Abstract
The hypoxia-inducible factors (HIFs) coordinate cellular adaptations to low oxygen stress by regulating transcriptional programs in erythropoiesis, angiogenesis and metabolism. These programs promote the growth and progression of many tumours, making HIFs attractive anticancer targets. Transcriptionally active HIFs consist of HIF-α and ARNT (also called HIF-1β) subunits. Here we describe crystal structures for each of mouse HIF-2α–ARNT and HIF-1α–ARNT heterodimers in states that include bound small molecules and their hypoxia response element. A highly integrated quaternary architecture is shared by HIF-2α–ARNT and HIF-1α–ARNT, wherein ARNT spirals around the outside of each HIF-α subunit. Five distinct pockets are observed that permit small-molecule binding, including PAS domain encapsulated sites and an interfacial cavity formed through subunit heterodimerization. The DNA-reading head rotates, extends and cooperates with a distal PAS domain to bind hypoxia response elements. HIF-α mutations linked to human cancers map to sensitive sites that establish DNA binding and the stability of PAS domains and pockets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Semenza, G. L. Regulation of mammalian O2 homeostasis by hypoxia-inducible factor 1. Annu. Rev. Cell Dev. Biol. 15, 551–578 (1999)
Semenza, G. L. Hypoxia-inducible factors in physiology and medicine. Cell 148, 399–408 (2012)
Wang, G. L., Jiang, B. H., Rue, E. A. & Semenza, G. L. Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc. Natl Acad. Sci. USA 92, 5510–5514 (1995)
Jiang, B. H., Rue, E., Wang, G. L., Roe, R. & Semenza, G. L. Dimerization, DNA binding, and transactivation properties of hypoxia-inducible factor 1. J. Biol. Chem. 271, 17771–17778 (1996)
Peng, J., Zhang, L., Drysdale, L. & Fong, G. H. The transcription factor EPAS-1/hypoxia-inducible factor 2alpha plays an important role in vascular remodeling. Proc. Natl Acad. Sci. USA 97, 8386–8391 (2000)
Bersten, D. C., Sullivan, A. E., Peet, D. J. & Whitelaw, M. L. bHLH-PAS proteins in cancer. Nature Rev. Cancer 13, 827–841 (2013)
Kewley, R. J., Whitelaw, M. L. & Chapman-Smith, A. The mammalian basic helix-loop-helix/PAS family of transcriptional regulators. Int. J. Biochem. Cell Biol. 36, 189–204 (2004)
McIntosh, B. E., Hogenesch, J. B. & Bradfield, C. A. Mammalian Per-Arnt-Sim proteins in environmental adaptation. Annu. Rev. Physiol. 72, 625–645 (2010)
Keith, B., Johnson, R. S. & Simon, M. C. HIF1α and HIF2α: sibling rivalry in hypoxic tumour growth and progression. Nature Rev. Cancer 12, 9–22 (2012)
Heikkilä, M., Pasanen, A., Kivirikko, K. I. & Myllyharju, J. Roles of the human hypoxia-inducible factor (HIF)-3alpha variants in the hypoxia response. Cell. Mol. Life Sci. 68, 3885–3901 (2011)
Harris, A. L. Hypoxia—a key regulatory factor in tumour growth. Nature Rev. Cancer 2, 38–47 (2002)
Bruick, R. K. & McKnight, S. L. A conserved family of prolyl-4-hydroxylases that modify HIF. Science 294, 1337–1340 (2001)
Lando, D., Peet, D. J., Whelan, D. A., Gorman, J. J. & Whitelaw, M. L. Asparagine hydroxylation of the HIF transactivation domain a hypoxic switch. Science 295, 858–861 (2002)
Dames, S. A., Martinez-Yamout, M., De Guzman, R. N., Dyson, H. J. & Wright, P. E. Structural basis for Hif-1α/CBP recognition in the cellular hypoxic response. Proc. Natl Acad. Sci. USA 99, 5271–5276 (2002)
Semenza, G. L. HIF-1 mediates metabolic responses to intratumoral hypoxia and oncogenic mutations. J. Clin. Invest. 123, 3664–3671 (2013)
Semenza, G. L. Hypoxia-inducible factor 1 and cardiovascular disease. Annu. Rev. Physiol. 76, 39–56 (2014)
Girgis, C. M., Cheng, K., Scott, C. H. & Gunton, J. E. Novel links between HIFs, type 2 diabetes, and metabolic syndrome. Trends Endocrinol. Metab. 23, 372–380 (2012)
Eltzschig, H. K., Bratton, D. L. & Colgan, S. P. Targeting hypoxia signalling for the treatment of ischaemic and inflammatory diseases. Nature Rev. Drug Discov. 13, 852–869 (2014)
Semenza, G. L. Defining the role of hypoxia-inducible factor 1 in cancer biology and therapeutics. Oncogene 29, 625–634 (2010)
Semenza, G. L. Hypoxia-inducible factors: mediators of cancer progression and targets for cancer therapy. Trends Pharmacol. Sci. 33, 207–214 (2012)
Hewitson, K. S. & Schofield, C. J. The HIF pathway as a therapeutic target. Drug Discov. Today 9, 704–711 (2004)
Huang, N. et al. Crystal structure of the heterodimeric CLOCK:BMAL1 transcriptional activator complex. Science 337, 189–194 (2012)
Wenger, R. H., Stiehl, D. P. & Camenisch, G. Integration of oxygen signaling at the consensus HRE. Sci. STKE 2005, re12 (2005)
Murray, I. A., Patterson, A. D. & Perdew, G. H. Aryl hydrocarbon receptor ligands in cancer: friend and foe. Nature Rev. Cancer 14, 801–814 (2014)
Dioum, E. M. et al. NPAS2: a gas-responsive transcription factor. Science 298, 2385–2387 (2002)
Erbel, P. J., Card, P. B., Karakuzu, O., Bruick, R. K. & Gardner, K. H. Structural basis for PAS domain heterodimerization in the basic helix–loop–helix-PAS transcription factor hypoxia-inducible factor. Proc. Natl Acad. Sci. USA 100, 15504–15509 (2003)
Scheuermann, T. H. et al. Artificial ligand binding within the HIF2α PAS-B domain of the HIF2 transcription factor. Proc. Natl Acad. Sci. USA 106, 450–455 (2009)
Guo, Y., Scheuermann, T. H., Partch, C. L., Tomchick, D. R. & Gardner, K. H. Coiled-coil coactivators play a structural role mediating interactions in hypoxia inducible factor heterodimerization. J. Biol. Chem. 290, 7707–7721 (2015)
Denison, M. S., Soshilov, A. A., He, G., DeGroot, D. E. & Zhao, B. Exactly the same but different: promiscuity and diversity in the molecular mechanisms of action of the aryl hydrocarbon (dioxin) receptor. Toxicol. Sci. 124, 1–22 (2011)
Möglich, A., Ayers, R. A. & Moffat, K. Structure and signaling mechanism of Per-ARNT-Sim domains. Structure 17, 1282–1294 (2009)
Henry, J. T. & Crosson, S. Ligand-binding PAS domains in a genomic, cellular, and structural context. Annu. Rev. Microbiol. 65, 261–286 (2011)
Cardoso, R. et al. Identification of Cys255 in HIF-1α as a novel site for development of covalent inhibitors of HIF-1α/ARNT PasB domain protein-protein interaction. Protein Sci. 21, 1885–1896 (2012)
Rogers, J. L. et al. Development of inhibitors of the PAS-B domain of the HIF-2α transcription factor. J. Med. Chem. 56, 1739–1747 (2013)
Scheuermann, T. H. et al. Allosteric inhibition of hypoxia inducible factor-2 with small molecules. Nature Chem. Biol. 9, 271–276 (2013)
Miranda, E. et al. A cyclic peptide inhibitor of HIF-1 heterodimerization that inhibits hypoxia signaling in cancer cells. J. Am. Chem. Soc. 135, 10418–10425 (2013)
Guo, Y. et al. Regulating the ARNT/TACC3 axis: multiple approaches to manipulating protein/protein interactions with small molecules. ACS Chem. Biol. 8, 626–635 (2013)
Key, J., Scheuermann, T. H., Anderson, P. C., Daggett, V. & Gardner, K. H. Principles of ligand binding within a completely buried cavity in HIF2alpha PAS-B. J. Am. Chem. Soc. 131, 17647–17654 (2009)
Lee, K. et al. Acriflavine inhibits HIF-1 dimerization, tumor growth, and vascularization. Proc. Natl Acad. Sci. USA 106, 17910–17915 (2009)
Wang, Z., Wu, Y., Li, L. & Su, X. D. Intermolecular recognition revealed by the complex structure of human CLOCK-BMAL1 basic helix-loop-helix domains with E-box DNA. Cell Res. 23, 213–224 (2013)
Latif, F. et al. Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 260, 1317–1320 (1993)
Maxwell, P. H. et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399, 271–275 (1999)
Li, L. et al. Hypoxia-inducible factor linked to differential kidney cancer risk seen with type 2A and type 2B VHL mutations. Mol. Cell. Biol. 27, 5381–5392 (2007)
Kaelin, W. G., Jr Molecular basis of the VHL hereditary cancer syndrome. Nature Rev. Cancer 2, 673–682 (2002)
Shen, C. et al. Genetic and functional studies implicate HIF1alpha as a 14q kidney cancer suppressor gene. Cancer Discov. 1, 222–235 (2011)
Kroeger, N. et al. Deletions of chromosomes 3p and 14q molecularly subclassify clear cell renal cell carcinoma. Cancer 119, 1547–1554 (2013)
Ollerenshaw, M., Page, T., Hammonds, J. & Demaine, A. Polymorphisms in the hypoxia inducible factor-1α gene (HIF1A) are associated with the renal cell carcinoma phenotype. Cancer Genet. Cytogenet. 153, 122–126 (2004)
Morris, M. R. et al. Mutation analysis of hypoxia-inducible factors HIF1A and HIF2A in renal cell carcinoma. Anticancer Res. 29, 4337–4343 (2009)
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–D811 (2015)
To, K. K., Sedelnikova, O. A., Samons, M., Bonner, W. M. & Huang, L. E. The phosphorylation status of PAS-B distinguishes HIF-1alpha from HIF-2α in NBS1 repression. EMBO J. 25, 4784–4794 (2006)
Kalousi, A. et al. Casein kinase 1 regulates human hypoxia-inducible factor HIF-1. J. Cell Sci. 123, 2976–2986 (2010)
Wu, D., Potluri, N., Kim, Y. & Rastinejad, F. Structure and dimerization properties of the aryl hydrocarbon receptor PAS-A domain. Mol. Cell. Biol. 33, 4346–4356 (2013)
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. D 62, 859–866 (2006)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Joosten, R. P., Long, F., Murshudov, G. N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 1, 213–220 (2014)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Dundas, J. et al. CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res. 34, W116–W118 (2006)
Abaan, O. D. et al. The exomes of the NCI-60 panel: a genomic resource for cancer biology and systems pharmacology. Cancer Res. 73, 4372–4382 (2013)
Guichard, C. et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nature Genet. 44, 694–698 (2012)
Sato, Y. et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nature Genet. 45, 860–867 (2013)
Seo, J. S. et al. The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res. 22, 2109–2119 (2012)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014)
Acknowledgements
We thank Y. Zhang and M. Wang for isolation and identification of trypaflavin.
Author information
Authors and Affiliations
Contributions
D.W. and F.R. conceived the study; D.W. isolated the proteins, carried out crystallizations and conducted biochemical studies; N.P. produced the expression and mutation constructs; Y.K. and J.L. collected synchrotron diffraction data; D.W., Y.K. and F.R. analysed the data; F.R. and D.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 The mammalian bHLH-PAS family of transcription factors.
a, Diagram showing the partner choices for heterodimerization between members of this family. ARNT is ubiquitously expressed, while ARNT2 (paralogue of ARNT) is mainly expressed in the central nervous system6,7. b, Sequence identity comparison among the same domains from mouse ARNT, HIF-1α, HIF-2α, HIF-3α, CLOCK and BMAL1 proteins.
Extended Data Figure 2 Structure of the HIF-2α–ARNT complex in cartoon mode.
Overall structure of HIF-2α–ARNT complex (centre), and individual domains of ARNT (top) and HIF-2α (bottom) with their secondary structures labelled.
Extended Data Figure 3 Comparison of the overall structures of HIF-2α–ARNT and CLOCK–BMAL1 complexes.
Two complexes are superposed by aligning the bHLH domains (a) or PAS-B domains (b) of ARNT and BMAL1, respectively. Colours used in labels match those used in figures for the same components.
Extended Data Figure 4 Domain interfaces of the HIF-2α–ARNT complex and the comparison with other structures.
a, Overall structure of HIF-2α–ARNT complex with domain interfaces numbered (left), and the spatial arrangement of each interface (right). b, The dimer interfaces formed by equivalent domains in the CLOCK–BMAL1 complex, corresponding to the interfaces 1, 2 and 4 of HIF-2α–ARNT heterodimer. c, The β-sheet-mediated antiparallel interface of isolated HIF-2α–ARNT PAS-B complex with mutations. d, The crystal structure showing the interaction between ARNT and TACC3 (left), and its superimpositions with HIF-2α–ARNT complex through the PAS-B domain of ARNT in two views (right). Colours used in labels match those used in figures for the same components. The binding position of TACC3 peptide with respect to ARNT PAS-B cannot be fully accommodated in our quaternary structure, as some steric clashes would result involving TACC3 and the HIF-2α PAS-B domain.
Extended Data Figure 5 Ligand binding sites in the HIF-2α–ARNT complex.
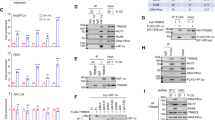
a, Acriflavine is a mixture of proflavine and trypaflavin. b, Binding tests of acriflavine, proflavine and trypaflavin to protein complexes of HIF-2α–ARNT (left) and HIF-1α–ARNT (right). The calculated Kd values for acriflavine, proflavine and trypaflavin were 41 nM, 31 nM and 34 nM for HIF-2α–ARNT, and 40 nM, 29 nM and 56 nM for HIF-1α–ARNT, respectively. Representative data of at least two experiments, shown as mean ± s.d. from three technical replicates. c, A proposed mechanism by which 0X3 binding can destabilize the HIF-2α–ARNT heterodimer. Shown is the proximity of 0X3 within the PAS-B domain of HIF-2α and R366 from the PAS-B domain of ARNT. 0X3 binding can potentially influence domain–domain interactions mediated by R366. We showed in Fig. 2b that R366 is a highly sensitive site for maintaining the stability of HIF-2α–ARNT heterodimer. 0X3 binding to the PAS-B domain of HIF-2α could further influence interfaces 3, 4 and 5 (shown in Extended Data Fig. 4a). d–g, 0X3 (d) binds at the inner pocket of HIF-2α PAS-B domain, while proflavine (f) binds at the interfacial pocket formed by ARNT PAS-A and HIF-2α PAS-B domains; corresponding positions in the structures of HIF-1α PAS-B (e) and HIF-1α–ARNT–DNA complex (g) are also shown. The yellow dotted lines represent hydrogen bonds. h, Pocket volumes calculated with CASTp program58 at default 1.4 Å probe radius.
Extended Data Figure 6 Interactions between the HIF-2α–ARNT complex and DNA.
a, Recognition of the HRE site by the bHLH domains. Overall structure of HIF-2α–ARNT–DNA with each domain labelled (left); and detailed interactions between the core HRE site (blue) and the bHLH domains of HIF-2α (upper right) or ARNT (lower right) are shown, with grey meshes showing 2Fo − Fc electron density contoured at 0.8σ. Hydrogen bonds (2.5–3.5 Å) are indicated by the brown dotted lines, while hydrophobic contact (3.6 Å) is shown by the green one. b, Schematic recognition diagram of HIF-2α–ARNT to the HRE sequence (blue) on DNA. The brown and green dotted lines represent hydrogen bonds and hydrophobic contact, respectively. The black arrow indicates the nucleotide interacting with residues N184 and K186 from HIF-2α PAS-A domain. Additional basic residues from HIF-2α and ARNT bHLH domains (HIF-2α K16, K18, R20, R24, R26 and ARNT R91, R99, R101) that can interact with DNA (mainly through the phosphate backbone) are also labelled in Extended Data Fig. 9. c, HRE DNA binding assay of HIF-2α–ARNT protein complex in wild type (WT) or point-mutated forms using fluorescence polarization. Representative data of at least two experiments are shown as mean ± s.d. from three technical replicates. Calculated approximate Kd values are shown in parentheses. d, The interactions between HIF-2α PAS-A domain and DNA, with the GH loop (including residues N184 and K186) and interacting nucleotides meshed by 2Fo − Fc map at 0.8σ.
Extended Data Figure 7 Locations of cancer-related missense mutations on the HIF-α–ARNT heterodimers.
a, Detailed information of the selected cancer-related mutations in HIF-2α and HIF-1α. The information about tissue and histology was adopted from the COSMIC database48 and other publications45,59,60,61,62. b, c, Spatial distribution of the HIF-2α (b) and HIF-1α (c) mutations in the heterodimers. Arrows point to close-up views of several regions in each heterodimer. The sequence positions of these mutations are also labelled in Extended Data Fig. 9.
Extended Data Figure 8 Spatial positions of phosphorylation sites at the HIF-α PAS-B domains in the context of HIF-α–ARNT complexes.
The phosphorylation sites T324 of HIF-2α (a) and S247 of HIF-1α (b) are shown as yellow sticks. Their corresponding residues T322 of HIF-1α and S249 of HIF-2α are also shown. Colours used in labels match those used in figures for the same components. HIF-2α is phosphorylated at residue T324 by protein kinase D1, but the equivalent residue in HIF-1α (T322) cannot be phosphorylated49. The differential positioning of their threonine residues next to a non-conserved loop between PAS-A and PAS-B domains may explain why they cannot be similarly phosphorylated. Casein kinase 1 (CK1) can phosphorylate HIF-1α at S247 (ref. 50). We predict that the equivalent residue S249 in HIF-2α may also be the target of CK1 (ref. 9), since the local environments for these residues are indistinguishable, with both residues being solvent accessible.
Extended Data Figure 9 Comparison of mouse ARNT, BMAL1, CLOCK, HIF-1α and HIF-2α proteins.
Sequence alignment of these five proteins includes the bHLH, PAS-A and PAS-B domains. Alignment was conducted by ClustalW263 and then processed by ESPript 3.064. The secondary structures of each domain of HIF-2α–ARNT complex, and residues involved in domain interfaces, protein–DNA or protein–compound interactions, are differently labelled above (for ARNT) or below (for HIF-2α) the sequences. In addition, the HIF-2α (magenta) or HIF-1α (blue) residues with cancer-related mutations (mainly selected from the COSMIC database48) and phosphorylation sites at the PAS-B domains are also labelled, further below the secondary structures of HIF-2α.
Supplementary information
Supplementary Information
This file contains uncropped scans with size marker indications for Figure 2b. (PDF 155 kb)
Rights and permissions
About this article
Cite this article
Wu, D., Potluri, N., Lu, J. et al. Structural integration in hypoxia-inducible factors. Nature 524, 303–308 (2015). https://doi.org/10.1038/nature14883
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14883
This article is cited by
-
Targeting hypoxia-inducible factors: therapeutic opportunities and challenges
Nature Reviews Drug Discovery (2024)
-
Design of hypoxia responsive CRISPR-Cas9 for target gene regulation
Scientific Reports (2023)
-
Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions
Signal Transduction and Targeted Therapy (2023)
-
Hypoxia-induced signaling in the cardiovascular system: pathogenesis and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Hypoxia signaling in human health and diseases: implications and prospects for therapeutics
Signal Transduction and Targeted Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.