone cytokine in health and disease. Nature Immunol. 16, 448–457 (2015)
Ma, F. et al. The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-γ. Nature Immunol. 12, 861–869 (2011)
Liu, Y. et al. Histone lysine methyltransferase Ezh1 promotes TLR-triggered inflammatory cytokine production by suppressing Tollip. J. Immunol. 194, 2838–2846 (2015)
Chen, W. et al. Induction of Siglec-G by RNA viruses inhibits the innate immune response by promoting RIG-I degradation. Cell 152, 467–478 (2013)
Xia, M. et al. Histone methyltransferase Ash1l suppresses interleukin-6 production and inflammatory autoimmune diseases by inducing the ubiquitin-editing enzyme A20. Immunity 39, 470–481 (2013)
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010)
Acknowledgements
This work was supported by the National Key Basic Research Program of China (2013CB530503) and the National Natural Science Foundation of China (31200654, 31390431, 81230074, 81123006).
Author information
Authors and Affiliations
Contributions
Q.Z., K.Z., Q.S. and Y.H. performed the experiments, analysed the data and contributed equally for the whole study. Y.G., X.L., D.Z., Y.L., C.W., X.Z., X.S., J.L., W.G. and N.L. provided reagents and performed experiments; RL.L. provided Tet2 KO mice. X.C. and Q.Z. analysed the data and wrote the manuscript. X.C. designed and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
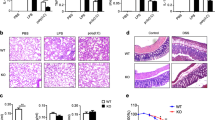
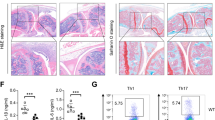
Extended Data Figure 1 Loss of Tet2 specifically enhances expression of IL-6 but not TNF-α.
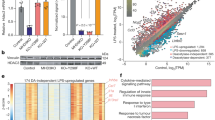
a, Hierarchical cluster of transcription level of chromatin modifiers with significant variation in LPS-stimulated murine BMDC, RPKMs were centred by the mean. b,Tet2 mRNA in human dendritic cells (hDC), LPS-stimulated murine BMDC, BMM and primary peritoneal macrophages (pMϕ). c–f, j, Human imDCs in which TET2 were lentivirally silenced (LV-siTET2), BMDC, BMM and peritoneal macrophages from Tet2-deficient mice (c–f), peritoneal macrophages from conditional Tet2-deficient mice (f, j) and their respective controls were stimulated by LPS for 8 h (c) or the indicated time (d, e, j). Protein levels of TET2 were detected by immunoblot, with lamin A/C as the loading control (c). IL-6 protein levels were analysed by ELISA (d), and Tnfa and Il6 mRNA levels were analysed by qPCR (e, j). Percentage of Iab+CD11c+BMDC or F4/80+ CD11b+peritoneal macrophages were analysed (f). g–i, Tet2 (g, h), Tet3 (g, i) were silenced in peritoneal macrophages for 48 h and then peritoneal macrophages were stimulated with LPS for 8 h (g, i) or the indicated time (h), and the protein levels of Tet2 or Tet3 were detected by immunoblot, with lamin A/C as the loading control (g).Tnfa and Il6 mRNA were analysed by qPCR (h, i). Full scans of blots are shown in Supplementary Fig. 1. Error bars represent s.e.m. of triplicate biological replicates and are representative of three independent experiments. **P < 0.01.
Extended Data Figure 2 Loss of Tet2 barely affects development of immune cells.
Percentages of the indicated immune cells were analysed by their specific lineage markers from control and Tet2-deficient mice. Data were from one representative of three independent experiments.
Extended Data Figure 3 Gene expression variation in Tet2-deficient dendritic cells.
a, b, mRNA variations of indicated genes in RNA-seq analysis of BMDC stimulated with LPS. RPKM of each of the genes in 4 h group were compared with 0 h group, and calculated to log2 ratio. c, d, qPCR analysis of mRNA levels of indicated genes in wild type and Tet2-deficient naive BMDC (c) or LPS-stimulated BMDC (d). e, BMDC were stimulated with LPS for 8 h, and then endogenous Tet2 enrichments at proximal promoter of indicated genes were analysed by ChIP-qPCR. Error bars represent s.d. of triplicate technical (e) or s.e.m. of triplicate biological (c, d) replicates and are representative of three independent experiments. **P < 0.01.
Extended Data Figure 4 Potential targets of Tet2 during immune response.
a–d, ELISA of cytokines in supernatants of splenic activated CD4+ T cells (a), CD8+ T cells (b) and NK cells (c), and conventional splenic CD19+ B cells (d) from Tet2-deficient (KO) and control mice (WT). T cells were stimulated with 10 μg ml−1 anti-CD3 and 1 μg ml−1 anti-CD28 monoclonal antibodies for 24 h. NK cells were stimulated with a combination of 10 ng ml−1 IL-12 plus 5 ng ml−1 IL-18 for 12 h. B cells were stimulated with 5 μg ml−1 LPS for 24 h, Error bars represent s.d. of triplicate technical replicates and are representative of three independent experiments. **P < 0.01.
Extended Data Figure 5 Loss of Tet2 barely affects TLR4 signalling pathways.
a, b, Immunoblot assays of the phosphorylated (p-) or total proteins in lysates of wild-type and Tet2-deficent BMDC (a) and peritoneal macrophages (b) stimulated with LPS for indicated time. Full scans of blots are shown in Supplementary Fig. 1. Data were from one representative of three independent experiments.
Extended Data Figure 6 Tet2 binding partners for cytokine regulation.
a, Co-immunoprecipitation assay using Tet2 antibody of nuclear fraction of murine BMDC stimulated with LPS for 8 h. Gene symbols of transcription factor deposited in KEGG database and their unique peptides identified by mass spectrometry analysis were tabled. b, IκBζ was silenced in BMDC using specific siRNA for 36 h and the BMDC were stimulated with LPS for 8 h. The protein levels of IκBζ were detected by immunoblot, with lamin A/C as the loading control. c, Hdac1/2 (blue circles) were subjected to interaction analysis with genes which had significant expression variations (red circles for upregulated, and green circles for downregulated) in BMDC 4 h after LPS stimulation. The linkers indicate interaction or regulation relationship between the two genes. d, e, Flag-tagged Hdac1 (d) or Hdac2 (e) were overexpressed in HEK293T cells together with Myc-tagged Tet2. Cell lysates were examined by IP and immunoblot with indicated antibodies. The whole-cell lysates (WCL) were used to examine the input of overexpressed proteins. f, BMDC were pretreated with 100 nM TSA for 1 h, and then stimulated by LPS for indicated time. mRNA levels of indicated genes were analysed by qPCR. g, h, BMDC (g) and peritoneal macrophages (h) were stimulated with LPS for the indicated time. Enrichments of H3Ac (g), H4Ac (h) at Tnf promoters were analysed by ChIP-qPCR. i, Hdac2 was silenced in BMDC for 48 h and then the BMDC were stimulated with LPS for 8 h. The protein levels of Hdac2 were detected by immunoblot, with lamin A/C as the loading control. Full scans of blots are shown in Supplementary Fig. 1. Error bars represent s.d. of triplicate technical (g, h) or s.e.m. of triplicate biological (f) replicates and are representative of three independent experiments. **P < 0.01.
Extended Data Figure 7 Tet2 represses Il6 promoter activity through recruiting Hdac2.
a, Luciferase activities in lysates of HEK293T cells transfected with indicated plasmids and luciferase reporter for murine Il6 promoter (TSS to upstream 3K). b, ChIP-qPCR analysis of Myc-tagged Tet3 in murine Il6 promoter in HEK293T cells which were transfected with indicated plasmids and luciferase reporter. c, ChIP-qPCR of Myc-tagged Tet2 or endogenous Hdac2 in murine Il6 promoter in HEK293T cells transfected with indicated plasmids and luciferase reporter. d, Hdac2 was silenced for 36 h in HEK293T cells. Protein levels of Hdac2 were detected by immunoblot, with lamin A/C as the loading control. e, Luciferase activities in lysates of Hdac2-silenced HEK293T cells transfected with indicated plasmids and luciferase reporter containing murine Il6 promoter. f, Wild-type (WT) and Tet2-deficient (KO) peritoneal macrophages were stimulated with LPS for 8 h. IP using O-GlcNAcylation antibody was performed. The whole-cell lysates (WCL) were used as the input. Full scans of blots are shown in Supplementary Fig. 1. Error bars represent s.d. of triplicate technical replicates and are representative of three independent experiments. **P < 0.01.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1, which shows the full scans of the blots and gels. (PDF 1032 kb)
Supplementary Data
This file contains Supplementary Table 1. (XLSX 18 kb)
Rights and permissions
About this article
Cite this article
Zhang, Q., Zhao, K., Shen, Q. et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 525, 389–393 (2015). https://doi.org/10.1038/nature15252
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15252
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.