Abstract
The E3 ubiquitin ligase PARKIN (encoded by PARK2) and the protein kinase PINK1 (encoded by PARK6) are mutated in autosomal-recessive juvenile Parkinsonism (AR-JP) and work together in the disposal of damaged mitochondria by mitophagy1,2,3. PINK1 is stabilized on the outside of depolarized mitochondria and phosphorylates polyubiquitin4,5,6,7,8 as well as the PARKIN ubiquitin-like (Ubl) domain9,10. These phosphorylation events lead to PARKIN recruitment to mitochondria, and activation by an unknown allosteric mechanism4,5,6,7,8,9,10,11,12. Here we present the crystal structure of Pediculus humanus PARKIN in complex with Ser65-phosphorylated ubiquitin (phosphoUb), revealing the molecular basis for PARKIN recruitment and activation. The phosphoUb binding site on PARKIN comprises a conserved phosphate pocket and harbours residues mutated in patients with AR-JP. PhosphoUb binding leads to straightening of a helix in the RING1 domain, and the resulting conformational changes release the Ubl domain from the PARKIN core; this activates PARKIN. Moreover, phosphoUb-mediated Ubl release enhances Ubl phosphorylation by PINK1, leading to conformational changes within the Ubl domain and stabilization of an open, active conformation of PARKIN. We redefine the role of the Ubl domain not only as an inhibitory13 but also as an activating element that is restrained in inactive PARKIN and released by phosphoUb. Our work opens up new avenues to identify small-molecule PARKIN activators.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
21 August 2015
The final figures, corresponding to the PDF, were updated online.
References
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nature Rev. Mol. Cell Biol. 12, 9–14 (2011)
Corti, O., Lesage, S. & Brice, A. What genetics tells us about the causes and mechanisms of Parkinson’s disease. Physiol. Rev. 91, 1161–1218 (2011)
Corti, O. & Brice, A. Mitochondrial quality control turns out to be the principal suspect in parkin and PINK1-related autosomal recessive Parkinson’s disease. Curr. Opin. Neurobiol. 23, 100–108 (2013)
Koyano, F. et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510, 162–166 (2014)
Kane, L. A. et al. PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J. Cell Biol. 205, 143–153 (2014)
Kazlauskaite, A. et al. Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem. J. 460, 127–139 (2014)
Ordureau, A. et al. Quantitative proteomics reveal a feedforward mechanism for mitochondrial PARKIN translocation and ubiquitin chain synthesis. Mol. Cell 56, 360–375 (2014)
Wauer, T. et al. Ubiquitin Ser65 phosphorylation affects ubiquitin structure, chain assembly and hydrolysis. EMBO J. 34, 307–325 (2015)
Kondapalli, C. et al. PINK1 is activated by mitochondrial membrane potential depolarization and stimulates Parkin E3 ligase activity by phosphorylating serine 65. Open Biol. 2, 120080 (2012)
Shiba-Fukushima, K. et al. PINK1-mediated phosphorylation of the Parkin ubiquitin-like domain primes mitochondrial translocation of Parkin and regulates mitophagy. Sci. Rep. 2, 1002 (2012)
Shiba-Fukushima, K. et al. Phosphorylation of mitochondrial polyubiquitin by PINK1 promotes Parkin mitochondrial tethering. PLoS Genet. 10, e1004861 (2014)
Okatsu, K. et al. Phosphorylated ubiquitin chain is the genuine Parkin receptor. J. Cell Biol. 209, 111–128 (2015)
Chaugule, V. K. et al. Autoregulation of Parkin activity through its ubiquitin-like domain. EMBO J. 30, 2853–2867 (2011)
Wauer, T. & Komander, D. Structure of the human Parkin ligase domain in an autoinhibited state. EMBO J. 32, 2099–2112 (2013)
Riley, B. E. et al. Structure and function of Parkin E3 ubiquitin ligase reveals aspects of RING and HECT ligases. Nature Commun. 4, 1982 (2013)
Trempe, J.-F. et al. Structure of parkin reveals mechanisms for ubiquitin ligase activation. Science 340, 1451–1455 (2013)
Sarraf, S. A. et al. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496, 372–376 (2013)
Borodovsky, A. et al. Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme family. Chem. Biol. 9, 1149–1159 (2002)
Wang, T. et al. Evidence for bidentate substrate binding as the basis for the K48 linkage specificity of otubain 1. J. Mol. Biol. 386, 1011–1023 (2009)
Macedo, M. G. et al. Genotypic and phenotypic characteristics of Dutch patients with early onset Parkinson's disease. Mov. Disord. 24, 196–203 (2009)
Veeriah, S. et al. Somatic mutations of the Parkinson’s disease-associated gene PARK2 in glioblastoma and other human malignancies. Nature Genet. 42, 77–82 (2010)
Zheng, X. & Hunter, T. Parkin mitochondrial translocation is achieved through a novel catalytic activity coupled mechanism. Cell Res. 23, 886–897 (2013)
Duda, D. M. et al. Structure of HHARI, a RING-IBR-RING ubiquitin ligase: autoinhibition of an ariadne-family E3 and insights into ligation mechanism. Structure 21, 1030–1041 (2013)
Caulfield, T. R. et al. Phosphorylation by PINK1 releases the UBL domain and initializes the conformational opening of the E3 ubiquitin ligase Parkin. PLOS Comput. Biol. 10, e1003935 (2014)
Ordureau, A. et al. Defining roles of PARKIN and ubiquitin phosphorylation by PINK1 in mitochondrial quality control using a ubiquitin replacement strategy. Proc. Natl Acad. Sci. USA 112, 6637–6642 (2015)
Kazlauskaite, A. & Muqit, M. M. K. PINK1 and Parkin — mitochondrial interplay between phosphorylation and ubiquitylation in Parkinson’s disease. FEBS J. 282, 215–223 (2015)
Pickrell, A. M. & Youle, R. J. The roles of PINK1, Parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85, 257–273 (2015)
Koyano, F. & Matsuda, N. Molecular mechanisms underlying PINK1 and Parkin catalyzed ubiquitylation of substrates on damaged mitochondria. Biochim. Biophys. Acta. http://dx.doi.org/10.1016/j.bbamcr.2015.02.009 (2015)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Vijay-Kumar, S., Bugg, C. E. & Cook, W. J. Structure of ubiquitin refined at 1.8 Å resolution. J. Mol. Biol. 194, 531–544 (1987)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011)
Favier, A. & Brutscher, B. Recovering lost magnetization: polarization enhancement in biomolecular NMR. J. Biomol. NMR 49, 9–15 (2011)
Kazimierczuk, K. & Orekhov, V. Y. Accelerated NMR spectroscopy by using compressed sensing. Angew. Chem. Int. Edn Engl. 50, 5556–5559 (2011)
Dou, H., Buetow, L., Sibbet, G. J., Cameron, K. & Huang, D. T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nature Struct. Mol. Biol. 19, 876–883 (2012)
Sakata, E. et al. Parkin binds the Rpn10 subunit of 26S proteasomes through its ubiquitin-like domain. EMBO Rep. 4, 301–306 (2003)
Kelsall, I. R. et al. TRIAD1 and HHARI bind to and are activated by distinct neddylated Cullin-RING ligase complexes. EMBO J. 32, 2848–2860 (2013)
Acknowledgements
We would like to thank M. Yu and beamline staff at Diamond Light Source, beamline I-04 and I-04-1, S. Freund and J. Wagstaff for NMR data, C. Johnson and S. McLaughlin for help with biophysics, BostonBiochem for providing UbVs and UbVME, C. Gladkova for help with cloning, and N. Birsa and J. Kittler (UCL London) for providing YFP–HsPARKIN plasmids. We thank members of the D.K. laboratory for reagents and discussions, and D. Barford, J. Pruneda and P. Elliott for comments on the manuscript. This work was supported by the Medical Research Council (U105192732), the European Research Council (309756), the Lister Institute for Preventive Medicine, the EMBO Young Investigator Program (to D.K.), and an EMBO Long-term Fellowship (to M.S.).
Author information
Authors and Affiliations
Contributions
T.W. and D.K. designed the research, and T.W. performed all experiments. M.S. performed cell-based studies. A.S. contributed to characterization of Ubl and ubiquitin phosphorylation. T.W. and D.K. analysed the data and wrote the manuscript with help from all authors.
Corresponding author
Ethics declarations
Competing interests
D.K. is part of the DUB Alliance that includes Cancer Research Technology and FORMA Therapeutics, and is a consultant for FORMA Therapeutics.
Extended data figures and tables
Extended Data Figure 1 Autoinhibited PARKIN and phosphoUb probes.
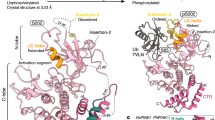
a, Structure of RnPARKIN (PDB 4K95 (ref. 16), chain A is used for all representations of RnPARKIN) with Ubl domain coloured in green, the unique PARKIN domain (UPD) in dark blue, RING1 in blue, IBR in light blue, REP in red, and RING2 in cyan. Zinc atoms are shown as grey spheres. The catalytic Cys431 is shown in ball-and-stick representation. Disordered linkers are indicated as dotted lines. The three mechanisms of PARKIN inhibition indicated in red. b, Schematic diagram of the closed, autoinhibited conformation of full-length PARKIN. c, Superposition of the E2–Ub complex from the BIRC7 structure (PDB 4AUQ (ref. 35)), superposed via its RING domain onto RING1 of full-length RnPARKIN (PDB 4K95 (ref. 16)) to indicate the position of the E2–Ub on PARKIN RING1. Assuming that E2–Ub adopts a canonical conformation on RING1, the E2 would clash with the Ubl domain and partially with the REP, while the E2-linked ubiquitin would clash with the REP. Hence, Ubl and REP have to be released to enable PARKIN E2–Ub binding at RING1. d, Time-course analysis of an exemplary reaction of UbC3Br (0.2 mg ml−1) phosphorylated by GST–PhPINK1 (5 µM) as described previously for ubiquitin8. Phosphorylation of proteins was monitored by a band shift on Coomassie-stained Phos-Tag gels. The experiment was performed two times with consistent results. Molecular weight markers are in kDa. e, Data collection and refinement statistics.
Extended Data Figure 2 PhPARKIN similarity with human and rat PARKIN and map quality.
a, Structure-based sequence alignment of HsPARKIN from human (top), RnPARKIN (middle) and Pediculus humanus corporis (PhPARKIN, bottom). The domains are indicated in boxes coloured according to structural figures. HsPARKIN and PhPARKIN are 45% identical within their crystallized constructs. Secondary structure elements are shown for HsPARKIN Ubl domain (PDB 1IYF (ref. 36)) and for HsPARKIN core domain (4BM9 (ref. 14)). Red asterisks denote the phospho-Ser65 ubiquitin binding pocket, and yellow spheres the residues contacting phosphoUb (see Fig. 2). A black asterisk denotes the catalytic Cys in RING2. b, Stereo representation of the asymmetric unit of PhPARKIN–pUb crystals, showing 2|Fo| − |Fc| electron density at 1σ, in blue for PhPARKIN and in green for phosphoUb. c, Electron density detail, shown as in b, zooming in on phosphoUb phospho-Ser65, in stereo representation. d, Superposition of the two PhPARKIN–pUb complexes in the asymmetric unit, coloured as in Fig. 1. The r.m.s.d. is 0.76 Å. Electron density is missing for parts of the flexible linker between IBR and RING2. e, Superposition of available PARKIN structures (PDB 4BM9 (ref. 14); 4I1H (ref. 15); 4K95 (ref. 16) and two PhPARKIN–pUb complexes) in different colours, showing similar domain positions with respect to each other, with the exception of the IBR domain. Only the structure of full-length RnPARKIN16 contains the Ubl domain.
Extended Data Figure 3 Conservation of phosphoUb interacting residues and biochemical analysis.
a, b, An alignment of PARKIN from species available in Ensembl (http://www.ensembl.org) was curated by removing sequences with truncations or poorly sequenced regions. The residues involved in phosphoUb binding, comprising (a) the phosphate pocket and (b) the pUBH, are shown, and residues contacting phosphoUb are highlighted. c, Comparison of residues from PhPARKIN and HsPARKIN involved in phosphoUb binding. Highlighted in green is the HsPARKIN β-hairpin residue Gly284 (Gly286 in PhPARKIN), which is mutated to Arg in AR-JP. Other β-hairpin patient mutations, L283P (Phe285 in PhPARKIN) and H279P (Ser281 in PhPARKIN), introduce Pro residues, probably distorting the β-hairpin loop. The notation (bb) indicates backbone interaction. d, Fluorescence polarization assays performed with mutant PARKIN and phosphoUb. Assays from two independent experiments were combined to produce Fig. 2e. Measurements were performed in triplicate with error bars given as standard deviation from the mean. e, Coomassie-stained SDS–PAGE gel showing HsPARKIN proteins used in activity assays. f, Normalized proteins for PARKIN activity assays in Fig. 3d. g, Analogous to Fig. 2a, phosphoUb bound to PhPARKIN is shown with Lys residues in stick representation, and free amine groups as blue spheres. Several Lys side chains (Lys29, Lys33, Lys48 and Lys63) were disordered in the electron density maps, suggesting high flexibility and solvent accessibility, and were modelled in their preferred side-chain rotamer for illustrative purposes. When bound to PARKIN, each Lys residue can be ubiquitinated, and the C terminus which is covalently attached to PhPARKIN in the complex structure is probably more flexible and could also be attached to a more proximal (phospho)Ub in a chain. This indicates that PARKIN could interact with phosphoUb-containing polyubiquitin chains that were reported to be the PINK1 substrate on mitochondria7,8,11. Molecular weight markers are in kDa in e and f.
Extended Data Figure 4 PARKIN phosphoUb-binding mutants do not translocate to mitochondria.
a, b, HeLa cells that do not contain detectable levels of PARKIN were transiently transfected with YFP–HsPARKIN wild type and indicated mutants (green). Tom20 staining with anti-Tom20 antibody indicates mitochondria (red). Images are representative of three biological replicates. a, In DMSO-treated control cells, PARKIN does not co-localize with mitochondrial Tom20. b, After treatment with CCCP (10 µM) for 1 h, wild-type PARKIN but not phosphoUb-binding mutants co-localize with Tom20. Scale bars, 10 µm. Split channels are shown with overlay to illustrate co-localization. The YFP channel is identical to Fig. 2g. Digital zoom 10×. c, Quantification of cells with PARKIN localized at mitochondria in b, scored for 150 cells per condition in three biological replicates. PhosphoUb binding mutants do not show sustained mitochondrial localization. Error bars represent standard error of the mean. d, HeLa cells transiently transfected with HsPARKIN wild type and mutants were treated with CCCP and YFP–HsPARKIN immunoprecipitated (see Methods). Whole-cell lysates (WCL) or immunoprecipitates (IP) were western blotted for PARKIN, Tom20 and GAPDH (loading control) as indicated. Ubiquitinated forms of PARKIN and Tom20 can be observed with wild-type PARKIN but not with PARKIN mutants. See also Fig. 2h where the A320R mutant was also included. These experiments were performed three times as biological replicates with similar results. Note that PARKIN autoubiquitination varied and was weaker in some experiments, while Tom20 ubiquitination was more robust. Molecular weight markers are in kDa. See Supplementary Information for uncropped blots.
Extended Data Figure 5 The kink in the pUBH.
a–d, To understand whether the straight pUBH in PhPARKIN was a consequence of a helix-favouring mutation of Gly319 (HsPARKIN) to Ala (Ala321 in PhPARKIN), the HsPARKIN(ΔUbl) G319A mutant was crystallized and a structure determined at 2.35 Å resolution (see Extended Data Fig. 1e and Methods). a, Superposition of HsPARKIN(ΔUbl) G319A and wild-type HsPARKIN(ΔUbl) (PDB 4BM9 (ref. 14)). The structures are virtually identical, both containing a kinked helix. b, c, Electron density detail of the kinked helix for the mutant (b) and wild type (c), shown in stereo representation, with 2|Fo| − |Fc| density contoured at 1σ. The Cβ atom of Ala319 is clearly defined in electron density and does not induce helix straightening in this crystallographic setting. d, Comparison of pUBH helices in HsPARKIN(ΔUbl) G319A (left), HsPARKIN(ΔUbl) wild type (second from left), superposition of the two (second from right), and for comparison the straight pUBH in PhPARKIN–pUb. e, f, Structure of the RBR E3 ligase HHARI (e) in an autoinhibited form (PDB 4KBL (ref. 23)) showing an entirely different RBR module as compared to full-length RnPARKIN (PDB 4K95 (ref. 16)) in f. RING1, IBR, RING2 domains are coloured as for PARKIN in Fig. 1, and other domains (UBA-like domain and Ariadne domain in HHARI) are coloured in red. Interestingly, the pUBH equivalent helix in HHARI is kinked at a similar position as compared to mammalian PARKIN. g, Superposition of structures from e and f on their RING1 domains. In HHARI, the pUBH-equivalent helix kinks in a different manner as compared to HsPARKIN. h, In HHARI, the kinked helix seems to be stabilized by an interaction in cis with the UBA-like domain that binds NEDD8 (ref. 37). The sequence of the helix does not contain Gly residues, but is kinked at a Thr residue (Thr263). It will be interesting to see whether helix straightening occurs in active forms of HHARI that can be induced by binding NEDD8-modified cullins37.
Extended Data Figure 6 Structural detail and B-factor analysis.
a, PhosphoUb-induced pUBH straightening is energetically neutral as the two hydrogen bonds between helix residues and the RING1 core are not lost but only adjusted. Left: full-length RnPARKIN structure (PDB 4K95 (ref. 16)). Right: PhPARKIN–pUb structure. b, Structure of full-length RnPARKIN (PDB 4K95 (ref. 16), left), PhPARKIN–pUb (middle), and a superposition of the two (right), in which Cα atoms of structurally identical, ordered residues of the IBR–REP linker are shown as spheres. The distance between these residues is indicated by a dotted line. The linker sequence is highlighted in Extended Data Fig. 2. c, Structures of PARKIN in which B-factors were refined for individual atoms (PDB 4BM9 (ref. 14); 4I1H (ref. 15)) are shown in a ribbon representation, in which the ribbon thickness indicates B-factors differences, with thin ribbons indicating low and thick ribbon indicating high B-factors. The full-length RnPARKIN structure (PDB 4K95 (ref. 16)) is not included as in this 6.5 Å structure overall B-factors were assigned. d, Structures of PhPARKIN–pUb are shown as in c for each molecule of the asymmetric unit. The REP and RING2 elements are destabilized as indicated by higher B-factors. The rigid core of the protein has shifted from the UPD–RING1–RING2 interfaces (compare c) to the UPD–RING1–IBR–phosphoUb interfaces. B-factor analysis may be distorted by neighbouring molecules and crystal contacts, which are not indicated here.
Extended Data Figure 7 The Ile44 patch is essential for PINK1-mediated phosphorylation of Ub and PARKIN Ubl.
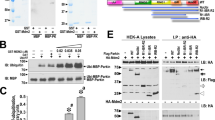
All assays were performed three times with consistent results. a, b, Coomassie-stained PhosTag gels comparing the phosphorylation of (a) the HsPARKIN Ubl domain (amino acids 1–72) and (b) ubiquitin. In both cases, the wild-type form is compared with the I44A mutant form of the protein. GST–TcPINK1 does not efficiently phosphorylate the I44A mutants of ubiquitin or of the HsPARKIN Ubl domain. This is important since the Ile44 patch in the PARKIN Ubl domain is inaccessible and binds to RING1 in the structure of full-length RnPARKIN (PDB 4K95 (ref. 16)) (see Fig. 4a). c, d, Controls for Fig. 4c. Coomassie-stained gel with proteins labelled (c), and full-size blot (d) for Fig. 4c. Molecular weight markers are in kDa for all panels.
Extended Data Figure 8 NMR analysis of phosphorylated Ubl.
a, BEST-TROSY spectra for isotope-labelled HsPARKIN Ubl domain (dark green) and phospho-Ubl domain (light green) with resonances assigned for the Ubl domain. b, Chemical shift perturbation of Ubl with respect to phospho-Ubl, showing significant perturbations in the region of phosphorylation (Ser65), the last β-strand and neighbouring β-strands. Grey bars, exchange broadened resonances. c, Mapping of perturbed resonances onto the previously determined NMR structure of HsPARKIN Ubl (PDB 1IYF (ref. 36)). The perturbed residues cluster in the Ser65-containing loop and in proximity to the Ile44 patch of the Ubl. d, Mapping of the perturbed resonances to the structure of RnPARKIN (PDB 4K95 (ref. 16)) shows that they perturb the interface between the Ubl domain and the PARKIN core. Thus, phosphorylated Ubl may not be able to (re)bind PARKIN at the same binding site.
Extended Data Figure 9 PARKIN phosphoUbl does not compete with phosphoUb.
a, A possible scenario is that the phosphoUbl competes with phosphoUb for the phosphoUb binding site on PARKIN. This could be favoured since the interaction would occur in cis. b, Fluorescence polarization competition experiment increasing the concentration of Ubl, phosphoUbl or phosphoUb with respect to full-length HsPARKIN in the presence of FlAsH-tagged phosphoUb. Measurements were performed in triplicate with error bars given as standard deviation from the mean. While unlabelled phosphoUb competes with labelled phosphoUb in the reaction, Ubl or phosphoUbl do not compete with phosphoUb. c, Binding of FlAsH-phosphoUb to HsPARKIN and phospho-HsPARKIN. The measurements were performed in the same experiment as samples in Fig. 2d. If phosphoUbl interacts with HsPARKIN in cis, phosphoUb binding should be inhibited. In contrast, binding of phosphoUb to phospho-PARKIN is slightly enhanced as also reported in ref. 7. Measurements were performed in triplicate and error bars represent standard deviation from the mean. d, PARKIN phosphorylation assays as in Fig. 4c, including phosphoUbl as well as phosphoUb (both at 10 μM). While addition of phosphoUb induces PARKIN phosphorylation, addition of phosphoUbl does not, indicating that the Ubl domain is not released from the PARKIN core. The experiment has been performed three times with consistent results. Molecular weight markers are in kDa.
Extended Data Figure 10 PhosphoUbl-induced ‘opening’ of PARKIN relies on a phosphate pocket in the UPD.
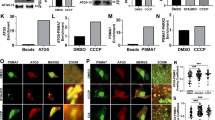
a, A second scenario would be that the phosphorylated Ubl domain rebinds to PARKIN at an alternative site. We previously speculated that the UPD contains a phosphate-binding site that is lined by two AR-JP patient mutations, K161N and K211N, and also contains Arg163 (ref. 14). b, Full-length HsPARKIN mutants as indicated were phosphorylated in vitro with GST–PhPINK1, resolved on SDS–PAGE and western blotted using an anti-pSer65 PARKIN antibody (Abcam cat no. ab154995). These proteins were used in c. c, Ub-vinyl sulfone (Ub-VS) modification of the active site Cys431 of HsPARKIN and HsPARKIN mutants with and without phosphoUb from a time course experiment is assessed on Coomassie-stained gels. Ub-VS reacts with ‘open’ forms of PARKIN, and was previously shown to modify phospho-PARKIN but not phosphoUb-activated PARKIN7. Phospho-PARKIN-mediated opening depends on the phosphate pocket present in the UPD, since phospho-PARKIN K211N, K161N (two AR-JP patient mutations) and R163E abrogated or impaired modification by Ub-VS and do not appear to have an accessible catalytic Cys, while the phosphorylated phosphoUb-binding-deficient mutant K151E is readily modified. The experiment was performed two times with consistent results and gels have been collated from two different assays (indicated by the gap). d, PARKIN ubiquitination reactions in presence of E1, UBE2L3, ubiquitin or ubiquitin S65A, ATP and GST-TcPINK1 for 2 h. PARKIN is activated by phosphoUb or by PARKIN Ubl phosphorylation in absence of phosphoUb (with ubiquitin S65A) (lanes 2/3). Mutants in the UPD phosphate pocket can still be activated by phosphoUb (albeit not to the same extent) (lanes 5, 8, 11) but are inactive when the Ubl domain is phosphorylated (lanes 6, 9, 12). This could suggest that the phosphoUbl binds back to the PARKIN UPD pocket. However, we cannot exclude the possibility that, for example, the linker between the Ubl and the UPD plays a more active role in PARKIN activation. The experiment was performed three times with consistent results. Molecular weight markers are in kDa for b–d.
Supplementary information
Supplementary Information
This file contains the uncropped blots. (PDF 1464 kb)
Rights and permissions
About this article
Cite this article
Wauer, T., Simicek, M., Schubert, A. et al. Mechanism of phospho-ubiquitin-induced PARKIN activation. Nature 524, 370–374 (2015). https://doi.org/10.1038/nature14879
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14879
This article is cited by
-
Reduced mitophagy is an early feature of NAFLD and liver-specific PARKIN knockout hastens the onset of steatosis, inflammation and fibrosis
Scientific Reports (2023)
-
Discovery of Small Molecule PARKIN Activator from Antipsychotic/Anti-neuropsychiatric Drugs as Therapeutics for PD: an In Silico Repurposing Approach
Applied Biochemistry and Biotechnology (2023)
-
Mitophagy and reactive oxygen species interplay in Parkinson’s disease
npj Parkinson's Disease (2022)
-
Mitophagy, a Form of Selective Autophagy, Plays an Essential Role in Mitochondrial Dynamics of Parkinson’s Disease
Cellular and Molecular Neurobiology (2022)
-
Mitochondrial Quality and Quantity Control: Mitophagy Is a Potential Therapeutic Target for Ischemic Stroke
Molecular Neurobiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.