Abstract
Mutation rates vary within genomes, but the causes of this remain unclear1. As many prior inferences rely on methods that assume an absence of selection, potentially leading to artefactual results2, we call mutation events directly using a parent–offspring sequencing strategy focusing on Arabidopsis and using rice and honey bee for replication. Here we show that mutation rates are higher in heterozygotes and in proximity to crossover events. A correlation between recombination rate and intraspecific diversity is in part owing to a higher mutation rate in domains of high recombination/diversity. Implicating diversity per se as a cause, we find an ∼3.5-fold higher mutation rate in heterozygotes than in homozygotes, with mutations occurring in closer proximity to heterozygous sites than expected by chance. In a genome that is a patchwork of heterozygous and homozygous domains, mutations occur disproportionately more often in the heterozygous domains. If segregating mutations predispose to a higher local mutation rate, clusters of genes dominantly under purifying selection (more commonly homozygous) and under balancing selection (more commonly heterozygous), might have low and high mutation rates, respectively. Our results are consistent with this, there being a ten times higher mutation rate in pathogen resistance genes, expected to be under positive or balancing selection. Consequently, we do not necessarily need to evoke extremely weak1,2 selection on the mutation rate to explain why mutational hot and cold spots might correspond to regions under positive/balancing and purifying selection, respectively3,4.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Data deposits
All Illumina reads have been deposited in the BioProject database under accession numbers PRJNA243018, PRJNA252997, PRJNA178613 and PRJNA232554.
References
Hodgkinson, A. & Eyre-Walker, A. Variation in the mutation rate across mammalian genomes. Nature Rev. Genet. 12, 756–766 (2011)
Chen, X. & Zhang, J. No gene-specific optimization of mutation rate in Escherichia coli. Mol. Biol. Evol. 30, 1559–1562 (2013)
Chuang, J. H. & Li, H. Functional bias and spatial organization of genes in mutational hot and cold regions in the human genome. PLoS Biol. 2, e29 (2004)
Martincorena, I., Seshasayee, A. S. N. & Luscombe, N. M. Evidence of non-random mutation rates suggests an evolutionary risk management strategy. Nature 485, 95–98 (2012)
Roach, J. C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010)
Wang, J., Fan, H. C., Behr, B. & Quake, S. R. Genome-wide single-cell analysis of recombination activity and de novo mutation rates in human sperm. Cell 150, 402–412 (2012)
Sung, W., Ackerman, M. S., Miller, S. F., Doak, T. G. & Lynch, M. Drift-barrier hypothesis and mutation-rate evolution. Proc. Natl Acad. Sci. USA 109, 18488–18492 (2012)
Ossowski, S. et al. The rate and molecular spectrum of spontaneous mutations in Arabidopsis thaliana. Science 327, 92–94 (2010)
Jiang, C. et al. Environmentally responsive genome-wide accumulation of de novo Arabidopsis thaliana mutations and epimutations. Genome Res. 24, 1821–1829 (2014)
Cutter, A. D. & Payseur, B. A. Genomic signatures of selection at linked sites: unifying the disparity among species. Nature Rev. Genet. 14, 262–274 (2013)
Lercher, M. J. & Hurst, L. D. Human SNP variability and mutation rate are higher in regions of high recombination. Trends Genet. 18, 337–340 (2002)
Magni, G. E. & Borstel, R. C. V. Different rates of spontaneous mutation during mitosis and meiosis in yeast. Genetics 47, 1097–1108 (1962)
Perry, J. & Ashworth, A. Evolutionary rate of a gene affected by chromosomal position. Curr. Biol. 9, 987 (1999)
Pratto, F. et al. Recombination initiation maps of individual human genomes. Science 346, 1256442 (2014)
Rattray, A., Santoyo, G., Shafer, B. & Strathern, J. N. Elevated mutation rate during meiosis in Saccharomyces cerevisiae. PLoS Genet. 11, e1004910 (2015)
Arbeithuber, B., Betancourt, A. J., Ebner, T. & Tiemann-Boege, I. Crossovers are associated with mutation and biased gene conversion at recombination hotspots. Proc. Natl Acad. Sci. USA 112, 2109–2114 (2015)
Liu, H. et al. Causes and consequences of crossing-over evidenced via a high-resolution recombinational landscape of the honey bee. Genome Biol. 16, 15 (2015)
Amos, W. Heterozygosity and mutation rate: evidence for an interaction and its implications. Bioessays 32, 82–90 (2010)
Hollister, J. D., Ross-Ibarra, J. & Gaut, B. S. Indel-associated mutation rate varies with mating system in flowering plants. Mol. Biol. Evol. 27, 409–416 (2010)
Amos, W. Even small SNP clusters are non-randomly distributed: is this evidence of mutational non-independence? Proc. R. Soc. Lond. B 277, 1443–1449 (2010)
Tian, D. et al. Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes. Nature 455, 105–108 (2008)
Pal, C., Maciá, M. D., Oliver, A., Schachar, I. & Buckling, A. Coevolution with viruses drives the evolution of bacterial mutation rates. Nature 450, 1079–1081 (2007)
Cox, E. C. On the organization of higher chromosomes. Nature New Biol. 239, 133–134 (1972)
Shee, C., Gibson, J. L. & Rosenberg, S. M. Two mechanisms produce mutation hotspots at DNA breaks in Escherichia coli. Cell Rep. 2, 714–721 (2012)
Pál, C. & Hurst, L. D. Evidence for co-evolution of gene order and recombination rate. Nature Genet. 33, 392–395 (2003)
Gladyshev, E. & Kleckner, N. Direct recognition of homology between double helices of DNA in Neurospora crassa. Nature Commun. 5, 3509 (2014)
Boateng, K. A., Bellani, M. A., Gregoretti, I. V., Pratto, F. & Camerini-Otero, R. D. Homologous pairing preceding SPO11-mediated double-strand breaks in mice. Dev. Cell 24, 196–205 (2013)
Malkova, A. & Haber, J. E. Mutations arising during repair of chromosome breaks. Annu. Rev. Genet. 46, 455–473 (2012)
Ichijima, Y. et al. MDC1 directs chromosome-wide silencing of the sex chromosomes in male germ cells. Genes Dev. 25, 959–971 (2011)
Cao, J. et al. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nature Genet. 43, 956–963 (2011)
Gao, Z.-Y. et al. Dissecting yield-associated loci in super hybrid rice by resequencing recombinant inbred lines and improving parental genome sequences. Proc. Natl Acad. Sci. USA 110, 14492–14497 (2013)
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at http://arxiv.org/abs/1303.3997 (2013)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Ghoneim, D. H., Myers, J. R., Tuttle, E. & Paciorkowski, A. R. Comparison of insertion/deletion calling algorithms on human next-generation sequencing data. BMC Res. Notes 7, 864 (2014)
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013)
Li, M. & Stoneking, M. A new approach for detecting low-level mutations in next-generation sequence data. Genome Biol. 13, R34 (2012)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Keightley, P. D., Ness, R. W., Halligan, D. L. & Haddrill, P. R. Estimation of the spontaneous mutation rate per nucleotide site in a Drosophila melanogaster full-sib family. Genetics 196, 313–320 (2014)
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279 (2004)
Yang, Z. PAML 4: Phylogenetic Analysis by Maximum Likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007)
R Development Core Team . R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2013)
North, B. V., Curtis, D. & Sham, P. C. A note on the calculation of empirical P values from Monte Carlo procedures. Am. J. Hum. Genet. 72, 498–499 (2003)
Acknowledgements
This work was supported by the National Natural Science Foundation of China (91331205, 91231102 and 31170210), Program for Changjiang Scholars and Innovative Research Team (IRT_14R27) and Jiangsu Collaborative Innovation Center for Modern Crop Production to D.T., S.H. and J.-Q.C., respectively.
Author information
Authors and Affiliations
Contributions
D.T., L.D.H., S.Y. and J.-Q.C. designed the experiments. S.Y., L.W., J.H., X.Z., L.D.H. and Y.Y. performed the experiments and analysed the data. L.D.H. and D.T. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Details of materials and methods.
a, Schematic diagram of the detection of de novo mutations. b, The calculation of the expected mutations in the meiosis of F2→F3 (EM3) and F3→F4 (EM4). For further explanation see Methods.
Extended Data Figure 2 Mutational properties.
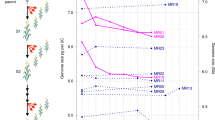
a, Spectra of nucleotide substitutions in Arabidopsis and rice. b, Co-occurrence of mutations and crossover break points in bees. By using the sequence data of 43 honey bee drones and their 3 corresponding queens17, a total of 27 base and 8 indel mutations were detected. Of note, 2 of 35 mutations are found in close proximity with crossover break points in the same sample (distance < 2 kb; P = 0.0012 with 10,000 randomizations), these being illustrated here. The crossover event is between the red and blue line with marker positions annotated. The positions of the mutations are annotated with arrows. c, A schematic diagram of the genomic structures and the possible pairings of two homologous chromosomes during the meiosis at two mutated LRR-TM genes (top) and one mutated NBS-LRR gene (bottom). The top panel shows the genomic structures between Col and Ler at the loci of AT3G23110, the receptor-like protein 37 with a non-synonymous mutation (Chr3:8224726, T→C) at sample of c74, and AT3G23120, the receptor-like protein 38 with a deletion mutation (Chr3:8228194, Del:C, frameshift) at sample of c70. The bottom panel illustrates the genomic structures between two Col chromosomes at AT1G59780 and the mutations detected in a homozygous plant of Col8. Red arrows represent the position of mutation; the hatched areas indicate the highly similar sequences, the other regions being highly diversified; the dotted lines indicate the paired length of the homologues at the highly identical regions. During meiosis, possible pairings between parental chromosomes are illustrated, where the loops indicate the unpaired regions.
Extended Data Figure 3 Correlation between mutations, recombination events, diversity and divergence.
a, The relationship between nucleotide diversity (Col versus Ler) and recombination rate. When the chromosomes were dissected into 100 kb non-overlapping windows, the diversity (polymorphism density) between Col and Ler and the recombination rates in 67 F2 and 32 F4 plants were calculated for each window. When sorting the windows by the diversity and dividing them into 8 equal intervals (for example, from 0 to 0.001, 0.001 to 0.002, 0.002 to 0.003, and so on), the relationships between the average diversity and recombination rate is displayed. Error bars indicate standard error of the mean. b, The relationship between diversity and divergence. The red line represents standard linear regression and is for illustrative purposes only. The statistic is the result of Spearman’s rank correlation. c, Relationship between mutation and distance to polymorphic sites. The mutation data were collected from our 67 F2 samples. Window 0 in the x-axis is the 2 × 100 bp sequence surrounding the position of any given de novo mutation and 1–9 is 100–900 bp away from the mutation on both sides. For each window of 2 × 100 bp sequence, the average diversity is calculated. The black squares denote the average pairwise diversity among the published 80 Arabidopsis ecotypes; the red circles denote the average diversity between Col and the 80 ecotypes; the blue triangles denote the average diversity between Ler and the 80 ecotypes. Error bars indicate standard error of the mean. d, Distribution of the mutations on the chromosomes. The grey vertical bars in the chromosomes denote the position of all collected mutations. When the chromosomes were dissected into 1 Mb non-overlapping windows, the mutation numbers (blue shadow in the figure) were counted in each window. The red lines denote the average pairwise diversity among the published 80 Arabidopsis ecotypes.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-9. (PDF 3767 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Yang, S., Wang, L., Huang, J. et al. Parent–progeny sequencing indicates higher mutation rates in heterozygotes. Nature 523, 463–467 (2015). https://doi.org/10.1038/nature14649
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14649
This article is cited by
-
Reply to: Re-evaluating evidence for adaptive mutation rate variation
Nature (2023)
-
High level of somatic mutations detected in a diploid banana wild relative Musa basjoo
Molecular Genetics and Genomics (2023)
-
Genome-wide identification, characterization, and evolutionary analysis of NBS genes and their association with disease resistance in Musa spp.
Functional & Integrative Genomics (2023)
-
Spontaneous mutation rate estimates for the principal malaria vectors Anopheles coluzzii and Anopheles stephensi
Scientific Reports (2022)
-
Genetic and chemotherapeutic influences on germline hypermutation
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.