Abstract
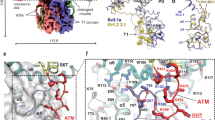
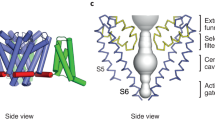
Na+-activated K+ channels are members of the Slo family of large conductance K+ channels that are widely expressed in the brain, where their opening regulates neuronal excitability. These channels fulfil a number of biological roles and have intriguing biophysical properties, including conductance levels that are ten times those of most other K+ channels and gating sensitivity to intracellular Na+. Here we present the structure of a complete Na+-activated K+ channel, chicken Slo2.2, in the Na+-free state, determined by cryo-electron microscopy at a nominal resolution of 4.5 ångströms. The channel is composed of a large cytoplasmic gating ring, in which resides the Na+-binding site and a transmembrane domain that closely resembles voltage-gated K+ channels. In the structure, the cytoplasmic domain adopts a closed conformation and the ion conduction pore is also closed. The structure reveals features that can explain the unusually high conductance of Slo channels and how contraction of the cytoplasmic gating ring closes the pore.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The 3D cryo-EM density maps of Slo2.2 with low-pass filter and amplitude modification have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-3062 (Slo2.2 whole channel), EMD-3063 (Slo2.2 gating ring) and EMD-3064 (Slo2.2 TMD). Atomic coordinates for the atomic model of full-length Slo2.2, Slo2.2 gating ring and Slo2.2 TMD have been deposited in the Protein Data Bank under accession numbers 5A6E, 5A6F and 5A6G, respectively.
References
Hille, B. Ionic Channels of Excitable Membranes 3rd edn 131–168 (Sinauer Associates, 2001)
Bader, C. R., Bernheim, L. & Bertrand, D. Sodium-activated potassium current in cultured avian neurones. Nature 317, 540–542 (1985)
Dryer, S. E., Fujii, J. T. & Martin, A. R. A Na+-activated K+ current in cultured brain stem neurones from chicks. J. Physiol. (Lond.) 410, 283–296 (1989)
Haimann, C. & Bader, C. R. Sodium-activated potassium channel in avian sensory neurons. Cell Biol. Int. Rep. 13, 1133–1139 (1989)
Kameyama, M. et al. Intracellular Na+ activates a K+ channel in mammalian cardiac cells. Nature 309, 354–356 (1984)
Schwindt, P. C., Spain, W. J. & Crill, W. E. Long-lasting reduction of excitability by a sodium-dependent potassium current in cat neocortical neurons. J. Neurophysiol. 61, 233–244 (1989)
Yan, Y., Yang, Y., Bian, S. & Sigworth, F. J. Expression, purification and functional reconstitution of slack sodium-activated potassium channels. J. Membr. Biol. 245, 667–674 (2012)
Yuan, A. et al. The sodium-activated potassium channel is encoded by a member of the Slo gene family. Neuron 37, 765–773 (2003)
Bhattacharjee, A., Gan, L. & Kaczmarek, L. K. Localization of the Slack potassium channel in the rat central nervous system. J. Comp. Neurol. 454, 241–254 (2002)
Kaczmarek, L. K. et al. Regulation of the timing of MNTB neurons by short-term and long-term modulation of potassium channels. Hear. Res. 206, 133–145 (2005)
Wallén, P. et al. Sodium-dependent potassium channels of a Slack-like subtype contribute to the slow afterhyperpolarization in lamprey spinal neurons. J. Physiol. (Lond.) 585, 75–90 (2007)
Yang, B., Desai, R. & Kaczmarek, L. K. Slack and Slick KNa channels regulate the accuracy of timing of auditory neurons. J. Neurosci. 27, 2617–2627 (2007)
Allen, A. S. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013)
Barcia, G. et al. De novo gain-of-function KCNT1 channel mutations cause malignant migrating partial seizures of infancy. Nature Genet. 44, 1255–1259 (2012)
Ishii, A. et al. A recurrent KCNT1 mutation in two sporadic cases with malignant migrating partial seizures in infancy. Gene 531, 467–471 (2013)
McTague, A. et al. Migrating partial seizures of infancy: expansion of the electroclinical, radiological and pathological disease spectrum. Brain 136, 1578–1591 (2013)
Vanderver, A. et al. Identification of a novel de novo p.Phe932Ile KCNT1 mutation in a patient with leukoencephalopathy and severe epilepsy. Pediatr. Neurol. 50, 112–114 (2014)
Heron, S. E. et al. Missense mutations in the sodium-gated potassium channel gene KCNT1 cause severe autosomal dominant nocturnal frontal lobe epilepsy. Nature Genet. 44, 1188–1190 (2012)
Martin, H. C. et al. Clinical whole-genome sequencing in severe early-onset epilepsy reveals new genes and improves molecular diagnosis. Hum. Mol. Genet. 23, 3200–3211 (2014)
Huang, F. et al. TMEM16C facilitates Na+-activated K+ currents in rat sensory neurons and regulates pain processing. Nature Neurosci. 16, 1284–1290 (2013)
Lu, R. et al. Slack channels expressed in sensory neurons control neuropathic pain in mice. J. Neurosci. 35, 1125–1135 (2015)
Lu, S., Das, P., Fadool, D. A. & Kaczmarek, L. K. The Slack sodium-activated potassium channel provides a major outward current in olfactory neurons of Kv1.3−/− super-smeller mice. J. Neurophysiol. 103, 3311–3319 (2010)
Paulais, M., Lachheb, S. & Teulon, J. A Na+- and Cl−-activated K+ channel in the thick ascending limb of mouse kidney. J. Gen. Physiol. 127, 205–215 (2006)
Wu, Y., Yang, Y., Ye, S. & Jiang, Y. Structure of the gating ring from the human large-conductance Ca2+-gated K+ channel. Nature 466, 393–397 (2010)
Yuan, P., Leonetti, M. D., Hsiung, Y. & MacKinnon, R. Open structure of the Ca2+ gating ring in the high-conductance Ca2+-activated K+ channel. Nature 481, 94–97 (2012)
Yuan, P., Leonetti, M. D., Pico, A. R., Hsiung, Y. & MacKinnon, R. Structure of the human BK channel Ca2+-activation apparatus at 3.0 Å resolution. Science 329, 182–186 (2010)
Garg, P., Gardner, A., Garg, V. & Sanguinetti, M. C. Structural basis of ion permeation gating in Slo2.1 K+ channels. J. Gen. Physiol. 142, 523–542 (2013)
Wilkens, C. M. & Aldrich, R. W. State-independent block of BK channels by an intracellular quaternary ammonium. J. Gen. Physiol. 128, 347–364 (2006)
Zhou, Y., Xia, X. M. & Lingle, C. J. Cysteine scanning and modification reveal major differences between BK channels and Kv channels in the inner pore region. Proc. Natl Acad. Sci. USA 108, 12161–12166 (2011)
Long, S. B., Tao, X., Campbell, E. B. & MacKinnon, R. Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature 450, 376–382 (2007)
Zhou, Y., Morais-Cabral, J. H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by a K+ channel–Fab complex at 2.0 Å resolution. Nature 414, 43–48 (2001)
Joiner, W. J. et al. Formation of intermediate-conductance calcium-activated potassium channels by interaction of Slack and Slo subunits. Nature Neurosci. 1, 462–469 (1998)
Zhang, X., Zeng, X. & Lingle, C. J. Slo3 K+ channels: voltage and pH dependence of macroscopic currents. J. Gen. Physiol. 128, 317–336 (2006)
Dai, L., Garg, V. & Sanguinetti, M. C. Activation of Slo2.1 channels by niflumic acid. J. Gen. Physiol. 135, 275–295 (2010)
Long, S. B., Campbell, E. B. & Mackinnon, R. Crystal structure of a mammalian voltage-dependent Shaker family K+ channel. Science 309, 897–903 (2005)
Tao, X., Avalos, J. L., Chen, J. & MacKinnon, R. Crystal structure of the eukaryotic strong inward-rectifier K+ channel Kir2.2 at 3.1 Å resolution. Science 326, 1668–1674 (2009)
Whorton, M. R. & MacKinnon, R. Crystal structure of the mammalian GIRK2 K+ channel and gating regulation by G proteins, PIP2, and sodium. Cell 147, 199–208 (2011)
Guo, D. & Lu, Z. Interaction mechanisms between polyamines and IRK1 inward rectifier K+ channels. J. Gen. Physiol. 122, 485–500 (2003)
Heginbotham, L. & MacKinnon, R. Conduction properties of the cloned Shaker K+ channel. Biophys. J. 65, 2089–2096 (1993)
Budelli, G., Geng, Y., Butler, A., Magleby, K. L. & Salkoff, L. Properties of Slo1 K+ channels with and without the gating ring. Proc. Natl Acad. Sci. USA 110, 16657–16662 (2013)
Leonetti, M. D., Yuan, P., Hsiung, Y. & Mackinnon, R. Functional and structural analysis of the human SLO3 pH- and voltage-gated K+ channel. Proc. Natl Acad. Sci. USA 109, 19274–19279 (2012)
Zhang, Z., Rosenhouse-Dantsker, A., Tang, Q. Y., Noskov, S. & Logothetis, D. E. The RCK2 domain uses a coordination site present in Kir channels to confer sodium sensitivity to Slo2.2 channels. J. Neurosci. 30, 7554–7562 (2010)
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002)
Thomson, S. J., Hansen, A. & Sanguinetti, M. C. Identification of the intracellular Na+ sensor in Slo2.1 potassium channels. J. Biol. Chem. 290, 14528–14535 (2015)
Yang, H. et al. Activation of Slo1 BK channels by Mg2+ coordinated between the voltage sensor and RCK1 domains. Nature Struct. Mol. Biol. 15, 1152–1159 (2008)
Doyle, D. A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998)
Webster, S. M., Del Camino, D., Dekker, J. P. & Yellen, G. Intracellular gate opening in Shaker K+ channels defined by high-affinity metal bridges. Nature 428, 864–868 (2004)
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Penczek, P. A. et al. CTER-rapid estimation of CTF parameters with error assessment. Ultramicroscopy 140, 9–19 (2014)
Yang, Z., Fang, J., Chittuluru, J., Asturias, F. J. & Penczek, P. A. Iterative stable alignment and clustering of 2D transmission electron microscope images. Structure 20, 237–247 (2012)
Hohn, M. et al. SPARX, a new environment for Cryo-EM image processing. J. Struct. Biol. 157, 47–55 (2007)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011)
Ho, B. K. & Gruswitz, F. HOLLOW: generating accurate representations of channel and interior surfaces in molecular structures. BMC Struct. Biol. 8, 49 (2008)
Dolinsky, T. J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013)
Schmidt, D., Jiang, Q. X. & MacKinnon, R. Phospholipids and the origin of cationic gating charges in voltage sensors. Nature 444, 775–779 (2006)
Miller, C. (ed.) Ion Channel Reconstitution (Plenum, 1986)
Ruta, V., Jiang, Y., Lee, A., Chen, J. & MacKinnon, R. Functional analysis of an archaebacterial voltage-dependent K+ channel. Nature 422, 180–185 (2003)
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015)
Acknowledgements
We thank Z. Yu and J. de la Cruz at the Howard Hughes Medical Institute Janelia Cryo-EM facility for assistance in data collection, S. Harrison and S. Jenni for assistance with Phenix refinement of cryo-EM density maps, and members of the MacKinnon laboratory for discussions. This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation grant number ACI-1053575. This work was supported in part by GM43949. R.K.H. is a Howard Hughes Medical Institute postdoctoral fellow of the Helen Hay Whitney Foundation and T.W. and R.M. are investigators of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
R.K.H. performed the experiments. P.Y. provided assistance with protein expression and purification. Z.L. aided with sample preparation and data collection. Y.H. provided assistance with protein expression. T.W. aided with initial model generation and map interpretation. R.K.H and R.M. designed the experiments and analysed the results. R.K.H. and R.M. prepared the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of Slo channels.
a, Sequence alignment of chicken Slo2.2 with human Slo2.2 and human Slo2.1. b, Predicted position of transmembrane helices in Slo2.2 S1–S4 domain on the basis of hydropothy analysis using Jpred 4 (ref. 72). c, d, Structure-based sequence alignment of chicken Slo2.2 TMD with rat Kv chimaera (c) and chicken Slo2.2 gating ring with human Slo1 gating ring (d). Helices are blue and β-strands are red.
Extended Data Figure 2 Full channel 3D reconstruction of chicken Slo2.2.
a, Representative micrograph of detergent- and lipid-solubilized Slo2.2 in vitreous ice. b, Selected 2D class averages. c, Ab initio model of Slo2.2. d, FSC curve of the full channel reconstruction with the nominal resolution estimated to be 4.5 Å on the basis of the FSC = 0.143 (dashed line) cut-off criterion.
Extended Data Figure 3 Focused refinement of the gating ring and the TMD.
a, 3D density map of the full channel reconstruction, coloured according to local resolution (in ångströms). b, c, 3D density map calculated following focused refinement using a mask to only include the gating ring (b) and the TMD (c), coloured according to local resolution (in ångströms). d, FSC of the full channel reconstruction (estimated resolution of 4.5 Å), the gating-ring-focused refinement reconstruction (4.2 Å) and the TMD-focused refinement reconstruction (5.2 Å).
Extended Data Figure 4 Validation of the Slo2.2 model.
a, Refinement statistics for the Slo2.2 full channel, TMD and gating-ring models. b, c, FSC curves for cross-validation of the refined gating ring (b) and TMD (c) models. The black curves are the refined model compared to the full data set, the red curves are the refined model compared to half map 1 (used during test refinement) and the blue curves are the refined model compared to half map 2 (not used during test refinement).
Extended Data Figure 5 K+ ions in Slo2.2.
a, Central section of the density maps of the two independently calculated half maps (coloured in green and red) with densities corresponding to K+ ions labelled. b, Superposition of the Slo2.2 selectivity filter (green) with KcsA (PDB code 1K4C) selectivity filter (yellow). Density peaks resolved in the Slo2.2 selectivity filter at 6.5 σ are shown as blue meshes. K+ ions resolved in KcsA are shown as grey spheres.
Extended Data Figure 6 Representative segments of the cryo-EM density map.
a–d, Selected regions of the gating-ring density (a, b) and the TMD density (c, d) maps with the refined model.
Extended Data Figure 7 Single channel conductance of Slo2.2.
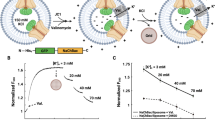
a, Single channel current–voltage relationship (mean ± s.e.m.) for Slo2.2 in planar lipid bilayers. Single channel conductance is about 200 pS. b, Representative recordings of Slo2.2 held at −80 mV, −40 mV and 0 mV in planar lipid bilayers. Chamber solution contained 135 mM NaCl and 15 mM KCl, and cup solution contained 150 mM KCl. c, Histogram of Slo2.2 currents when held at −80 mV, −40 mV and 0 mV, as labelled.
Extended Data Figure 8 Inner helix gate.
a, Ribbon diagram of the Slo2.2 pore with Met333 side chains modelled as spheres. b, Pore radius plot as a function of distance from the extracellular surface for Slo2.2 with Met333 modelled as each of the six most frequently observed rotamers, as labelled. For distances less than about 40 Å, the curves coincide.
Extended Data Figure 9 Slo2.2 gating ring is in a closed conformation.
Wire diagrams of Slo1 gating ring in the open (top left) and closed (top right) conformations. The mobile RCK1 N lobe is black and the rest of the gating ring is grey. The N-terminal residue of the gating ring, Lys343, is shown as a pink sphere. Wire diagram of the Slo2.2 gating ring (bottom) with the RCK1 N-lobe blue and the rest of the gating ring light blue. The N-terminal residue of the gating ring, Lys351, is shown as a pink sphere.
Rights and permissions
About this article
Cite this article
Hite, R., Yuan, P., Li, Z. et al. Cryo-electron microscopy structure of the Slo2.2 Na+-activated K+ channel. Nature 527, 198–203 (2015). https://doi.org/10.1038/nature14958
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14958
This article is cited by
-
Conformational plasticity of NaK2K and TREK2 potassium channel selectivity filters
Nature Communications (2023)
-
Interaction Between HCN and Slack Channels Regulates mPFC Pyramidal Cell Excitability in Working Memory Circuits
Molecular Neurobiology (2023)
-
Impaired motor skill learning and altered seizure susceptibility in mice with loss or gain of function of the Kcnt1 gene encoding Slack (KNa1.1) Na+-activated K+ channels
Scientific Reports (2020)
-
Deep Learning to Predict Protein Backbone Structure from High-Resolution Cryo-EM Density Maps
Scientific Reports (2020)
-
Ca2+-regulated Ca2+ channels with an RCK gating ring control plant symbiotic associations
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.