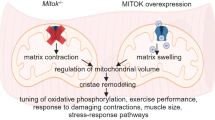
Abstract
Intracellular energy distribution has attracted much interest and has been proposed to occur in skeletal muscle via metabolite-facilitated diffusion1,2; however, genetic evidence suggests that facilitated diffusion is not critical for normal function3,4,5,6,7. We hypothesized that mitochondrial structure minimizes metabolite diffusion distances in skeletal muscle. Here we demonstrate a mitochondrial reticulum providing a conductive pathway for energy distribution, in the form of the proton-motive force, throughout the mouse skeletal muscle cell. Within this reticulum, we find proteins associated with mitochondrial proton-motive force production preferentially in the cell periphery and proteins that use the proton-motive force for ATP production in the cell interior near contractile and transport ATPases. Furthermore, we show a rapid, coordinated depolarization of the membrane potential component of the proton-motive force throughout the cell in response to spatially controlled uncoupling of the cell interior. We propose that membrane potential conduction via the mitochondrial reticulum is the dominant pathway for skeletal muscle energy distribution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bessman, S. P. & Geiger, P. J. Transport of energy in muscle: The phosphorylcreatine shuttle. Science 211, 448–452 (1981)
Wittenberg, J. B. Myoglobin-facilitated oxygen diffusion: Role of myoglobin in oxygen entry into muscle. Physiol. Rev. 50, 559–632 (1970)
Garry, D. J. et al. Mice without myoglobin. Nature 395, 905–908 (1998)
van Deursen, J. et al. Skeletal muscles of mice deficient in muscle creatine kinase lack burst activity. Cell 74, 621–631 (1993)
Lygate, C. A. et al. Living without creatine: unchanged exercise capacity and response to chronic myocardial infarction in creatine-deficient mice. Circ. Res. 112, 945–955 (2013)
Kernec, F., Unlu, M., Labeikovsky, W., Minden, J. S. & Koretsky, A. P. Changes in the mitochondrial proteome from mouse hearts deficient in creatine kinase. Physiol. Genomics 6, 117–128 (2001)
Glancy, B. et al. In vivo microscopy reveals extensive embedding of capillaries within the sarcolemma of skeletal muscle fibers. Microcirculation 21, 131–147 (2014)
Narayan, K. et al. Multi-resolution correlative focused ion beam scanning electron microscopy: Applications to cell biology. J. Struct. Biol. 185, 278–284 (2014)
Dahl, R. et al. Three dimensional reconstruction of the human skeletal muscle mitochondrial network as a tool to assess mitochondrial content and structural organization. Acta Physiol. 213, 145–155 (2015)
Bakeeva, L. E., Chentsov, Y. S. & Skulachev, V. P. Ontogenesis of mitochondrial reticulum in rat diaphragm muscle. Eur. J. Cell Biol. 25, 175–181 (1981)
Bakeeva, L. E., Chentsov Yu, S. & Skulachev, V. P. Mitochondrial framework (reticulum mitochondriale) in rat diaphragm muscle. Biochim. Biophys. Acta 501, 349–369 (1978)
Kayar, S. R., Hoppeler, H., Mermod, L. & Weibel, E. R. Mitochondrial size and shape in equine skeletal muscle: a three-dimensional reconstruction study. Anat. Rec. 222, 333–339 (1988)
Kirkwood, S. P., Munn, E. A. & Brooks, G. A. Mitochondrial reticulum in limb skeletal muscle. Am. J. Physiol. 251, C395–C402 (1986)
Kirkwood, S. P., Packer, L. & Brooks, G. A. Effects of endurance training on a mitochondrial reticulum in limb skeletal muscle. Arch. Biochem. Biophys. 255, 80–88 (1987)
Ogata, T. & Yamasaki, Y. Scanning electron-microscopic studies on the three-dimensional structure of mitochondria in the mammalian red, white and intermediate muscle fibers. Cell Tissue Res. 241, 251–256 (1985)
Denk, W. & Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol. 2, e329 (2004)
Bakalar, M. et al. Three-dimensional motion tracking for high-resolution optical microscopy, in vivo. J. Microsc. 246, 237–247 (2012)
Rothstein, E. C., Carroll, S., Combs, C. A., Jobsis, P. D. & Balaban, R. S. Skeletal muscle NAD(P)H two-photon fluorescence microscopy in vivo: topology and optical inner filters. Biophys. J. 88, 2165–2176 (2005)
Bubenzer, H. J. Die Dunnen Und Die Dicken Muskelfasern Des Zwerchfells Der Ratte. Z. Zellforsch. Mikrosk. Anat. 69, 520–550 (1966)
Skulachev, V. P. Energy transformation in the respiratory chain. Curr. Top. Bioenerg. 4, 127–190 (1971)
Amchenkova, A. A., Bakeeva, L. E., Chentsov, Y. S., Skulachev, V. P. & Zorov, D. B. Coupling membranes as energy-transmitting cables. I. Filamentous mitochondria in fibroblasts and mitochondrial clusters in cardiomyocytes. J. Cell Biol. 107, 481–495 (1988)
Picard, M. et al. Acute exercise remodels mitochondrial membrane interactions in mouse skeletal muscle. J. Appl. Physiol. 115, 1562–1571 (2013)
Saito, A., Smigel, M. & Fleischer, S. Membrane junctions in the intermembrane space of mitochondria from mammalian tissues. J. Cell Biol. 60, 653–663 (1974)
Eisner, V., Lenaers, G. & Hajnoczky, G. Mitochondrial fusion is frequent in skeletal muscle and supports excitation-contraction coupling. J. Cell Biol. 205, 179–195 (2014)
Chalmers, S. et al. Selective uncoupling of individual mitochondria within a cell using a mitochondria-targeted photoactivated protonophore. J. Am. Chem. Soc. 134, 758–761 (2012)
Fan, X., Hussien, R. & Brooks, G. A. H2O2-induced mitochondrial fragmentation in C2C12 myocytes. Free Radic. Biol. Med. 49, 1646–1654 (2010)
Weibel, E. R. & Hoppeler, H. Exercise-induced maximal metabolic rate scales with muscle aerobic capacity. J. Exp. Biol. 208, 1635–1644 (2005)
Bakalar, M. et al. Three-dimensional motion tracking for high-resolution optical microscopy, in vivo. J. Microsc. 246, 237–247 (2012)
Narayan, K. et al. Multi-resolution correlative focused ion beam scanning electron microscopy: applications to cell biology. J. Struct. Biol. 185, 278–284 (2014)
Murphy, G. E. et al. Correlative 3D imaging of whole mammalian cells with light and electron microscopy. J. Struct. Biol. 176, 268–278 (2011)
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996)
Glancy, B. & Balaban, R. S. Protein composition and function of red and white skeletal muscle mitochondria. Am. J. Physiol. Cell Physiol. 300, C1280–C1290 (2011)
Balaban, R. S., Mootha, V. K. & Arai, A. Spectroscopic determination of cytochrome c oxidase content in tissues containing myoglobin or hemoglobin. Anal. Biochem. 237, 274–278 (1996)
Wittig, I., Karas, M. & Schagger, H. High resolution clear native electrophoresis for in-gel functional assays and fluorescence studies of membrane protein complexes. Mol. Cell. Proteomics 6, 1215–1225 (2007)
Hein, B. et al. Stimulated emission depletion nanoscopy of living cells using SNAP-Tag fusion proteins. Biophys. J. 98, 158–163 (2010)
Acknowledgements
We would like to thank E. Tyler for his assistance with image rendering, J. Taraska for providing experimental advice, T. Karpova and the NCI Core Fluorescence Imaging Facility for access to a microscope with a 355-nm laser, and P. Kellman for assistance with image analysis. We would also like to thank S. Caldwell and R. Hartley for providing us with MitoPhotoDNP. This study was supported by intramural funding of the Division of Intramural Research, National Heart, Lung, and Blood Institute and the Center for Cancer Research, National Cancer Institute.
Author information
Authors and Affiliations
Contributions
R.S.B., S.S., B.G. and L.M.H. designed and L.M.H performed the FIB-SEM experiments. R.S.B. and B.G. designed and B.G. performed the MPM experiments. R.S.B., B.G., Z.-X.Y. and D.M. designed and B.G., Z.-X.Y. and D.M. performed the dual immunolabelling experiments. P.S.C. performed the TEM imaging. R.S.B. and B.G. performed the segmentations and analysed the data. R.S.B., C.A.C. and B.G. designed and performed the isolated muscle fibre experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Mitochondrial coupling assessment.
a, b, Single image from the middle of a FIB-SEM muscle volume including two nuclei (N) and a blood vessel (V). a, Direct coupling. PVM tagged green are those where a single mitochondrion projects from a PVM into an IBM with a continuous outer membrane (see inset) within the imaged volume. Red-tagged PVM did not project into IBM. Blue-tagged PVM continued out of the field of view and could not be classified. b, Electron-dense contact site (EDCS) coupling. Magenta-tagged mitochondria were connected by EDCS (see yellow arrows in inset). Additional green dots were added to PVM which project into IBM (see a). Greater than 99% of tracked PVM were coupled by EDCS. c, d, Single image from the middle of the intra-fibrillar region of a FIB-SEM volume. c, Direct coupling. FPM tagged green are those where a single mitochondrion projects from an FPM into an IBM with a continuous outer membrane (see inset). Yellow-tagged FPM did not project into IBM. d, EDCS coupling. FPM tagged blue were connected to an adjacent FPM through an EDCS (see yellow arrows in inset). Images are representative of 4 samples with significant paravascular and intra-fibrillar mitochondrial content from 4 animals. e, A longitudinal TEM mouse tibialis anterior muscle image showing the close association between adjacent mitochondria. f, Close up view of an EDCS highlighted in e showing the convergence of mitochondrial membranes between two adjacent mitochondria. Scale bars, 750 nm.
Extended Data Figure 2 Multi-photon microscopy (MPM) images of fresh muscle fibres in situ.
a, Endogenous mitochondrial NAD(P)H signal from a single XY image within a 3D volume of a muscle fibre. In this orientation, paravascular mitochondria (PVM) are apparent as clusters around the embedded capillary (V) and nuclei (N). Fibre parallel mitochondria (FPM) are seen as horizontal lines while I-band mitochondria (IBM) are seen as discrete spots. There are very few vertical lines in this image due to the infrequency of cross-fibre connection mitochondria (CFCM). The full volume can be seen in Supplementary Video 3. b, A YZ image from the same fibre volume as in a. PVM appear lateral to the embedded capillary (V), while IBM appear as vertical lines and FPM are seen as discrete spots. Note the lack of horizontal lines (CFCM) in this image or the accompanying video (Supplementary Video 4). c, An XZ image of the same muscle volume as in a and b. PVM are located on the cell periphery; IBM are seen as vertical lines; FPM are seen as horizontal lines; and CFCM as discrete spots. In the inset, pairs of IBM projecting out from PVM are highlighted. d, A 3D rendering of the endogenous mitochondrial NAD(P)H signal within the muscle fibre shown in a–c. 360° views of this 3D rendering are shown in Supplementary Video 5. e, A view of the interior of the 3D rendering from d showing the regularity of the mitochondrial network within skeletal muscle. The field of view for the muscle volume in this image is 102.4 × 51.2 × 36.8 µm in x, y and z, respectively. Scale bars, 15 μm. Mean fibre volume from these MPM images was 159,431 ± 15,507 μm3. Images are representative of 4 fibres from 3 mice.
Extended Data Figure 3 Super resolution microscopy allows improved visualization of individual mitochondria.
a, Single confocal microscopy image of an isolated muscle fibre loaded with mitochondrial membrane potential dye, TMRM. PVM, paravascular mitochondria; IFM, intra-fibrillar mitochondria; V, vessel. b, Single stimulated emission depletion (STED) microscopy image of the same fibre as in a. c, The average of a confocal microscopy image stack of a TMRM loaded isolated muscle fibre. d, The average of a STED microscopy image stack of the same fibre as in c. For c and d, image volume was 24.30 × 27.47 × 1.75 µm in x, y and z, and images shown are the average of 9 images taken with 219 nm z-steps. Confocal images were acquired before STED images at each image depth. All images were acquired with x and y pixel sizes of 30 nm. Scale bars, 2 μm. As most muscle mitochondria are larger than 300 nm, the improved resolution with STED is primarily noted by the increased clarity of the spaces between individual mitochondria, particularly the PVM. Images are representative of 13 fibres from 3 mice.
Extended Data Figure 4 Complex V expression is relatively higher than complex IV in the muscle fibre interior.
a, Representative confocal microscopy image of a cross-section of a muscle fibre immunostained for complex IV. b, The same muscle section depicted in a immunostained for complex V. Scale bars, 15 μm. All images analysis was performed on the raw images. Brightness and contrast have been increased for the images shown here to improve presentation. c, Distance map from the boundary of the cell (yellow line) towards the interior of the fibre where image intensity corresponds to distance from the cell boundary. The mean fluorescence signal for all pixels of a given distance from the cell boundary was assessed for both complex IV and complex V. Images representative of 4 fibres from 4 mice. d, Results from intensity profile analysis of 4 muscle fibres from 4 animals show relatively increased complex V in the fibre interior or, conversely, relatively higher complex IV near the fibre periphery. The red, solid line shows the mean intensity profile for complex V from the outside to the inside of the fibre. The green, dotted line shows the mean intensity profile for complex IV from the outside to the inside of the fibre. Values are normalized to the maximal intensity from each intensity profile to allow comparison of images of varying intensities. Grey shading around each line shows the standard error.
Extended Data Figure 5 Primary antibody specificity.
a, Coomassie-stained CN-PAGE gel after transferring isolated mouse skeletal muscle mitochondrial proteins to PVDF membrane for western blotting. Mitochondrial oxidative phosphorylation complexes I–V can be resolved as individual bands. b, Western blot dual immunostained for complex V, β-subunit (upper band), and complex IV, subunit I (lower band), with the same primary antibodies as used for the muscle section dual immunostaining as shown in Fig. 3 and Extended Data Fig. 4.
Extended Data Figure 6 Sarcolemmal capillary grooves remain even after removal of capillaries.
a, Longitudinal view of a 3D rendering of the outer 10 µm of an enzymatically isolated muscle fibre imaged as the mitochondrial TMRM signal by confocal microscopy. Blood vessels (V) and nuclei (N) are apparent as negative signals. b, c, Rotation of the volume in a around the longitudinal (b) and cross-sectional (c) axes reveals the grooves in the sarcolemma where capillaries were once present. Scale bars, 10 μm. Images representative of 11 fibres from 4 mice.
Extended Data Figure 7 Additional analyses of mitochondrial membrane potential conduction.
a, Confocal image of an isolated muscle fibre loaded with TMRM showing regions chosen for representative line profile analysis. Images are representative of 11 fibres from 4 mice. b, Line profile of TMRM fluorescence in the PVM+nuclear (N) region of the cell (marked by yellow dotted lines in a) before and after activation of MitoPhotoDNP shows a decrease in PVM signal, increase in N signal, and no change outside of the cell where a blood vessel (V) was located before enzymatic fibre isolation. c, Line profile of TMRM fluorescence in the region of the cell irradiated with UV light (irradiated, marked by white dotted lines in a) showing a decrease in mitochondrial signal (peaks) and an increase in cytosolic signal (troughs) after MitoPhotoDNP activation. d, Line profile of TMRM fluorescence in the non-irradiated IFM region of the cell (intra-fibrillar, marked by red dotted lines in a) showing a decrease in mitochondrial signal (peaks) and an increase in cytosolic signal (troughs) after MitoPhotoDNP activation. c and d take advantage of the regular pattern of IBM and cytosol in the intra-fibrillar space. However, any FPM in the regions selected for analysis may confound these results. A more robust analysis that selects only the IBM is shown in e and f. e, Representative fast Fourier transform (FFT) power spectrum of IFM TMRM signal reveals a 61 ± 3% decrease (n = 9 experiments) in the amplitude of IBM component (0.5 μm−1 frequency) after MitoPhotoDNP activation. Inset: log scale full FFT power spectrum. f, Representative FFT analyses of IFM mitochondria reveals no change (pre/post = 1.0 ± 0.03, n = 5 experiments) in the TMRM amplitude of IBM (0.5 μm−1 frequency) after UV exposure when MitoPhotoDNP is not present. Inset: log scale full FFT power spectrum.
Extended Data Figure 8 Effect of UV exposure on membrane potential without MitoPhotoDNP.
a, Representative heat map image of an isolated mouse soleus muscle fibre loaded with 5 nM TMRM. N, nucleus; V, vessel. b, c, Image from a thresholded to include only the TMRM signal in the mitochondria (b) or cytosol+nuclei (c). d–f, Whole cell (d), mitochondrial (e) and cytosolic+nuclear (f) TMRM signals immediately after UV exposure in the centre region of the cell (outlined by the white dotted lines). g–i, Post/pre ratiometric whole-cell (g), mitochondrial (h), and cytosolic+nuclear (i) TMRM images showing little change outside of the UV-exposed area. j, Greyscale image of a highlighting the different cell regions used for quantitative analysis. All pixels were analysed. k, TMRM signal decreased in both the mitochondria and cytosol in the UV-exposed region indicative of slight photobleaching. Images are representative of 10 fibres from 3 mice. Data represent the mean ± s.e. from 10 experiments from 3 animals. Asterisk indicates significantly different (ANOVA, P < 0.05) from a post/pre ratio of 1.0. Plus symbol indicates significantly different (P < 0.05) from IFM+UV region. See Supplementary Video 10 for time course video.
Extended Data Figure 9 UV light was restricted to the drawn ROI.
Difference XY image of a uniformly blue fluorescent slide before and after exposure to UV light in the drawn ROI (white outline). Inset: YZ view shows that the UV light was largely maintained to the ROI drawn (black line). Scale bars, 5 μm. Images are representative of 3 experiments.
Related audio
Supplementary information
Supplementary Information
This file contains the full gel image of data shown in Extended Data Figure 5a (highlighted by red box) and full western blot image of data shown in Extended Data Figure 5a (highlighted by red box). (PDF 549 kb)
Begins with a 3D FIB/SEM image stack of mouse Tibialis anterior muscle.
Longitudinal images are shown and time represents sequential images moving deeper into the fiber (15 nm steps). These are the data from which Figures 1 and 2a-f were created. Then the same image stack after automated mitochondrial segmentation is shown followed by a 3D rendering of the segmented mitochondria. Close up views of IBM projections from PVM as well as the fiber interior are also shown, respectively. Raw image stack is available as Supplementary Dataset 1 at http://labalaban.nhlbi.nih.gov/files/SuppDataset.tif. (AVI 21769 kb)
3D rendering of the data in Figure 2a.
3D rendering of the data in Figure 2a. (AVI 15661 kb)
Longitudinal MPM image stack for the data shown Extended Data Figure 2.
Time represents sequential images moving deeper into the muscle fiber (100 nm steps). (AVI 32049 kb)
Cross-sectional view of the MPM image stack shown in Extended Data Figure 2.
Time represents sequential images moving down the barrel of the fiber (100 nm steps). (AVI 31990 kb)
3D rendering of the MPM muscle volume shown in Extended Data Figure 2.
3D rendering of the MPM muscle volume shown in Extended Data Figure 2. (AVI 23510 kb)
Longitudinal STED image stack of an isolated muscle cell stained with mitochondrial membrane potential probe TMRM.
Time represents sequential images moving deeper into the muscle fiber (219 nm steps). Images shown are raw data. (AVI 3270 kb)
360° rotation of a 3D rendering of the ratiometric determination of the spatial distribution of mitochondrial Complexes IV and V as shown in Figure 3.
360° rotation of a 3D rendering of the ratiometric determination of the spatial distribution of mitochondrial Complexes IV and V as shown in Figure 3. (AVI 11637 kb)
Individual channel and merged images of the raw data for the muscle section dual immunostaining shown in Figure 3.
Upper left (green) – Complex IV, upper right (red) – Complex V, lower left (blue) – nuclei, lower right – merged images. (AVI 2616 kb)
Timecourse loop of confocal microscopy images of the TMRM signal in a live, isolated muscle fiber before and after UV light induced release of MitoPhotoDNP in the cell interior.
Timecourse loop of confocal microscopy images of the TMRM signal in a live, isolated muscle fiber before and after UV light induced release of MitoPhotoDNP in the cell interior. (AVI 6552 kb)
Timecourse loop of confocal microscopy images of the TMRM signal in a live, isolated muscle fiber before and after UV light exposure in the cell interior of a fiber without MitoPhotoDNP.
Timecourse loop of confocal microscopy images of the TMRM signal in a live, isolated muscle fiber before and after UV light exposure in the cell interior of a fiber without MitoPhotoDNP. (AVI 15427 kb)
Rights and permissions
About this article
Cite this article
Glancy, B., Hartnell, L., Malide, D. et al. Mitochondrial reticulum for cellular energy distribution in muscle. Nature 523, 617–620 (2015). https://doi.org/10.1038/nature14614
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14614
This article is cited by
-
Hypoxia induces mitochondrial protein lactylation to limit oxidative phosphorylation
Cell Research (2024)
-
Cholic and deoxycholic acids induce mitochondrial dysfunction, impaired biogenesis and autophagic flux in skeletal muscle cells
Biological Research (2023)
-
Exercise metabolism and adaptation in skeletal muscle
Nature Reviews Molecular Cell Biology (2023)
-
Leafhopper males compensate for unclear directional cues in vibration-mediated mate localization
Scientific Reports (2023)
-
Changes in cAMP signaling are associated with age-related downregulation of spontaneously beating atrial tissue energetic indices
GeroScience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.